+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0988 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Dimethylformamidase, tetramer | ||||||||||||
 Map data Map data | One of the primary map from the refinement in Relion | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | ab polypeptide / mononuclear iron / amidohydrolase / tetramer / HYDROLASE | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationN,N-dimethylformamidase / N,N-dimethylformamidase activity / metal ion binding Similarity search - Function | ||||||||||||
| Biological species |  Paracoccus sp. SSG05 (bacteria) Paracoccus sp. SSG05 (bacteria) | ||||||||||||
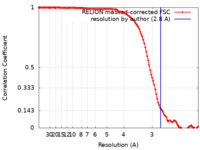
| Method | single particle reconstruction / cryo EM / Resolution: 2.8 Å | ||||||||||||
 Authors Authors | Arya CA / Yadav S | ||||||||||||
| Funding support |  India, 3 items India, 3 items
| ||||||||||||
 Citation Citation | Journal: Prog Biophys Mol Biol / Year: 2021 Title: Comparison of CryoEM and X-ray structures of dimethylformamidase. Authors: Kutti R Vinothkumar / Chetan Kumar Arya / Gurunath Ramanathan / Ramaswamy Subramanian /   Abstract: Dimethylformamidase (DMFase) catalyzes the hydrolysis of dimethylformamide, an industrial solvent, introduced into the environment by humans. Recently, we determined the structures of ...Dimethylformamidase (DMFase) catalyzes the hydrolysis of dimethylformamide, an industrial solvent, introduced into the environment by humans. Recently, we determined the structures of dimethylformamidase by electron cryo microscopy and X-ray crystallography revealing a tetrameric enzyme with a mononuclear iron at the active site. DMFase from Paracoccus sp. isolated from a waste water treatment plant around the city of Kanpur in India shows maximal activity at 54 °C and is halotolerant. The structures determined by both techniques are mostly identical and the largest difference is in a loop near the active site. This loop could play a role in co-operativity between the monomers. A number of non-protein densities are observed in the EM map, which are modelled as water molecules. Comparison of the structures determined by the two methods reveals conserved water molecules that could play a structural role. The higher stability, unusual active site and negligible activity at low temperature makes this a very good model to study enzyme mechanism by cryoEM. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0988.map.gz emd_0988.map.gz | 116.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0988-v30.xml emd-0988-v30.xml emd-0988.xml emd-0988.xml | 23.3 KB 23.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0988_fsc.xml emd_0988_fsc.xml | 11.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_0988.png emd_0988.png | 49.2 KB | ||
| Filedesc metadata |  emd-0988.cif.gz emd-0988.cif.gz | 7.1 KB | ||
| Others |  emd_0988_half_map_1.map.gz emd_0988_half_map_1.map.gz emd_0988_half_map_2.map.gz emd_0988_half_map_2.map.gz | 96.6 MB 96.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0988 http://ftp.pdbj.org/pub/emdb/structures/EMD-0988 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0988 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0988 | HTTPS FTP |
-Related structure data
| Related structure data |  6lvbMC  0989C  0990C  0991C  6lvcC  6lvdC  6lveC  6lvvC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_0988.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0988.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | One of the primary map from the refinement in Relion | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: One of the half map from the refinement in Relion
| File | emd_0988_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | One of the half map from the refinement in Relion | ||||||||||||
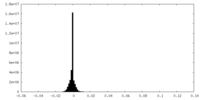
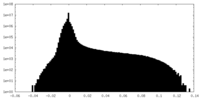
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: One of the half map from the refinement in Relion
| File | emd_0988_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | One of the half map from the refinement in Relion | ||||||||||||
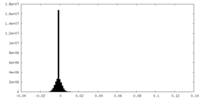
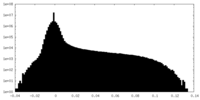
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Dimethylformamidase
| Entire | Name: Dimethylformamidase |
|---|---|
| Components |
|
-Supramolecule #1: Dimethylformamidase
| Supramolecule | Name: Dimethylformamidase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 / Details: Tetramer, 2x(a2b2) |
|---|---|
| Source (natural) | Organism:  Paracoccus sp. SSG05 (bacteria) Paracoccus sp. SSG05 (bacteria) |
| Molecular weight | Theoretical: 400 KDa |
-Macromolecule #1: N,N-dimethylformamidase large subunit
| Macromolecule | Name: N,N-dimethylformamidase large subunit / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number: N,N-dimethylformamidase |
|---|---|
| Source (natural) | Organism:  Paracoccus sp. SSG05 (bacteria) Paracoccus sp. SSG05 (bacteria) |
| Molecular weight | Theoretical: 86.341758 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKDIAIRGYC DRPSVATGET IRFYVSANET RGTFDAELVR LIHGDSNPAG PGYKEEAIKS DLEGQYPARF QRTQFGSYVE VADPDAGLQ PDGAFSVHLF LWSTTPSRGR QGIASRWNDE RQSGWNLAIE DGRVVFTIGD GSGATSSVVS DRPLFQQIWY S ITGVYDPE ...String: MKDIAIRGYC DRPSVATGET IRFYVSANET RGTFDAELVR LIHGDSNPAG PGYKEEAIKS DLEGQYPARF QRTQFGSYVE VADPDAGLQ PDGAFSVHLF LWSTTPSRGR QGIASRWNDE RQSGWNLAIE DGRVVFTIGD GSGATSSVVS DRPLFQQIWY S ITGVYDPE KKQLRLYQKS VVNRTNSRFG LVVPLDSDCA VSADATVKAA DSETSLLIAG LGEAAAQDGR TWCIAHYNGK VD APKIYGC ALGQDDAEKL SRGEIVRPIS RLAHWDFSAG IGLNGIPTDH VVDASGYGHH GRCMNQPSRG STGWNWDGHE ENF IHCPEQ YGALWFHEDC LDDCRWEKDF EFTVPEGLKS DFYAVKIRYE DTEDYIPFFV LPPRGTATAP ILVIASTLSY LAYA NEQIM HKADIGQAVA GHTPVLNEND VELHKNLSYY GLSTYDGHID GRGVQYTSWR RPIMNLRPKH RQGFGSIWEL PADLH LIDW LNHNGFEYDV ATEHDLNDQG AELLRRYKVV LTGSHPEYQT WANADAWEDY LADGGRGMYL AANGMYWIVE VHPEKP WVM EVRKELGVTA WEAPPGEYHY STNGRRGGRF RGRARATQKI WGTGMSSFGF DHSGYFVQMP DSQDERVAWI MEGIDPE ER IGDGGLVGGG AGGYELDRYD LALGTPPNTL LLASSVEHSV VYTVIPDDKA FPHPGMNGGE HPFVRADITY FSTANGGG M FATSSISWLG SLSWNDYDNN VSKMTKNVLN QFIKDEPAPR VKLAAALEHH HHHH UniProtKB: N,N-dimethylformamidase large subunit |
-Macromolecule #2: N,N-dimethylformamidase small subunit
| Macromolecule | Name: N,N-dimethylformamidase small subunit / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO / EC number: N,N-dimethylformamidase |
|---|---|
| Source (natural) | Organism:  Paracoccus sp. SSG05 (bacteria) Paracoccus sp. SSG05 (bacteria) |
| Molecular weight | Theoretical: 16.083823 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTEASESCVR DPSNYRDRSA DWYAFYDERR RKEIIDIIDE HPEIVEEHAA NPFGYRKHPS PYLQRVHNYF RMQPTFGRYY IYSEREWDA YRIATIREFG ELPELGDERF KTEEEAMHAV FLRRIEDVRA ELA UniProtKB: N,N-dimethylformamidase small subunit |
-Macromolecule #3: FE (III) ION
| Macromolecule | Name: FE (III) ION / type: ligand / ID: 3 / Number of copies: 4 / Formula: FE |
|---|---|
| Molecular weight | Theoretical: 55.845 Da |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 4 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL | ||||||
|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.2 / Component:
| ||||||
| Grid | Model: Quantifoil R0.6/1 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 90 sec. / Pretreatment - Atmosphere: AIR Details: Glow discharge was carried out in Quorum Glocube instrument at 25mA for 90 seconds. | ||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 291 K / Instrument: FEI VITROBOT MARK IV Details: Blot force was 0 and blotting was done for 3.5 seconds. | ||||||
| Details | Sample at higher salt exists mostly as tetramer |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 77.7 K / Max: 77.7 K |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 793 / Average exposure time: 60.0 sec. / Average electron dose: 27.0 e/Å2 Details: A total of 25 frames were saved from the 60 second exposure, resulting in ~1.08 electron/frame |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated defocus max: 4.1457 µm / Calibrated defocus min: 0.9315 µm / Calibrated magnification: 130841 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.1457 µm / Nominal defocus min: 0.9315 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER / Overall B value: 38.8 |
|---|---|
| Output model |  PDB-6lvb: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)