[English] 日本語
 Yorodumi
Yorodumi- EMDB-14004: Specific features and methylation sites of a plant 80S ribosome -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Specific features and methylation sites of a plant 80S ribosome | |||||||||||||||
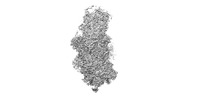
 Map data Map data | 60S, 40S body and 40S head combined map. | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Solanum lycopersicum / cytosolic ribosome / 80S / plant / rRNA / RIBOSOME | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationpectinesterase / pectinesterase activity / : / cell wall modification / pectin catabolic process / MAP kinase activity / endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / protein-RNA complex assembly / translation regulator activity / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) ...pectinesterase / pectinesterase activity / : / cell wall modification / pectin catabolic process / MAP kinase activity / endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / protein-RNA complex assembly / translation regulator activity / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of LSU-rRNA / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / chloroplast / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ribosomal large subunit biogenesis / maturation of SSU-rRNA / small-subunit processome / modification-dependent protein catabolic process / rRNA processing / protein tag activity / ribosome biogenesis / ribosome binding / ribosomal small subunit biogenesis / ribosomal small subunit assembly / small ribosomal subunit / small ribosomal subunit rRNA binding / 5S rRNA binding / large ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / cytosolic large ribosomal subunit / cytoplasmic translation / rRNA binding / negative regulation of translation / ribosome / intracellular signal transduction / protein ubiquitination / structural constituent of ribosome / translation / ribonucleoprotein complex / mRNA binding / ubiquitin protein ligase binding / nucleolus / RNA binding / zinc ion binding / ATP binding / nucleus / metal ion binding / cytoplasm / cytosol Similarity search - Function | |||||||||||||||
| Biological species |  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.38 Å | |||||||||||||||
 Authors Authors | Cottilli P / Itoh Y / Amunts A | |||||||||||||||
| Funding support | European Union, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Plant Commun / Year: 2022 Journal: Plant Commun / Year: 2022Title: Cryo-EM structure and rRNA modification sites of a plant ribosome. Authors: Patrick Cottilli / Yuzuru Itoh / Yuko Nobe / Anton S Petrov / Purificación Lisón / Masato Taoka / Alexey Amunts /     Abstract: Protein synthesis in crop plants contributes to the balance of food and fuel on our planet, which influences human metabolic activity and lifespan. Protein synthesis can be regulated with respect to ...Protein synthesis in crop plants contributes to the balance of food and fuel on our planet, which influences human metabolic activity and lifespan. Protein synthesis can be regulated with respect to changing environmental cues via the deposition of chemical modifications into rRNA. Here, we present the structure of a plant ribosome from tomato and a quantitative mass spectrometry analysis of its rRNAs. The study reveals fine features of the ribosomal proteins and 71 plant-specific rRNA modifications, and it re-annotates 30 rRNA residues in the available sequence. At the protein level, isoAsp is found in position 137 of uS11, and a zinc finger previously believed to be universal is missing from eL34, suggesting a lower effect of zinc deficiency on protein synthesis in plants. At the rRNA level, the plant ribosome differs markedly from its human counterpart with respect to the spatial distribution of modifications. Thus, it represents an additional layer of gene expression regulation, highlighting the molecular signature of a plant ribosome. The results provide a reference model of a plant ribosome for structural studies and an accurate marker for molecular ecology. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14004.map.gz emd_14004.map.gz | 75 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14004-v30.xml emd-14004-v30.xml emd-14004.xml emd-14004.xml | 97.9 KB 97.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_14004.png emd_14004.png | 52.6 KB | ||
| Filedesc metadata |  emd-14004.cif.gz emd-14004.cif.gz | 20 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14004 http://ftp.pdbj.org/pub/emdb/structures/EMD-14004 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14004 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14004 | HTTPS FTP |
-Validation report
| Summary document |  emd_14004_validation.pdf.gz emd_14004_validation.pdf.gz | 465.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_14004_full_validation.pdf.gz emd_14004_full_validation.pdf.gz | 464.7 KB | Display | |
| Data in XML |  emd_14004_validation.xml.gz emd_14004_validation.xml.gz | 8.1 KB | Display | |
| Data in CIF |  emd_14004_validation.cif.gz emd_14004_validation.cif.gz | 9.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14004 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14004 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14004 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14004 | HTTPS FTP |
-Related structure data
| Related structure data |  7qizMC  7qiwC  7qixC  7qiyC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14004.map.gz / Format: CCP4 / Size: 600.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14004.map.gz / Format: CCP4 / Size: 600.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 60S, 40S body and 40S head combined map. | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
+Entire : 80S ribosome
+Supramolecule #1: 80S ribosome
+Macromolecule #1: 60S ribosomal protein L8
+Macromolecule #2: Ribos_L4_asso_C domain-containing protein
+Macromolecule #3: Ribosomal protein L3
+Macromolecule #4: Ribosomal_L18_c domain-containing protein
+Macromolecule #5: Ribosomal_L6e_N domain-containing protein
+Macromolecule #6: Thaliana 60S ribosomal protein L7
+Macromolecule #7: Ribosomal_L7Ae domain-containing protein
+Macromolecule #8: 60S ribosomal protein uL6
+Macromolecule #9: 60S ribosomal protein L10
+Macromolecule #10: 60S ribosomal protein uL5
+Macromolecule #11: 60S ribosomal protein L13
+Macromolecule #12: Ribosomal_L14e domain-containing protein
+Macromolecule #13: Ribosomal protein L15
+Macromolecule #14: Pectinesterase
+Macromolecule #15: 50S ribosomal protein L22, chloroplastic
+Macromolecule #16: Ribosomal_L18e/L15P domain-containing protein
+Macromolecule #17: Ribosomal protein L19
+Macromolecule #18: 60S ribosomal protein L18a
+Macromolecule #19: 60S ribosomal protein eL21
+Macromolecule #20: 60S ribosomal protein eL22
+Macromolecule #21: 60S ribosomal protein uL14
+Macromolecule #22: TRASH domain-containing protein
+Macromolecule #23: Ribosomal_L23eN domain-containing protein
+Macromolecule #24: KOW domain-containing protein
+Macromolecule #25: 60S ribosomal protein L27
+Macromolecule #26: Ribosomal_L18e/L15P domain-containing protein
+Macromolecule #27: 60S ribosomal protein L29
+Macromolecule #28: 60S ribosomal protein eL30
+Macromolecule #29: 60S ribosomal protein eL31
+Macromolecule #30: 60S ribosomal protein eL32
+Macromolecule #31: 60S ribosomal protein eL33
+Macromolecule #32: 60S ribosomal protein eL34
+Macromolecule #33: Similar to 60S ribosomal protein L35
+Macromolecule #34: 60S ribosomal protein L36
+Macromolecule #35: Ribosomal protein L37
+Macromolecule #36: 60S ribosomal protein L38
+Macromolecule #37: 60S ribosomal protein eL39
+Macromolecule #38: Ubiquitin
+Macromolecule #39: 60S ribosomal protein eL42
+Macromolecule #40: 60S ribosomal protein eL43
+Macromolecule #41: Ribosomal_L28e domain-containing protein
+Macromolecule #47: KH type-2 domain-containing protein
+Macromolecule #48: Ribosomal_S7 domain-containing protein
+Macromolecule #49: S10_plectin domain-containing protein
+Macromolecule #50: 40S ribosomal protein uS19
+Macromolecule #51: 40S ribosomal protein uS9
+Macromolecule #52: 40S ribosomal protein uS13
+Macromolecule #53: 40S ribosomal protein eS19
+Macromolecule #54: Ribosomal_S10 domain-containing protein
+Macromolecule #55: 40S ribosomal protein eS28
+Macromolecule #56: 40S ribosomal protein uS14
+Macromolecule #57: Mitogen-activated protein kinase
+Macromolecule #58: 40S ribosomal protein S25
+Macromolecule #61: 60S ribosomal protein L41
+Macromolecule #62: 40S ribosomal protein SA
+Macromolecule #63: 40S ribosomal protein S3a
+Macromolecule #64: 40S ribosomal protein S4
+Macromolecule #65: 40S ribosomal protein S7
+Macromolecule #66: 40S ribosomal protein S8
+Macromolecule #67: Ribosomal_S17_N domain-containing protein
+Macromolecule #68: 40S ribosomal protein S17
+Macromolecule #69: 40S ribosomal protein S21
+Macromolecule #70: 40S body ribosomal protein uS12
+Macromolecule #71: 40S ribosomal protein S26
+Macromolecule #72: S5 DRBM domain-containing protein
+Macromolecule #73: 40S ribosomal protein S6
+Macromolecule #74: 40S body ribosomal protein uS4
+Macromolecule #75: 30S ribosomal protein S15, chloroplastic
+Macromolecule #76: Ribosomal protein S14
+Macromolecule #77: 40S ribosomal protein S24
+Macromolecule #78: 40S ribosomal protein S27
+Macromolecule #79: 40S ribosomal protein S30
+Macromolecule #80: 40S ribosomal protein S15a-1
+Macromolecule #42: tRNA
+Macromolecule #43: 25S rRNA
+Macromolecule #44: 5S rRNA
+Macromolecule #45: 5.8S rRNA
+Macromolecule #46: 18S
+Macromolecule #59: tRNA_1
+Macromolecule #60: mRNA
+Macromolecule #81: POTASSIUM ION
+Macromolecule #82: MAGNESIUM ION
+Macromolecule #83: ZINC ION
+Macromolecule #84: SPERMINE
+Macromolecule #85: SPERMIDINE
+Macromolecule #86: 1,4-DIAMINOBUTANE
+Macromolecule #87: beta-D-glucopyranose
+Macromolecule #88: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 30.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.8000000000000003 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP EMDB ID: |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 2.38 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 335806 |
| Initial angle assignment | Type: PROJECTION MATCHING / Software - Name: RELION (ver. 3.0) |
| Final angle assignment | Type: ANGULAR RECONSTITUTION / Software - Name: RELION (ver. 3.1) |
 Movie
Movie Controller
Controller