[English] 日本語
 Yorodumi
Yorodumi- EMDB-12223: Vps35/Vps29 arch of fungal membrane-assembled retromer:Grd19 complex -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12223 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Vps35/Vps29 arch of fungal membrane-assembled retromer:Grd19 complex | ||||||||||||||||||
 Map data Map data | LAFTER-filtered map of Vps35/Vps29 arch of fungal retromer:Grd19 complex on the membrane | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | endosomes / coat proteins / membrane trafficking / cargo-sorting / ENDOCYTOSIS | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationretromer, cargo-selective complex / retromer complex / retrograde transport, endosome to Golgi / intracellular protein transport / late endosome / protein transport / cytosol Similarity search - Function | ||||||||||||||||||
| Biological species |  Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) / Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) /  Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) | ||||||||||||||||||
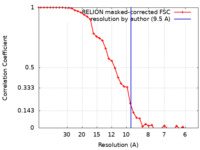
| Method | subtomogram averaging / cryo EM / Resolution: 9.5 Å | ||||||||||||||||||
 Authors Authors | Leneva N / Kovtun O | ||||||||||||||||||
| Funding support |  United Kingdom, 5 items United Kingdom, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2021 Journal: Sci Adv / Year: 2021Title: Architecture and mechanism of metazoan retromer:SNX3 tubular coat assembly. Authors: Natalya Leneva / Oleksiy Kovtun / Dustin R Morado / John A G Briggs / David J Owen /  Abstract: Retromer is a master regulator of cargo retrieval from endosomes, which is critical for many cellular processes including signaling, immunity, neuroprotection, and virus infection. The retromer core ...Retromer is a master regulator of cargo retrieval from endosomes, which is critical for many cellular processes including signaling, immunity, neuroprotection, and virus infection. The retromer core (VPS26/VPS29/VPS35) is present on cargo-transporting, tubular carriers along with a range of sorting nexins. Here, we elucidate the structural basis of membrane tubulation and coupled cargo recognition by metazoan and fungal retromer coats assembled with the non-Bin1/Amphiphysin/Rvs (BAR) sorting nexin SNX3 using cryo-electron tomography. The retromer core retains its arched, scaffolding structure but changes its mode of membrane recruitment when assembled with different SNX adaptors, allowing cargo recognition at subunit interfaces. Thus, membrane bending and cargo incorporation can be modulated to allow retromer to traffic cargoes along different cellular transport routes. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12223.map.gz emd_12223.map.gz | 4.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12223-v30.xml emd-12223-v30.xml emd-12223.xml emd-12223.xml | 20.8 KB 20.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12223_fsc.xml emd_12223_fsc.xml | 3.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_12223.png emd_12223.png | 102.6 KB | ||
| Masks |  emd_12223_msk_1.map emd_12223_msk_1.map | 4.3 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-12223.cif.gz emd-12223.cif.gz | 6.8 KB | ||
| Others |  emd_12223_half_map_1.map.gz emd_12223_half_map_1.map.gz emd_12223_half_map_2.map.gz emd_12223_half_map_2.map.gz | 4.1 MB 4.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12223 http://ftp.pdbj.org/pub/emdb/structures/EMD-12223 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12223 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12223 | HTTPS FTP |
-Related structure data
| Related structure data |  7blpMC  7blnC  7bloC  7blqC  7blrC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10631 (Title: Cryo-electron tomography of the fungal membrane-assembled retromer:Grd19 coat containing Kex2 cargo motif EMPIAR-10631 (Title: Cryo-electron tomography of the fungal membrane-assembled retromer:Grd19 coat containing Kex2 cargo motifData size: 347.7 Data #1: Raw image frames for the fungal retromer:Grd19 coat assembled on the Kex2 cargo-containing membranes [micrographs - multiframe] Data #2: Corrected, aligned and order-sorted tilt series for the membrane-reconstituted fungal retromer:Grd19 complex in the presence of cargo-signal containing C-portion of the Kex2 cargo. [tilt series] Data #3: Corrected, aligned, dose-filtered and order-sorted tilt series for the membrane-reconstituted fungal retromer:Grd19 complex in the presence of cargo-signal containing C-portion of the Kex2 cargo. [tilt series]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12223.map.gz / Format: CCP4 / Size: 4.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12223.map.gz / Format: CCP4 / Size: 4.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | LAFTER-filtered map of Vps35/Vps29 arch of fungal retromer:Grd19 complex on the membrane | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.758 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12223_msk_1.map emd_12223_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
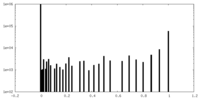
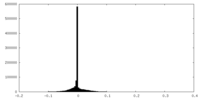
| Density Histograms |
-Half map: half-map1
| File | emd_12223_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
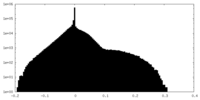
| Density Histograms |
-Half map: half-map2
| File | emd_12223_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
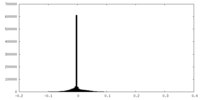
| Density Histograms |
- Sample components
Sample components
-Entire : Vps35/Vps29 arch of fungal membrane-assembled retromer:Grd19 complex.
| Entire | Name: Vps35/Vps29 arch of fungal membrane-assembled retromer:Grd19 complex. |
|---|---|
| Components |
|
-Supramolecule #1: Vps35/Vps29 arch of fungal membrane-assembled retromer:Grd19 complex.
| Supramolecule | Name: Vps35/Vps29 arch of fungal membrane-assembled retromer:Grd19 complex. type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: fungal retromer:Grd19 complex assembled on liposomes containing Kex2 cargo peptide. |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) |
-Macromolecule #1: Vacuolar protein sorting-associated protein 35
| Macromolecule | Name: Vacuolar protein sorting-associated protein 35 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus)Strain: DSM 1495 / CBS 144.50 / IMI 039719 |
| Molecular weight | Theoretical: 98.566781 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSTPAPPEEQ ARLLEDALIA VRQQTAMMRK FLDTPGKLMD ALKCCSTLVS ELRTSSLSPK QYYELYMAVF DALRYLSAHL RENHPVNHL ADLYELVQYA GNIIPRLYLM ITVGTAYMSI DGAPVKELMK DMMDMSRGVQ HPVRGLFLRY YLSGQARDYL P TGDSDGPE ...String: MSTPAPPEEQ ARLLEDALIA VRQQTAMMRK FLDTPGKLMD ALKCCSTLVS ELRTSSLSPK QYYELYMAVF DALRYLSAHL RENHPVNHL ADLYELVQYA GNIIPRLYLM ITVGTAYMSI DGAPVKELMK DMMDMSRGVQ HPVRGLFLRY YLSGQARDYL P TGDSDGPE GNLQDSINFI LTNFVEMNKL WVRLQHQGHS RERDLRTQER RELQLLVGSN IVRLSQLVDL PTYRDSILGP LL EQIVQCR DILAQEYLLE VITQVFPDEY HLHTLDQFLG AVSRLNPHVN VKAIVIGMMN RLSDYAERES QNEPEEDRAK LEE EALAKL LEKTKLGQNS ELEPQNGDHP DTEVSSTTDS AQAPSTADSD TTAVNGEEEP VRKRRGIPVN VPLYDIFFDQ VQHL VQAQH LPIQDTIALC CSLANLSLNI YPERLDYVDG ILAYALAKVK EHANSADLHS QPAQQSLLSL LQSPLRRYVS IFTAL SLPT YVSLFQAQTY PTRRAIAGEI VRTLLKNQTL ISTPAHLENV LEILKVLIKE GSQPPAGYPG VVQPRARPLE TDETME EQG WLARLVHLIH SDDNDTQFRL LQMTRKAYAE GNERIRTTTP PLITAGLKLA RRFKAREHYD DNWSSQSSSL FKFLHSA IS TLYTRVNGPG VADLCLRLFC SCGQVADMTE FEEVAYEFFA QAFTVYEESI SDSKAQFQAV CVIASALHRT RNFGRENY D TLITKCAQHA SKLLRKPDQC RAVYLASHLW WATPIAARGE TEDTELYRDG KRVLECLQRA LRVADSCMET ATSIELFVE ILDRYVYYFD QRNESVTTKY LNGLIELIHS NLAGNQQDSA SVEASRKHFI QTLEMIQSKE FEGIVVAPK UniProtKB: Vacuolar protein sorting-associated protein 35 |
-Macromolecule #2: Vacuolar protein sorting-associated protein 29
| Macromolecule | Name: Vacuolar protein sorting-associated protein 29 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus)Strain: DSM 1495 / CBS 144.50 / IMI 039719 |
| Molecular weight | Theoretical: 22.330602 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SMAFLILVIG NLHIPDRALD IPPKFKKLLS PGKISQTLCL GNLTDRATYD YLRSISPDLK IVRGRMDVEA TSLPLMQVVT HGSLRIGFL EGFTLVSEEP DVLLAEANKL DVDVLCWAGG SHRFECFEYM DKFFVNPGSA TGAFTTDWLA EGEEVVPSFC L MDVQGISL ...String: SMAFLILVIG NLHIPDRALD IPPKFKKLLS PGKISQTLCL GNLTDRATYD YLRSISPDLK IVRGRMDVEA TSLPLMQVVT HGSLRIGFL EGFTLVSEEP DVLLAEANKL DVDVLCWAGG SHRFECFEYM DKFFVNPGSA TGAFTTDWLA EGEEVVPSFC L MDVQGISL TLYVYQLRKD ENGTENVAVE KVTYTKPVEP TGAS UniProtKB: Vacuolar protein sorting-associated protein 29 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | 3D array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 3.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)