[English] 日本語
 Yorodumi
Yorodumi- EMDB-1132: A partial atomic structure for the flagellar hook of Salmonella t... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1132 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | A partial atomic structure for the flagellar hook of Salmonella typhimurium. | |||||||||
 Map data Map data | This is a map of the Salmonell Typhimurium Flagellar hook | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationbacterial-type flagellum basal body / bacterial-type flagellum-dependent swarming motility Similarity search - Function | |||||||||
| Biological species |  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 9.0 Å | |||||||||
 Authors Authors | Shaikh TR / Thomas DR / Chen JZ / Samatey FA / Matsunami H / Imada K / Namba K / DeRosier DJ | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2005 Journal: Proc Natl Acad Sci U S A / Year: 2005Title: A partial atomic structure for the flagellar hook of Salmonella typhimurium. Authors: Tanvir R Shaikh / Dennis R Thomas / James Z Chen / Fadel A Samatey / Hideyuki Matsunami / Katsumi Imada / Keiichi Namba / David J Derosier /  Abstract: The axial proteins of the bacterial flagellum function as a drive shaft, universal joint, and propeller driven by the flagellar rotary motor; they also form the putative protein export channel. The N- ...The axial proteins of the bacterial flagellum function as a drive shaft, universal joint, and propeller driven by the flagellar rotary motor; they also form the putative protein export channel. The N- and C-terminal sequences of the eight axial proteins were predicted to form interlocking alpha-domains generating an axial tube. We report on an approximately 1-nm resolution map of the hook from Salmonella typhimurium, which reveals such a tube made from interdigitated, 1-nm rod-like densities similar to those seen in maps of the filament. Atomic models for the two outer domains of the hook subunit were docked into the corresponding outermost features of the map. The N and C termini of the hook subunit fragment are positioned next to each other and face toward the axis of the hook. The placement of these termini would permit the residues missing in the fragment to form the rod-like features that form the core domain of the hook. We also fit the hook atomic model to an approximately 2-nm resolution map of the hook from Caulobacter crescentus. The hook protein sequence from C. crescentus is largely homologous to that of S. typhimurium except for a large insertion (20 kDa). According to difference maps and our fitting, this insertion is found on the outer surface of the hook, consistent with our modeling of the hook. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1132.map.gz emd_1132.map.gz | 3.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1132-v30.xml emd-1132-v30.xml emd-1132.xml emd-1132.xml | 10.8 KB 10.8 KB | Display Display |  EMDB header EMDB header |
| Images |  1132.gif 1132.gif | 17.6 KB | ||
| Filedesc layerLines |  emd_1132_ll.cif.gz emd_1132_ll.cif.gz | 23.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1132 http://ftp.pdbj.org/pub/emdb/structures/EMD-1132 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1132 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1132 | HTTPS FTP |
-Validation report
| Summary document |  emd_1132_validation.pdf.gz emd_1132_validation.pdf.gz | 231.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1132_full_validation.pdf.gz emd_1132_full_validation.pdf.gz | 230.2 KB | Display | |
| Data in XML |  emd_1132_validation.xml.gz emd_1132_validation.xml.gz | 4.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1132 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1132 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1132 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1132 | HTTPS FTP |
-Related structure data
| Related structure data |  2bgyMC  2bgzMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1132.map.gz / Format: CCP4 / Size: 4.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1132.map.gz / Format: CCP4 / Size: 4.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is a map of the Salmonell Typhimurium Flagellar hook | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.18 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
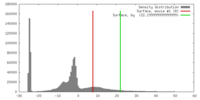
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Salmonella typhimurium Flagellar hook
| Entire | Name: Salmonella typhimurium Flagellar hook |
|---|---|
| Components |
|
-Supramolecule #1000: Salmonella typhimurium Flagellar hook
| Supramolecule | Name: Salmonella typhimurium Flagellar hook / type: sample / ID: 1000 Details: sample is derived from a polyhook mutant in the hook length controling gene fliK Oligomeric state: Segment of helix containing 75 hooksubunits Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 3.1 MDa / Method: amino acid sequence |
-Macromolecule #1: flagellar hook protein
| Macromolecule | Name: flagellar hook protein / type: protein_or_peptide / ID: 1 / Name.synonym: FlgE / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria)Strain: SJW880 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 8 / Details: 10 mM tris, 5 mM EDTA, 0.1% triton X-100 |
|---|---|
| Grid | Details: 300 mesh copper grids with holey carbon films |
| Vitrification | Cryogen name: ETHANE / Instrument: HOMEMADE PLUNGER Details: Vitrification instrument: gravity plunger. vitrification done in cold room at 4 deg C. Polyhooks are straight at 4 deg. |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM200FEG |
|---|---|
| Temperature | Average: 100 K |
| Alignment procedure | Legacy - Astigmatism: corrected at 300,000 |
| Image recording | Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 21 µm / Average electron dose: 10 e/Å2 / Bits/pixel: 8 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 2.7 µm / Nominal defocus min: 1.3 µm / Nominal magnification: 66000 |
| Sample stage | Specimen holder: cryo / Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| Details | Hook is naturally helical when assembled |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 9.0 Å / Resolution method: OTHER / Software - Name: BRANDEIS HELICAL PACKAGE Details: average includes 262 datasets and included 46 layerlines out of 84 collected. |
| CTF correction | Details: corrected averaged layerlines |
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A |
|---|---|
| Software | Name: RSRef |
| Details | PDBEntryID_givenInChain. Protocol: semi-rigid body. Two domains are kept as independent rigid bodies, connected by flexible peptide. Refinement is via simulated annealing molecular dynamics, including the adjacent subunits in the helix. |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Target criteria: correl. coeff. |
| Output model |  PDB-2bgy:  PDB-2bgz: |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)