+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12587 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Trimeric SARS-CoV-2 spike ectodomain bound to P008_056 Fab | |||||||||
 Map data Map data | main map | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationMaturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / entry receptor-mediated virion attachment to host cell ...Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / entry receptor-mediated virion attachment to host cell / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / receptor-mediated endocytosis of virus by host cell / membrane fusion / Attachment and Entry / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / receptor ligand activity / host cell surface receptor binding / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Rosa A / Pye VE / Nans A / Cherepanov P | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2021 Journal: Sci Adv / Year: 2021Title: SARS-CoV-2 can recruit a heme metabolite to evade antibody immunity. Authors: Annachiara Rosa / Valerie E Pye / Carl Graham / Luke Muir / Jeffrey Seow / Kevin W Ng / Nicola J Cook / Chloe Rees-Spear / Eleanor Parker / Mariana Silva Dos Santos / Carolina Rosadas / ...Authors: Annachiara Rosa / Valerie E Pye / Carl Graham / Luke Muir / Jeffrey Seow / Kevin W Ng / Nicola J Cook / Chloe Rees-Spear / Eleanor Parker / Mariana Silva Dos Santos / Carolina Rosadas / Alberto Susana / Hefin Rhys / Andrea Nans / Laura Masino / Chloe Roustan / Evangelos Christodoulou / Rachel Ulferts / Antoni G Wrobel / Charlotte-Eve Short / Michael Fertleman / Rogier W Sanders / Judith Heaney / Moira Spyer / Svend Kjær / Andy Riddell / Michael H Malim / Rupert Beale / James I MacRae / Graham P Taylor / Eleni Nastouli / Marit J van Gils / Peter B Rosenthal / Massimo Pizzato / Myra O McClure / Richard S Tedder / George Kassiotis / Laura E McCoy / Katie J Doores / Peter Cherepanov /     Abstract: The coronaviral spike is the dominant viral antigen and the target of neutralizing antibodies. We show that SARS-CoV-2 spike binds biliverdin and bilirubin, the tetrapyrrole products of heme ...The coronaviral spike is the dominant viral antigen and the target of neutralizing antibodies. We show that SARS-CoV-2 spike binds biliverdin and bilirubin, the tetrapyrrole products of heme metabolism, with nanomolar affinity. Using cryo-electron microscopy and x-ray crystallography, we mapped the tetrapyrrole interaction pocket to a deep cleft on the spike N-terminal domain (NTD). At physiological concentrations, biliverdin significantly dampened the reactivity of SARS-CoV-2 spike with immune sera and inhibited a subset of neutralizing antibodies. Access to the tetrapyrrole-sensitive epitope is gated by a flexible loop on the distal face of the NTD. Accompanied by profound conformational changes in the NTD, antibody binding requires relocation of the gating loop, which folds into the cleft vacated by the metabolite. Our results indicate that SARS-CoV-2 spike NTD harbors a dominant epitope, access to which can be controlled by an allosteric mechanism that is regulated through recruitment of a metabolite. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12587.map.gz emd_12587.map.gz | 168.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12587-v30.xml emd-12587-v30.xml emd-12587.xml emd-12587.xml | 25 KB 25 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12587_fsc.xml emd_12587_fsc.xml | 12.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_12587.png emd_12587.png | 167.5 KB | ||
| Masks |  emd_12587_msk_1.map emd_12587_msk_1.map | 178 MB |  Mask map Mask map | |
| Others |  emd_12587_additional_1.map.gz emd_12587_additional_1.map.gz emd_12587_half_map_1.map.gz emd_12587_half_map_1.map.gz emd_12587_half_map_2.map.gz emd_12587_half_map_2.map.gz | 160.1 MB 164.9 MB 164.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12587 http://ftp.pdbj.org/pub/emdb/structures/EMD-12587 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12587 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12587 | HTTPS FTP |
-Validation report
| Summary document |  emd_12587_validation.pdf.gz emd_12587_validation.pdf.gz | 484.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_12587_full_validation.pdf.gz emd_12587_full_validation.pdf.gz | 483.9 KB | Display | |
| Data in XML |  emd_12587_validation.xml.gz emd_12587_validation.xml.gz | 20.1 KB | Display | |
| Data in CIF |  emd_12587_validation.cif.gz emd_12587_validation.cif.gz | 25.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12587 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12587 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12587 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-12587 | HTTPS FTP |
-Related structure data
| Related structure data |  7ntcMC  7b62C  7nt9C  7ntaC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12587.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12587.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | main map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.38 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12587_msk_1.map emd_12587_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: deep EM enhancer, tight target
| File | emd_12587_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | deep EM enhancer, tight target | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map A
| File | emd_12587_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B
| File | emd_12587_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Stabilised SARS-CoV-2 spike ectodomain trimer bound to one P008_0...
| Entire | Name: Stabilised SARS-CoV-2 spike ectodomain trimer bound to one P008_056 Fab |
|---|---|
| Components |
|
-Supramolecule #1: Stabilised SARS-CoV-2 spike ectodomain trimer bound to one P008_0...
| Supramolecule | Name: Stabilised SARS-CoV-2 spike ectodomain trimer bound to one P008_056 Fab type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 Details: natural source below: Spike is SARS-Cov-2 and Fab is Human - there is only option for one natural source below! |
|---|---|
| Molecular weight | Theoretical: 478.86 KDa |
-Supramolecule #2: Stabilised SARS-CoV-2 spike ectodomain trimer
| Supramolecule | Name: Stabilised SARS-CoV-2 spike ectodomain trimer / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: Expi293 Homo sapiens (human) / Recombinant cell: Expi293 |
-Supramolecule #3: P008_056 Fab
| Supramolecule | Name: P008_056 Fab / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: Expi293 Homo sapiens (human) / Recombinant cell: Expi293 |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 142.269859 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGILPSPGMP ALLSLVSLLS VLLMGCVAET GMFVFLVLLP LVSSQCVNLT TRTQLPPAYT NSFTRGVYYP DKVFRSSVLH STQDLFLPF FSNVTWFHAI HVSGTNGTKR FDNPVLPFND GVYFASTEKS NIIRGWIFGT TLDSKTQSLL IVNNATNVVI K VCEFQFCN ...String: MGILPSPGMP ALLSLVSLLS VLLMGCVAET GMFVFLVLLP LVSSQCVNLT TRTQLPPAYT NSFTRGVYYP DKVFRSSVLH STQDLFLPF FSNVTWFHAI HVSGTNGTKR FDNPVLPFND GVYFASTEKS NIIRGWIFGT TLDSKTQSLL IVNNATNVVI K VCEFQFCN DPFLGVYYHK NNKSWMESEF RVYSSANNCT FEYVSQPFLM DLEGKQGNFK NLREFVFKNI DGYFKIYSKH TP INLVRDL PQGFSALEPL VDLPIGINIT RFQTLLALHR SYLTPGDSSS GWTAGAAAYY VGYLQPRTFL LKYNENGTIT DAV DCALDP LSETKCTLKS FTVEKGIYQT SNFRVQPTES IVRFPNITNL CPFGEVFNAT RFASVYAWNR KRISNCVADY SVLY NSASF STFKCYGVSP TKLNDLCFTN VYADSFVIRG DEVRQIAPGQ TGKIADYNYK LPDDFTGCVI AWNSNNLDSK VGGNY NYLY RLFRKSNLKP FERDISTEIY QAGSTPCNGV EGFNCYFPLQ SYGFQPTNGV GYQPYRVVVL SFELLHAPAT VCGPKK STN LVKNKCVNFN FNGLTGTGVL TESNKKFLPF QQFGRDIADT TDAVRDPQTL EILDITPCSF GGVSVITPGT NTSNQVA VL YQDVNCTEVP VAIHADQLTP TWRVYSTGSN VFQTRAGCLI GAEHVNNSYE CDIPIGAGIC ASYQTQTNSP SRASSVAS Q SIIAYTMSLG AENSVAYSNN SIAIPTNFTI SVTTEILPVS MTKTSVDCTM YICGDSTECS NLLLQYGSFC TQLNRALTG IAVEQDKNTQ EVFAQVKQIY KTPPIKDFGG FNFSQILPDP SKPSKRSFIE DLLFNKVTLA DAGFIKQYGD CLGDIAARDL ICAQKFNGL TVLPPLLTDE MIAQYTSALL AGTITSGWTF GAGAALQIPF AMQMAYRFNG IGVTQNVLYE NQKLIANQFN S AIGKIQDS LSSTASALGK LQDVVNQNAQ ALNTLVKQLS SNFGAISSVL NDILSRLDPP EAEVQIDRLI TGRLQSLQTY VT QQLIRAA EIRASANLAA TKMSECVLGQ SKRVDFCGKG YHLMSFPQSA PHGVVFLHVT YVPAQEKNFT TAPAICHDGK AHF PREGVF VSNGTHWFVT QRNFYEPQII TTDNTFVSGN CDVVIGIVNN TVYDPLQPEL DSFKEELDKY FKNHTSPDVD LGDI SGINA SVVNIQKEID RLNEVAKNLN ESLIDLQELG KYEQSGRENL YFQGGGGSGY IPEAPRDGQA YVRKDGEWVL LSTFL GHHH HHH |
-Macromolecule #2: P008_056 Fab Heavy chain
| Macromolecule | Name: P008_056 Fab Heavy chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 27.2394 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGWSCIILFL VATATGVHSE VQLVESGGGL VKPGGSLRLS CAASGFSFSS YSMNWVRQAP GKGLEWVSSI SSNSNYIYYA DSMKGRFTI SRDNAKNSLY LQMNSLRAED TAVYYCASNR SPYDSSNYYF DYWGQGTRVT ISSASTKGPS VFPLAPSSKS T SGGTAALG ...String: MGWSCIILFL VATATGVHSE VQLVESGGGL VKPGGSLRLS CAASGFSFSS YSMNWVRQAP GKGLEWVSSI SSNSNYIYYA DSMKGRFTI SRDNAKNSLY LQMNSLRAED TAVYYCASNR SPYDSSNYYF DYWGQGTRVT ISSASTKGPS VFPLAPSSKS T SGGTAALG CLVKDYFPEP VTVSWNSGAL TSGVHTFPAV LQSSGLYSLS SVVTVPSSSL GTQTYICNVN HKPSNTKVDK RV EPKSGTK HHHHHH |
-Macromolecule #3: P008_056 Fab Light chain
| Macromolecule | Name: P008_056 Fab Light chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.25318 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGWSCIILFL VATATGVHCA IRMTQSPSSL SASVGDRVTI TCQASQDISN YLNWYQQKPG KAPKLLIYDA SNLETGVPSR FSGSGSGTD FTFTISSLQP EDIATYYCQH HDSLPLTFGG GTKVEIKRTV AAPSVFIFPP SDEQLKSGTA SVVCLLNNFY P REAKVQWK ...String: MGWSCIILFL VATATGVHCA IRMTQSPSSL SASVGDRVTI TCQASQDISN YLNWYQQKPG KAPKLLIYDA SNLETGVPSR FSGSGSGTD FTFTISSLQP EDIATYYCQH HDSLPLTFGG GTKVEIKRTV AAPSVFIFPP SDEQLKSGTA SVVCLLNNFY P REAKVQWK VDNALQSGNS QESVTEQDSK DSTYSLSSTL TLSKADYEKH KVYACEVTHQ GLSSPVTKSF NRGE |
-Macromolecule #5: BILIVERDINE IX ALPHA
| Macromolecule | Name: BILIVERDINE IX ALPHA / type: ligand / ID: 5 / Number of copies: 2 / Formula: BLA |
|---|---|
| Molecular weight | Theoretical: 582.646 Da |
| Chemical component information | 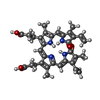 ChemComp-BLA: |
-Macromolecule #6: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 6 / Number of copies: 29 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.6 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 150 mM NaCl, 1 mM ethylenediaminetetraacetic acid (EDTA), 25 mM Tris-HCl, pH 8.0 |
| Grid | Model: Quantifoil R2/2 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE |
| Details | SARS-CoV-2 spike ectodomain (0.6 mg/ml final concentration in TBSE supplemented with 0.1% n-octylglucoside) with 0.2 mg/ml P008_056 Fab |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 17010 / Average electron dose: 51.0 e/Å2 Details: A total of 17010 movies were collected with a pixel size of 1.38 Angstrom and total electron exposure of 51 electrons per square Angstrom, spread over 40 frames. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


















 Z
Z Y
Y X
X



































