+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13643 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Delta-latroinsectotoxin dimer | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Pore-forming toxin / TOXIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationother organism cell membrane / host cell presynaptic membrane / exocytosis / toxin activity / extracellular region Similarity search - Function | ||||||||||||
| Biological species |  Latrodectus tredecimguttatus (black widow) Latrodectus tredecimguttatus (black widow) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.63 Å | ||||||||||||
 Authors Authors | Chen M / Gatsogiannis C | ||||||||||||
| Funding support |  Japan, 3 items Japan, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Molecular architecture of black widow spider neurotoxins. Authors: Minghao Chen / Daniel Blum / Lena Engelhard / Stefan Raunser / Richard Wagner / Christos Gatsogiannis /  Abstract: Latrotoxins (LaTXs) are presynaptic pore-forming neurotoxins found in the venom of Latrodectus spiders. The venom contains a toxic cocktail of seven LaTXs, with one of them targeting vertebrates (α- ...Latrotoxins (LaTXs) are presynaptic pore-forming neurotoxins found in the venom of Latrodectus spiders. The venom contains a toxic cocktail of seven LaTXs, with one of them targeting vertebrates (α-latrotoxin (α-LTX)), five specialized on insects (α, β, γ, δ, ε- latroinsectotoxins (LITs), and one on crustaceans (α-latrocrustatoxin (α-LCT)). LaTXs bind to specific receptors on the surface of neuronal cells, inducing the release of neurotransmitters either by directly stimulating exocytosis or by forming Ca-conductive tetrameric pores in the membrane. Despite extensive studies in the past decades, a high-resolution structure of a LaTX is not yet available and the precise mechanism of LaTX action remains unclear. Here, we report cryoEM structures of the α-LCT monomer and the δ-LIT dimer. The structures reveal that LaTXs are organized in four domains. A C-terminal domain of ankyrin-like repeats shields a central membrane insertion domain of six parallel α-helices. Both domains are flexibly linked via an N-terminal α-helical domain and a small β-sheet domain. A comparison between the structures suggests that oligomerization involves major conformational changes in LaTXs with longer C-terminal domains. Based on our data we propose a cyclic mechanism of oligomerization, taking place prior membrane insertion. Both recombinant α-LCT and δ-LIT form channels in artificial membrane bilayers, that are stabilized by Ca ions and allow calcium flux at negative membrane potentials. Our comparative analysis between α-LCT and δ-LIT provides first crucial insights towards understanding the molecular mechanism of the LaTX family. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13643.map.gz emd_13643.map.gz | 9.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13643-v30.xml emd-13643-v30.xml emd-13643.xml emd-13643.xml | 12.8 KB 12.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_13643.png emd_13643.png | 66.8 KB | ||
| Filedesc metadata |  emd-13643.cif.gz emd-13643.cif.gz | 6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13643 http://ftp.pdbj.org/pub/emdb/structures/EMD-13643 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13643 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13643 | HTTPS FTP |
-Related structure data
| Related structure data |  7ptyMC  7ptxC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13643.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13643.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.9 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
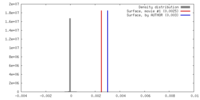
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Delta-latroinsectotoxin-Lt1a in soluble dimeric state
| Entire | Name: Delta-latroinsectotoxin-Lt1a in soluble dimeric state |
|---|---|
| Components |
|
-Supramolecule #1: Delta-latroinsectotoxin-Lt1a in soluble dimeric state
| Supramolecule | Name: Delta-latroinsectotoxin-Lt1a in soluble dimeric state / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Latrodectus tredecimguttatus (black widow) Latrodectus tredecimguttatus (black widow) |
| Molecular weight | Theoretical: 139.8 KDa |
-Macromolecule #1: Delta-latroinsectotoxin-Lt1a
| Macromolecule | Name: Delta-latroinsectotoxin-Lt1a / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Latrodectus tredecimguttatus (black widow) Latrodectus tredecimguttatus (black widow) |
| Molecular weight | Theoretical: 145.047547 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MKFLVNVALV FMVVYISYIY AMWSHPQFEK GSAGSAAGSG AGWSHPQFEK GAGLEVLFQG PPWMHSKELQ TISAAVARKA VPNTMVIRL KRDEEDGEMT LEERQAQCKA IEYSNSVFGM IADVANDIGS IPVIGEVVGI VTAPIAIVSH ITSAGLDIAS T ALDCDDIP ...String: MKFLVNVALV FMVVYISYIY AMWSHPQFEK GSAGSAAGSG AGWSHPQFEK GAGLEVLFQG PPWMHSKELQ TISAAVARKA VPNTMVIRL KRDEEDGEMT LEERQAQCKA IEYSNSVFGM IADVANDIGS IPVIGEVVGI VTAPIAIVSH ITSAGLDIAS T ALDCDDIP FDEIKEILEE RFNEIDRKLD KNTAALEEVS KLVSKTFVTV EKTRNEMNEN FKLVLETIES KEIKSIVFKI ND FKKFFEK ERQRIKGLPK DRYVAKLLEQ KGILGSLKEV REPSGNSLSS ALNELLDKNN NYAIPKVVDD NKAFQALYAL FYG TQTYAA VMFFLLEQHS YLADYYYQKG DDVNFNAEFN NVAIIFDDFK SSLTGGDDGL IDNVIEVLNT VKALPFIKNA DSKL YRELV TRTKALETLK NQIKTTDLPL IDDIPETLSQ VNFPNDENQL PTPIGNWVDG VEVRYAVQYE SKGMYSKFSE WSEPF TVQG NACPTIKVRV DPKKRNRLIF RKFNSGKPQF AGTMTHSQTN FKDIHRDLYD AALNINKLKA VDEATTLIEK GADIEA KFD NDRSAMHAVA YRGNNKIALR FLLKNQSIDI ELKDKNGFTP LHIAAEAGQA GFVKLLINHG ADVNAKTSKT NLTPLHL AT RSGFSKTVRN LLESPNIKVN EKEDDGFTPL HTAVMSTYMV VDALLNHPDI DKNAQSTSGL TPFHLAIINE SQEVAESL V ESNADLNIQD VNHMAPIHFA ASMGSIKMLR YLISIKDKVS INSVTENNNW TPLHFAIYFK KEDAAKELLK QDDINLTIV ADGNLTVLHL AVSTGQINII KELLKRGSNI EEKTGEGYTS LHIAAMRKEP EIAVVLIENG ADIEARSADN LTPLHSAAKI GRKSTVLYL LEKGADIGAK TADGSTALHL AVSGRKMKTV ETLLNKGANL KEYDNNKYLP IHKAIINDDL DMVRLFLEKD P SLKDDETE EGRTSIMLIV QKLLLELYNY FINNYAETLD EEALFNRLDE QGKLELAYIF HNKEGDAKEA VKPTILVTIK LM EYCLKKL REESGAPEGS FDSPSSKQCI STFSEDEMFR RTLPEIVKET NSRYLPLKGF SRSLNKFLPS LKFAESKNSY RSE NFVSNI DSNGALLLLD VFIRKFTNEK YNLTGKEAVP YLEAKASSLR IASKFEELLT EVKGIPAGEL INMAEVSSNI HKAI ASGKP VSKVLCSYLD TFSELNSQQM EELVNTYLST KPSVITSASA DYQKLPNLLT ATCLEPERMA QLIDVHQKMF LRLES SGLV PRGSHHHHHH HH UniProtKB: Delta-latroinsectotoxin-Lt1a |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 78.7 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-7pty: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)