[English] 日本語
 Yorodumi
Yorodumi- EMDB-42388: Cryo-EM map of the human CTF18-RFC-PCNA-DNA ternary complex with ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM map of the human CTF18-RFC-PCNA-DNA ternary complex with narrow PCNA opening state II (state 6) | |||||||||
 Map data Map data | EM sharpen map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | DNA clamp loader complex / REPLICATION-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of DNA-directed DNA polymerase activity / Elg1 RFC-like complex / DNA replication factor C complex / Ctf18 RFC-like complex / dinucleotide insertion or deletion binding / PCNA-p21 complex / mitotic telomere maintenance via semi-conservative replication / purine-specific mismatch base pair DNA N-glycosylase activity / DNA clamp loader activity / nuclear lamina ...positive regulation of DNA-directed DNA polymerase activity / Elg1 RFC-like complex / DNA replication factor C complex / Ctf18 RFC-like complex / dinucleotide insertion or deletion binding / PCNA-p21 complex / mitotic telomere maintenance via semi-conservative replication / purine-specific mismatch base pair DNA N-glycosylase activity / DNA clamp loader activity / nuclear lamina / Polymerase switching / Processive synthesis on the lagging strand / MutLalpha complex binding / PCNA complex / Telomere C-strand (Lagging Strand) Synthesis / Removal of the Flap Intermediate / Mismatch repair (MMR) directed by MSH2:MSH3 (MutSbeta) / Mismatch repair (MMR) directed by MSH2:MSH6 (MutSalpha) / Transcription of E2F targets under negative control by DREAM complex / Polymerase switching on the C-strand of the telomere / Processive synthesis on the C-strand of the telomere / replisome / Removal of the Flap Intermediate from the C-strand / response to L-glutamate / response to dexamethasone / HDR through Single Strand Annealing (SSA) / DNA strand elongation involved in DNA replication / histone acetyltransferase binding / leading strand elongation / G1/S-Specific Transcription / DNA polymerase processivity factor activity / DNA synthesis involved in DNA repair / Impaired BRCA2 binding to RAD51 / nuclear replication fork / SUMOylation of DNA replication proteins / replication fork processing / PCNA-Dependent Long Patch Base Excision Repair / Presynaptic phase of homologous DNA pairing and strand exchange / response to cadmium ion / ATP-dependent activity, acting on DNA / estrous cycle / Activation of ATR in response to replication stress / mismatch repair / cyclin-dependent protein kinase holoenzyme complex / translesion synthesis / base-excision repair, gap-filling / DNA polymerase binding / epithelial cell differentiation / liver regeneration / TP53 Regulates Transcription of Genes Involved in G2 Cell Cycle Arrest / positive regulation of DNA replication / nuclear estrogen receptor binding / positive regulation of DNA repair / replication fork / Translesion synthesis by REV1 / Translesion synthesis by POLK / Translesion synthesis by POLI / male germ cell nucleus / Gap-filling DNA repair synthesis and ligation in GG-NER / Termination of translesion DNA synthesis / Translesion Synthesis by POLH / Recognition of DNA damage by PCNA-containing replication complex / G2/M DNA damage checkpoint / receptor tyrosine kinase binding / DNA-templated DNA replication / HDR through Homologous Recombination (HRR) / cellular response to xenobiotic stimulus / Dual Incision in GG-NER / cellular response to hydrogen peroxide / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / cellular response to UV / response to estradiol / E3 ubiquitin ligases ubiquitinate target proteins / heart development / chromatin organization / Processing of DNA double-strand break ends / Regulation of TP53 Activity through Phosphorylation / damaged DNA binding / chromosome, telomeric region / DNA replication / nuclear body / DNA repair / chromatin binding / centrosome / chromatin / protein-containing complex binding / enzyme binding / negative regulation of transcription by RNA polymerase II / ATP hydrolysis activity / DNA binding / extracellular exosome / nucleoplasm / ATP binding / membrane / identical protein binding / nucleus / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
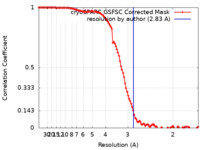
| Method | single particle reconstruction / cryo EM / Resolution: 2.83 Å | |||||||||
 Authors Authors | Wang F / He Q / Li H | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2024 Journal: Proc Natl Acad Sci U S A / Year: 2024Title: Cryo-EM reveals a nearly complete PCNA loading process and unique features of the human alternative clamp loader CTF18-RFC. Authors: Qing He / Feng Wang / Michael E O'Donnell / Huilin Li /  Abstract: The DNA sliding clamp PCNA is a multipurpose platform for DNA polymerases and many other proteins involved in DNA metabolism. The topologically closed PCNA ring needs to be cracked open and loaded ...The DNA sliding clamp PCNA is a multipurpose platform for DNA polymerases and many other proteins involved in DNA metabolism. The topologically closed PCNA ring needs to be cracked open and loaded onto DNA by a clamp loader, e.g., the well-studied pentameric ATPase complex RFC (RFC1-5). The CTF18-RFC complex is an alternative clamp loader found recently to bind the leading strand DNA polymerase ε and load PCNA onto leading strand DNA, but its structure and the loading mechanism have been unknown. By cryo-EM analysis of in vitro assembled human CTF18-RFC-DNA-PCNA complex, we have captured seven loading intermediates, revealing a detailed PCNA loading mechanism onto a 3'-ss/dsDNA junction by CTF18-RFC. Interestingly, the alternative loader has evolved a highly mobile CTF18 AAA+ module likely to lower the loading activity, perhaps to avoid competition with the RFC and to limit its role to leading strand clamp loading. To compensate for the lost stability due to the mobile AAA+ module, CTF18 has evolved a unique β-hairpin motif that reaches across RFC2 to interact with RFC5, thereby stabilizing the pentameric complex. Further, we found that CTF18 also contains a separation pin to locally melt DNA from the 3'-end of the primer; this ensures its ability to load PCNA to any 3'-ss/dsDNA junction, facilitated by the binding energy of the E-plug to the major groove. Our study reveals unique structural features of the human CTF18-RFC and contributes to a broader understanding of PCNA loading by the alternative clamp loaders. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42388.map.gz emd_42388.map.gz | 117.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42388-v30.xml emd-42388-v30.xml emd-42388.xml emd-42388.xml | 31.8 KB 31.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_42388_fsc.xml emd_42388_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_42388.png emd_42388.png | 61.6 KB | ||
| Filedesc metadata |  emd-42388.cif.gz emd-42388.cif.gz | 8.8 KB | ||
| Others |  emd_42388_additional_1.map.gz emd_42388_additional_1.map.gz emd_42388_additional_2.map.gz emd_42388_additional_2.map.gz emd_42388_half_map_1.map.gz emd_42388_half_map_1.map.gz emd_42388_half_map_2.map.gz emd_42388_half_map_2.map.gz | 110.8 MB 62.6 MB 116.1 MB 116.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42388 http://ftp.pdbj.org/pub/emdb/structures/EMD-42388 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42388 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42388 | HTTPS FTP |
-Related structure data
| Related structure data |  8umyMC  8umtC  8umuC  8umvC  8umwC  8un0C  8unjC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_42388.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42388.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EM sharpen map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.828 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: EM sharpen map by DeepEMhancer
| File | emd_42388_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EM sharpen map by DeepEMhancer | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Original unsharpen EM map
| File | emd_42388_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Original unsharpen EM map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: EM half map A
| File | emd_42388_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EM half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: EM half map B
| File | emd_42388_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | EM half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : the human clamp-clamp loader complex PCNA-CTF18
+Supramolecule #1: the human clamp-clamp loader complex PCNA-CTF18
+Macromolecule #1: Chromosome transmission fidelity protein 18 homolog
+Macromolecule #2: Replication factor C subunit 2
+Macromolecule #3: Replication factor C subunit 5
+Macromolecule #4: Replication factor C subunit 4
+Macromolecule #5: Replication factor C subunit 3
+Macromolecule #6: Proliferating cell nuclear antigen
+Macromolecule #7: DNA (40-MER)
+Macromolecule #8: DNA (20-MER)
+Macromolecule #9: MAGNESIUM ION
+Macromolecule #10: PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER
+Macromolecule #11: ADENOSINE-5'-DIPHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 280 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.9000000000000001 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Overall B value: 123 |
|---|---|
| Output model |  PDB-8umy: |
 Movie
Movie Controller
Controller



























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)