+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12979 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex | ||||||||||||||||||
 Map data Map data | Structure of the TELO2-TTI1-TTI2 complex and its function in TOR recruitment to the R2TP chaperone | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | R2TP / TTT / TELO2 / TTI1 / TTI2 / RUVBL1 / RUVBL2 / HSP90 chaperone / mTOR / PIKK / STRUCTURAL PROTEIN / CHAPERONE | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationprotein-containing complex stabilizing activity / positive regulation of DNA damage checkpoint / 'de novo' cotranslational protein folding / TTT Hsp90 cochaperone complex / promoter-enhancer loop anchoring activity / telomerase RNA localization to Cajal body / TORC2 complex / regulation of DNA strand elongation / positive regulation of telomere maintenance in response to DNA damage / establishment of protein localization to chromatin ...protein-containing complex stabilizing activity / positive regulation of DNA damage checkpoint / 'de novo' cotranslational protein folding / TTT Hsp90 cochaperone complex / promoter-enhancer loop anchoring activity / telomerase RNA localization to Cajal body / TORC2 complex / regulation of DNA strand elongation / positive regulation of telomere maintenance in response to DNA damage / establishment of protein localization to chromatin / TORC1 complex / R2TP complex / dynein axonemal particle / RPAP3/R2TP/prefoldin-like complex / regulation of TOR signaling / Swr1 complex / Ino80 complex / regulation of double-strand break repair / box C/D snoRNP assembly / regulation of chromosome organization / telomeric repeat DNA binding / NuA4 histone acetyltransferase complex / regulation of DNA replication / MLL1 complex / TFIID-class transcription factor complex binding / regulation of embryonic development / Telomere Extension By Telomerase / protein folding chaperone complex / regulation of DNA repair / RNA polymerase II core promoter sequence-specific DNA binding / positive regulation of double-strand break repair via homologous recombination / Deposition of new CENPA-containing nucleosomes at the centromere / TBP-class protein binding / telomere maintenance / positive regulation of DNA repair / DNA helicase activity / cellular response to estradiol stimulus / euchromatin / negative regulation of canonical Wnt signaling pathway / Hsp90 protein binding / Formation of the beta-catenin:TCF transactivating complex / ADP binding / chromatin DNA binding / beta-catenin binding / DNA Damage Recognition in GG-NER / kinase binding / nuclear matrix / cellular response to UV / unfolded protein binding / UCH proteinases / transcription corepressor activity / positive regulation of canonical Wnt signaling pathway / nucleosome / HATs acetylate histones / ATPase binding / protein folding / DNA recombination / regulation of apoptotic process / spermatogenesis / molecular adaptor activity / DNA helicase / transcription coactivator activity / chromosome, telomeric region / regulation of cell cycle / Ub-specific processing proteases / nuclear speck / protein stabilization / nuclear body / ciliary basal body / RNA polymerase II cis-regulatory region sequence-specific DNA binding / cadherin binding / chromatin remodeling / ribonucleoprotein complex / cell division / DNA repair / regulation of DNA-templated transcription / regulation of transcription by RNA polymerase II / centrosome / protein kinase binding / positive regulation of DNA-templated transcription / protein-containing complex binding / protein homodimerization activity / ATP hydrolysis activity / positive regulation of transcription by RNA polymerase II / extracellular exosome / nucleoplasm / ATP binding / membrane / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | ||||||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||||||||
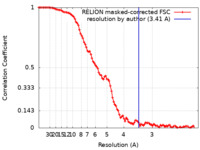
| Method | single particle reconstruction / cryo EM / Resolution: 3.41 Å | ||||||||||||||||||
 Authors Authors | Pal M / Llorca O | ||||||||||||||||||
| Funding support |  United Kingdom, United Kingdom,  Spain, 5 items Spain, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2021 Journal: Cell Rep / Year: 2021Title: Structure of the TELO2-TTI1-TTI2 complex and its function in TOR recruitment to the R2TP chaperone. Authors: Mohinder Pal / Hugo Muñoz-Hernandez / Dennis Bjorklund / Lihong Zhou / Gianluca Degliesposti / J Mark Skehel / Emma L Hesketh / Rebecca F Thompson / Laurence H Pearl / Oscar Llorca / Chrisostomos Prodromou /   Abstract: The R2TP (RUVBL1-RUVBL2-RPAP3-PIH1D1) complex, in collaboration with heat shock protein 90 (HSP90), functions as a chaperone for the assembly and stability of protein complexes, including RNA ...The R2TP (RUVBL1-RUVBL2-RPAP3-PIH1D1) complex, in collaboration with heat shock protein 90 (HSP90), functions as a chaperone for the assembly and stability of protein complexes, including RNA polymerases, small nuclear ribonucleoprotein particles (snRNPs), and phosphatidylinositol 3-kinase (PI3K)-like kinases (PIKKs) such as TOR and SMG1. PIKK stabilization depends on an additional complex of TELO2, TTI1, and TTI2 (TTT), whose structure and function are poorly understood. The cryoelectron microscopy (cryo-EM) structure of the human R2TP-TTT complex, together with biochemical experiments, reveals the mechanism of TOR recruitment to the R2TP-TTT chaperone. The HEAT-repeat TTT complex binds the kinase domain of TOR, without blocking its activity, and delivers TOR to the R2TP chaperone. In addition, TTT regulates the R2TP chaperone by inhibiting RUVBL1-RUVBL2 ATPase activity and by modulating the conformation and interactions of the PIH1D1 and RPAP3 components of R2TP. Taken together, our results show how TTT couples the recruitment of TOR to R2TP with the regulation of this chaperone system. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12979.map.gz emd_12979.map.gz | 78.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12979-v30.xml emd-12979-v30.xml emd-12979.xml emd-12979.xml | 51.9 KB 51.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12979_fsc.xml emd_12979_fsc.xml | 10 KB | Display |  FSC data file FSC data file |
| Images |  emd_12979.png emd_12979.png | 96.3 KB | ||
| Filedesc metadata |  emd-12979.cif.gz emd-12979.cif.gz | 10.1 KB | ||
| Others |  emd_12979_additional_1.map.gz emd_12979_additional_1.map.gz emd_12979_additional_2.map.gz emd_12979_additional_2.map.gz emd_12979_additional_3.map.gz emd_12979_additional_3.map.gz emd_12979_additional_4.map.gz emd_12979_additional_4.map.gz emd_12979_additional_5.map.gz emd_12979_additional_5.map.gz emd_12979_additional_6.map.gz emd_12979_additional_6.map.gz emd_12979_half_map_1.map.gz emd_12979_half_map_1.map.gz emd_12979_half_map_2.map.gz emd_12979_half_map_2.map.gz | 74.7 MB 74.4 MB 65.3 MB 65.4 MB 65.4 MB 50.4 MB 65.4 MB 65.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12979 http://ftp.pdbj.org/pub/emdb/structures/EMD-12979 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12979 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12979 | HTTPS FTP |
-Related structure data
| Related structure data |  7oleMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12979.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12979.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of the TELO2-TTI1-TTI2 complex and its function in TOR recruitment to the R2TP chaperone | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Cryo-EM structure of the TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex (dataset1)...
| File | emd_12979_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of the TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex (dataset1) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Cryo-EM structure of the TELO2-TTI1-TTI2 complex
| File | emd_12979_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of the TELO2-TTI1-TTI2 complex | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Cryo-EM structure of the TELO2-TTI1-TTI2 complex
| File | emd_12979_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of the TELO2-TTI1-TTI2 complex | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Cryo-EM structure of the TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex (dataset1 halfmap1)...
| File | emd_12979_additional_4.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of the TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex (dataset1 halfmap1) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Cryo-EM structure of the TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex (dataset1 halfmap2)...
| File | emd_12979_additional_5.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of the TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex (dataset1 halfmap2) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Composite map of the R2-TTT structure
| File | emd_12979_additional_6.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Composite map of the R2-TTT structure | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Cryo-EM structure of the TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex (dataset2)...
| File | emd_12979_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of the TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex (dataset2) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Cryo-EM structure of the TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex (dataset2)...
| File | emd_12979_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of the TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex (dataset2) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex
| Entire | Name: TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex |
|---|---|
| Components |
|
-Supramolecule #1: TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex
| Supramolecule | Name: TELO2-TTI1-TTI2-RUVBL1-RUVBL2 complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Molecular weight | Theoretical: 580 KDa |
-Macromolecule #1: RuvB-like 1
| Macromolecule | Name: RuvB-like 1 / type: protein_or_peptide / ID: 1 / Details: ADP / Number of copies: 3 / Enantiomer: LEVO / EC number: DNA helicase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 50.296914 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKIEEVKSTT KTQRIASHSH VKGLGLDESG LAKQAASGLV GQENAREACG VIVELIKSKK MAGRAVLLAG PPGTGKTALA LAIAQELGS KVPFCPMVGS EVYSTEIKKT EVLMENFRRA IGLRIKETKE VYEGEVTELT PCETENPMGG YGKTISHVII G LKTAKGTK ...String: MKIEEVKSTT KTQRIASHSH VKGLGLDESG LAKQAASGLV GQENAREACG VIVELIKSKK MAGRAVLLAG PPGTGKTALA LAIAQELGS KVPFCPMVGS EVYSTEIKKT EVLMENFRRA IGLRIKETKE VYEGEVTELT PCETENPMGG YGKTISHVII G LKTAKGTK QLKLDPSIFE SLQKERVEAG DVIYIEANSG AVKRQGRCDT YATEFDLEAE EYVPLPKGDV HKKKEIIQDV TL HDLDVAN ARPQGGQDIL SMMGQLMKPK KTEITDKLRG EINKVVNKYI DQGIAELVPG VLFVDEVHML DIECFTYLHR ALE SSIAPI VIFASNRGNC VIRGTEDITS PHGIPLDLLD RVMIIRTMLY TPQEMKQIIK IRAQTEGINI SEEALNHLGE IGTK TTLRY SVQLLTPANL LAKINGKDSI EKEHVEEISE LFYDAKSSAK ILADQQDKYM K UniProtKB: RuvB-like 1 |
-Macromolecule #2: RuvB-like 2
| Macromolecule | Name: RuvB-like 2 / type: protein_or_peptide / ID: 2 / Details: ADP / Number of copies: 3 / Enantiomer: LEVO / EC number: DNA helicase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 51.222465 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MATVTATTKV PEIRDVTRIE RIGAHSHIRG LGLDDALEPR QASQGMVGQL AARRAAGVVL EMIREGKIAG RAVLIAGQPG TGKTAIAMG MAQALGPDTP FTAIAGSEIF SLEMSKTEAL TQAFRRSIGV RIKEETEIIE GEVVEIQIDR PATGTGSKVG K LTLKTTEM ...String: MATVTATTKV PEIRDVTRIE RIGAHSHIRG LGLDDALEPR QASQGMVGQL AARRAAGVVL EMIREGKIAG RAVLIAGQPG TGKTAIAMG MAQALGPDTP FTAIAGSEIF SLEMSKTEAL TQAFRRSIGV RIKEETEIIE GEVVEIQIDR PATGTGSKVG K LTLKTTEM ETIYDLGTKM IESLTKDKVQ AGDVITIDKA TGKISKLGRS FTRARDYDAM GSQTKFVQCP DGELQKRKEV VH TVSLHEI DVINSRTQGF LALFSGDTGE IKSEVREQIN AKVAEWREEG KAEIIPGVLF IDEVHMLDIE SFSFLNRALE SDM APVLIM ATNRGITRIR GTSYQSPHGI PIDLLDRLLI VSTTPYSEKD TKQILRIRCE EEDVEMSEDA YTVLTRIGLE TSLR YAIQL ITAASLVCRK RKGTEVQVDD IKRVYSLFLD ESRSTQYMKE YQDAFLFNEL KGETMDTS UniProtKB: RuvB-like 2 |
-Macromolecule #3: TELO2-interacting protein 1 homolog,TELO2-interacting protein 1 h...
| Macromolecule | Name: TELO2-interacting protein 1 homolog,TELO2-interacting protein 1 homolog,TTI1 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 176.844344 KDa |
| Recombinant expression | Organism:  Spodoptera aff. frugiperda 1 BOLD-2017 (butterflies/moths) Spodoptera aff. frugiperda 1 BOLD-2017 (butterflies/moths) |
| Sequence | String: MAVFDTPEEA FGVLRPVCVQ LTKTQTVENV EHLQTRLQAV SDSALQELQQ YILFPLRFTL KTPGPKRERL IQSVVECLTF VLSSTCVKE QELLQELFSE LSACLYSPSS QKPAAVSEEL KLAVIQGLST LMHSAYGDII LTFYEPSILP RLGFAVSLLL G LAEQEKSK ...String: MAVFDTPEEA FGVLRPVCVQ LTKTQTVENV EHLQTRLQAV SDSALQELQQ YILFPLRFTL KTPGPKRERL IQSVVECLTF VLSSTCVKE QELLQELFSE LSACLYSPSS QKPAAVSEEL KLAVIQGLST LMHSAYGDII LTFYEPSILP RLGFAVSLLL G LAEQEKSK QIKIAALKCL QVLLLQCDCQ DHPRSLDELE QKQLGDLFAS FLPGISTALT RLITGDFKQG HSIVVSSLKI FY KTVSFIM ADEQLKRISK VQAKPAVEHR VAELMVYREA DWVKKTGDKL TILIKKIIEC VSVHPHWKVR LELVELVEDL LLK CSQSLV ECAGPLLKAL VGLVNDESPE IQAQCNKVLR HFADQKVVVG NKALADILSE SLHSLATSLP RLMNSQDDQG KFST LSLLL GYLKLLGPKI NFVLNSVAHL QRLSKALIQV LELDVADIKI VEERRWNSDD LNASPKTSAT QPWNRIQRRY FRFFT DERI FMLLRQVCQL LGYYGNLYLL VDHFMELYHQ SVVYRKQAAM ILNELVTGAA GLEVEDLHEK HIKTNPEELR EIVTSI LEE YTSQENWYLV TCLETEEMGE ELMMEHPGLQ AITSGEHTCQ VTSFLAFSKP SPTICSMNSN IWQICIQLEG IGQFAYA LG KDFCLLLMSA LYPVLEKAGD QTLLISQVAT STMMDVCRAC GYDSLQHLIN QNSDYLVNGI SLNLRHLALH PHTPKVLE V MLRNSDANLL PLVADVVQDV LATLDQFYDK RAASFVSVLH ALMAALAQWF PDTGNLGHLQ EQSLGEEGSH LNQRPAALE KSTTTAEDIE QFLLNYLKEK DVADGNVSDF DNEEEEQSVP PKVDENDTRP DVEPPLPLQI QIAMDVMERC IHLLSDKNLQ IRLKVLDVL DLCVVVLQSH KNQLLPLAHQ AWPSLVHRLT RDAPLAVLRA FKVLRTLGSK CGDFLRSRFC KDVLPKLAGS L VTQAPISA RAGPVYSHTL AFKLQLAVLQ GLGPLCERLD LGEGDLNKVA DACLIYLSVK QPVKLQEAAR SVFLHLMKVD PD STWFLLN ELYCPVQFTP PHPSLHPVQL HGASGQQNPY TTNVLQLLKE LQ(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)G(UNK)G(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)G(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)G (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)G(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)G(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) UniProtKB: TELO2-interacting protein 1 homolog |
-Macromolecule #4: TELO2-interacting protein 2,TELO2-interacting protein 2,TTI2
| Macromolecule | Name: TELO2-interacting protein 2,TELO2-interacting protein 2,TTI2 type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 95.633781 KDa |
| Recombinant expression | Organism:  Spodoptera aff. frugiperda 1 BOLD-2017 (butterflies/moths) Spodoptera aff. frugiperda 1 BOLD-2017 (butterflies/moths) |
| Sequence | String: MELDSALEAP SQEDSNLSEE LSHSAFGQAF SKILHCLARP EARRGNVKDA VLKDLGDLIE ATEFDRLFEG TGARLRGMPE TLGQVAKAL EKYAAPSKEE EGGGDGHSEA AEKAAQVGLL FLKLLGKVET AKNSLVGPAW QTGLHHLAGP VYIFAITHSL E QPWTTPRS ...String: MELDSALEAP SQEDSNLSEE LSHSAFGQAF SKILHCLARP EARRGNVKDA VLKDLGDLIE ATEFDRLFEG TGARLRGMPE TLGQVAKAL EKYAAPSKEE EGGGDGHSEA AEKAAQVGLL FLKLLGKVET AKNSLVGPAW QTGLHHLAGP VYIFAITHSL E QPWTTPRS REVAREVLTS LLQVTECGSV AGFLHGENED EKGRLSVILG LLKPDLYKES WKNNPAIKHV FSWTLQQVTR PW LSQHLER VLPASLVISD DYQTENKILG VHCLHHIVLN VPAADLLQYN RAQVLYHAIS NHLYTPEHHL IQAVLLCLLD LFP ILEKTL HWKGDGARPT THCDEVLRLI LTHMEPEHRL LLRRTYARNL PAFVNRLGIL TVRHLKRLER VIIGYLEVYD GPEE EARLK ILETLKLLMQ HTWPRVSCRL VVLLKALLKL ICDVARDPNL TPESVKSALL QEATDCLILL DRCSQGRVKG LLAKI PQSC EDRKVVNYIR KVQQVSEGAP YNGT(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)G(UNK)G(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)G(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)G (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)G (UNK)(UNK)G (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) UniProtKB: TELO2-interacting protein 2 |
-Macromolecule #5: Telomere length regulation protein TEL2 homolog
| Macromolecule | Name: Telomere length regulation protein TEL2 homolog / type: protein_or_peptide / ID: 5 / Details: LGEMEPPALPREKEEFASAHF / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 53.424195 KDa |
| Recombinant expression | Organism:  Spodoptera aff. frugiperda 1 BOLD-2017 (butterflies/moths) Spodoptera aff. frugiperda 1 BOLD-2017 (butterflies/moths) |
| Sequence | String: MEPAPSEVRL AVREAIHALS SSEDGGHIFC TLESLKRYLG EMEPPALPRE (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)KEEFASAHF SPVLRCLASR LSPAWLELLP HGRLE(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) ...String: MEPAPSEVRL AVREAIHALS SSEDGGHIFC TLESLKRYLG EMEPPALPRE (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)KEEFASAHF SPVLRCLASR LSPAWLELLP HGRLE(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)ELWASF FLEGPADQAF LVLMETIEGA AGPSFRLMKM ARLLARFLRE GRLAVLMEAQ CRQQTQPGF ILLRETLLGK VV(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)ALPDHLGNR LQQENLAEFF PQNYFRLLGE EVVRVLQAVV DSLQGGLDSS VSFVSQ VLG KACVHGRQQE ILGVLVPRLA ALTQGSYLHQ RVCWRLVEQV PDRAMEAVLT GLVEAALGPE VLSRLLGNLV VKNKKAQ FV MTQKLLFLQS RLTTPMLQSL LGHLAMDSQR RPLLLQVLKE LLETWGSSSA IRHTPLPQQR HVSKAVLICL AQLGEPEL R DSRDELLASM MAGVKCRLDS SLPPVRRLGM IVAEVVSARI HPEGPPLKFQ YEEDELSLEL LALASPQPAG DGASEAGT UniProtKB: Telomere length regulation protein TEL2 homolog, Telomere length regulation protein TEL2 homolog, Telomere length regulation protein TEL2 homolog, Telomere length regulation protein TEL2 homolog |
-Macromolecule #6: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 6 / Number of copies: 5 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: Solutions were made fresh for protein purification | |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | |||||||||
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 90 % / Chamber temperature: 287.15 K / Instrument: LEICA PLUNGER / Details: 3sec. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 90.0 K / Max: 100.0 K |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 1034 pixel / Digitization - Dimensions - Height: 1034 pixel / Digitization - Frames/image: 1-40 / Number grids imaged: 10 / Number real images: 6000 / Average exposure time: 9.0 sec. / Average electron dose: 60.17 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 3.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus min: 1.2 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)