+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | BcsH-BcsD 'beads-on-a-string' filament, local refine | |||||||||
 Map data Map data | BcsHD 'beads-on-a-string' filaments : Unsharpened locally refined map of a BcsH-bound dimer of BcsD octamers | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Bacterial cytoskeleton / cellulose secretion / STRUCTURAL PROTEIN | |||||||||
| Function / homology | Cellulose-complementing protein / Cellulose-complementing protein A / Cellulose synthase operon protein D, bacterial / Cellulose synthase subunit D superfamily / Cellulose synthase subunit D / cellulose biosynthetic process / Uncharacterized protein / Cellulose biosynthesis protein Function and homology information Function and homology information | |||||||||
| Biological species |  Komagataeibacter hansenii ATCC 23769 (bacteria) Komagataeibacter hansenii ATCC 23769 (bacteria) | |||||||||
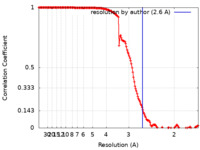
| Method | single particle reconstruction / cryo EM / Resolution: 2.6 Å | |||||||||
 Authors Authors | Krasteva PV / Abidi W / Decossas M | |||||||||
| Funding support | European Union,  France, 2 items France, 2 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: Bacterial crystalline cellulose secretion via a supramolecular BcsHD scaffold. Authors: Wiem Abidi / Marion Decossas / Lucía Torres-Sánchez / Lucie Puygrenier / Sylvie Létoffé / Jean-Marc Ghigo / Petya V Krasteva /  Abstract: Cellulose, the most abundant biopolymer on Earth, is not only the predominant constituent of plants but also a key extracellular polysaccharide in the biofilms of many bacterial species. Depending on ...Cellulose, the most abundant biopolymer on Earth, is not only the predominant constituent of plants but also a key extracellular polysaccharide in the biofilms of many bacterial species. Depending on the producers, chemical modifications, and three-dimensional assemblies, bacterial cellulose (BC) can present diverse degrees of crystallinity. Highly ordered, or crystalline, cellulose presents great economical relevance due to its ever-growing number of biotechnological applications. Even if some acetic acid bacteria have long been identified as BC superproducers, the molecular mechanisms determining the secretion of crystalline versus amorphous cellulose remain largely unknown. Here, we present structural and mechanistic insights into the role of the accessory subunits BcsH (CcpAx) and BcsD (CesD) that determine crystalline BC secretion in the lineage. We show that oligomeric BcsH drives the assembly of BcsD into a supramolecular cytoskeletal scaffold that likely stabilizes the cellulose-extruding synthase nanoarrays through an unexpected inside-out mechanism for secretion system assembly. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15039.map.gz emd_15039.map.gz | 174.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15039-v30.xml emd-15039-v30.xml emd-15039.xml emd-15039.xml | 25.1 KB 25.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15039_fsc.xml emd_15039_fsc.xml | 15.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_15039.png emd_15039.png | 94 KB | ||
| Masks |  emd_15039_msk_1.map emd_15039_msk_1.map | 347.6 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-15039.cif.gz emd-15039.cif.gz | 7.1 KB | ||
| Others |  emd_15039_additional_1.map.gz emd_15039_additional_1.map.gz emd_15039_half_map_1.map.gz emd_15039_half_map_1.map.gz emd_15039_half_map_2.map.gz emd_15039_half_map_2.map.gz | 328.2 MB 323.1 MB 323.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15039 http://ftp.pdbj.org/pub/emdb/structures/EMD-15039 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15039 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15039 | HTTPS FTP |
-Related structure data
| Related structure data |  7zzqMC  7zzyC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15039.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15039.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | BcsHD 'beads-on-a-string' filaments : Unsharpened locally refined map of a BcsH-bound dimer of BcsD octamers | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.839 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_15039_msk_1.map emd_15039_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: BcsHD 'beads-on-a-string' filaments : Autosharpened locally refined map...
| File | emd_15039_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | BcsHD 'beads-on-a-string' filaments : Autosharpened locally refined map of a BcsH-bound dimer of BcsD octamers | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_15039_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: BcsHD 'beads-on-a-string' filaments : half map B
| File | emd_15039_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | BcsHD 'beads-on-a-string' filaments : half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : BcsHD 'beads-on-a-string' cis-filaments
| Entire | Name: BcsHD 'beads-on-a-string' cis-filaments |
|---|---|
| Components |
|
-Supramolecule #1: BcsHD 'beads-on-a-string' cis-filaments
| Supramolecule | Name: BcsHD 'beads-on-a-string' cis-filaments / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 Details: G. hansenii BcsHD complex purified after the co-expression of hexahistidine tagged BcsH C-terminal domain and full-length BcsD. IMAC purification via BcsH, tag and linker removed during purification |
|---|---|
| Source (natural) | Organism:  Komagataeibacter hansenii ATCC 23769 (bacteria) / Strain: ATCC 23769 Komagataeibacter hansenii ATCC 23769 (bacteria) / Strain: ATCC 23769 |
-Macromolecule #1: Cellulose biosynthesis protein
| Macromolecule | Name: Cellulose biosynthesis protein / type: protein_or_peptide / ID: 1 Details: BcsD from Gluconacetobacter hansenii ATCC 23769 expressed recombinantly in BL21 Star DE3 cells Number of copies: 18 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Komagataeibacter hansenii ATCC 23769 (bacteria) Komagataeibacter hansenii ATCC 23769 (bacteria) |
| Molecular weight | Theoretical: 17.532963 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSTIFEKKP DFTLFLQTLS WEIDDQVGIE VRNELLREVG RGMGTRIMPP PCQTVDKLQI ELNALLALIG WGTVTLELLS EDQSLRIVH ENLPQVGSAG EPSGTWLAPV LEGLYGRWVT SQAGAFGDYV VTRDVDAEDL NAVPRQTIIM YMRVRSSAT UniProtKB: Cellulose biosynthesis protein |
-Macromolecule #2: BcsH fragment
| Macromolecule | Name: BcsH fragment / type: protein_or_peptide / ID: 2 Details: BcsH C-terminal domain from G. hansenii expressed from a pProEx-Htb vector with a HRV3c-cleavable hexahistidine tag Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Komagataeibacter hansenii ATCC 23769 (bacteria) Komagataeibacter hansenii ATCC 23769 (bacteria) |
| Molecular weight | Theoretical: 9.919921 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSYYHHHHHH DYDIPTTLEV LFQGPMGSTK TDTNSSQASR PGSPVASPDG SPTMAEVFMT LGGRATELLS PRPSLREALL RRRENEEES UniProtKB: Uncharacterized protein |
-Macromolecule #3: water
| Macromolecule | Name: water / type: ligand / ID: 3 / Number of copies: 105 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 8 / Details: 20 mM NaCl, 100 mM NaCL |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV |
| Details | Filaments of various sizes eluting in the void volume of a Superdex 200 Increase size-exclusion column. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 49.9 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.75 µm / Nominal defocus min: 0.48 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A / Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Details | Model building with Coot, refinement with Phenix and Namdinator |
| Refinement | Space: REAL / Protocol: BACKBONE TRACE |
| Output model |  PDB-7zzq: |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)