+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Goslar chimallin C4 tetramer localized reconstruction | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | phage / viral protein / STRUCTURAL PROTEIN | |||||||||||||||
| Function / homology | host cell cytoplasm / Chimallin Function and homology information Function and homology information | |||||||||||||||
| Biological species |  Goslarvirus / Goslarvirus /  Escherichia phage vB_EcoM_Goslar (virus) Escherichia phage vB_EcoM_Goslar (virus) | |||||||||||||||
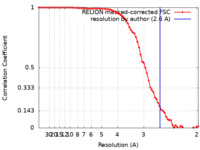
| Method | single particle reconstruction / cryo EM / Resolution: 2.6 Å | |||||||||||||||
 Authors Authors | Laughlin TG / Deep A / Prichard AM / Seitz C / Gu Y / Enustun E / Suslov S / Khanna K / Birkholz EA / Amaro RE ...Laughlin TG / Deep A / Prichard AM / Seitz C / Gu Y / Enustun E / Suslov S / Khanna K / Birkholz EA / Amaro RE / Pogliano J / Corbett KD / Villa E | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2022 Journal: Nature / Year: 2022Title: Architecture and self-assembly of the jumbo bacteriophage nuclear shell. Authors: Thomas G Laughlin / Amar Deep / Amy M Prichard / Christian Seitz / Yajie Gu / Eray Enustun / Sergey Suslov / Kanika Khanna / Erica A Birkholz / Emily Armbruster / J Andrew McCammon / Rommie ...Authors: Thomas G Laughlin / Amar Deep / Amy M Prichard / Christian Seitz / Yajie Gu / Eray Enustun / Sergey Suslov / Kanika Khanna / Erica A Birkholz / Emily Armbruster / J Andrew McCammon / Rommie E Amaro / Joe Pogliano / Kevin D Corbett / Elizabeth Villa /  Abstract: Bacteria encode myriad defences that target the genomes of infecting bacteriophage, including restriction-modification and CRISPR-Cas systems. In response, one family of large bacteriophages uses a ...Bacteria encode myriad defences that target the genomes of infecting bacteriophage, including restriction-modification and CRISPR-Cas systems. In response, one family of large bacteriophages uses a nucleus-like compartment to protect its replicating genomes by excluding host defence factors. However, the principal composition and structure of this compartment remain unknown. Here we find that the bacteriophage nuclear shell assembles primarily from one protein, which we name chimallin (ChmA). Combining cryo-electron tomography of nuclear shells in bacteriophage-infected cells and cryo-electron microscopy of a minimal chimallin compartment in vitro, we show that chimallin self-assembles as a flexible sheet into closed micrometre-scale compartments. The architecture and assembly dynamics of the chimallin shell suggest mechanisms for its nucleation and growth, and its role as a scaffold for phage-encoded factors mediating macromolecular transport, cytoskeletal interactions, and viral maturation. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25395.map.gz emd_25395.map.gz | 20.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25395-v30.xml emd-25395-v30.xml emd-25395.xml emd-25395.xml | 21.5 KB 21.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25395_fsc.xml emd_25395_fsc.xml | 7.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_25395.png emd_25395.png | 116.7 KB | ||
| Masks |  emd_25395_msk_1.map emd_25395_msk_1.map | 30.5 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-25395.cif.gz emd-25395.cif.gz | 6.3 KB | ||
| Others |  emd_25395_half_map_1.map.gz emd_25395_half_map_1.map.gz emd_25395_half_map_2.map.gz emd_25395_half_map_2.map.gz | 20.2 MB 20.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25395 http://ftp.pdbj.org/pub/emdb/structures/EMD-25395 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25395 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25395 | HTTPS FTP |
-Validation report
| Summary document |  emd_25395_validation.pdf.gz emd_25395_validation.pdf.gz | 839.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_25395_full_validation.pdf.gz emd_25395_full_validation.pdf.gz | 839.4 KB | Display | |
| Data in XML |  emd_25395_validation.xml.gz emd_25395_validation.xml.gz | 12.7 KB | Display | |
| Data in CIF |  emd_25395_validation.cif.gz emd_25395_validation.cif.gz | 17.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25395 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25395 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25395 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25395 | HTTPS FTP |
-Related structure data
| Related structure data |  7squMC  7sqqC  7sqrC  7sqsC  7sqtC  7sqvC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
| EM raw data |  EMPIAR-10863 (Title: Cryo-EM of recombinant Goslar chimallin / Data size: 432.9 EMPIAR-10863 (Title: Cryo-EM of recombinant Goslar chimallin / Data size: 432.9 Data #1: Unaligned frames as unormalized LZW-TIFF [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_25395.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25395.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.99019 Å | ||||||||||||||||||||||||||||||||||||
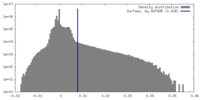
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_25395_msk_1.map emd_25395_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
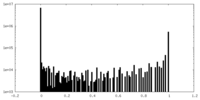
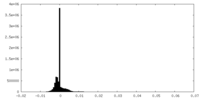
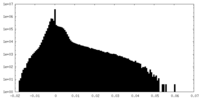
| Density Histograms |
-Half map: #1
| File | emd_25395_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
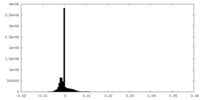
| Density Histograms |
-Half map: #2
| File | emd_25395_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
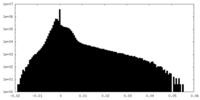
| Density Histograms |
- Sample components
Sample components
-Entire : Goslar chimallin tetramer localized reconstruction
| Entire | Name: Goslar chimallin tetramer localized reconstruction |
|---|---|
| Components |
|
-Supramolecule #1: Goslar chimallin tetramer localized reconstruction
| Supramolecule | Name: Goslar chimallin tetramer localized reconstruction / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Goslarvirus Goslarvirus |
-Macromolecule #1: Chimallin
| Macromolecule | Name: Chimallin / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Escherichia phage vB_EcoM_Goslar (virus) Escherichia phage vB_EcoM_Goslar (virus) |
| Molecular weight | Theoretical: 69.889445 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SNAMGLDVRN NGNDNVEIRA AETRTAQRAD EALETAADFA GQPKVTHTMR TINRTLSRRI SRNTGSEQVL NLRRLMEKYL EDTRFKDDF IFVAVDPNQY SVPYPTLVVM SGAKVGDHNH FFGYVLPLVA GLAPLPRREE QGPHGNILVP RTWVDNLNGT F INEVMAAM ...String: SNAMGLDVRN NGNDNVEIRA AETRTAQRAD EALETAADFA GQPKVTHTMR TINRTLSRRI SRNTGSEQVL NLRRLMEKYL EDTRFKDDF IFVAVDPNQY SVPYPTLVVM SGAKVGDHNH FFGYVLPLVA GLAPLPRREE QGPHGNILVP RTWVDNLNGT F INEVMAAM YAAIGGKSNG TARIAGLAVV TNEITAESAH LATTLLSAAD NAIQTAIEIR LGDKLGLPQF NLGMMASDQP IS SVQYNTS GMQDSDIVGN PVRSDITVTI SNRIRQAMSD YDSQQRLVAT TGYIDLTYSP QNPTFNQGPV LVNGYPVPPT VQY QPRYVM TSAYPLELDA FTPNTFVLGL IGTIATLNSG MAWAQSLISN AARGIGPHNP GALAMVLDPE VTAPLDLSTQ TNEQ IYKFL QQVLYPSLLI SIDVPEEGEY SWLLRMIPAA EKIYTGKVEG EVREISEGYK ALYRAFDDVT LGCFSKKYQY GLPLV YATG NRIPLGHYNH QDGHRHDIRD MDDLYMMNIT NPDTVEAWED SFDRTDMTMS QRVVARHEII DRVLSGSWEQ TGWAMR YDF DPLALQALIE AAADAGFTIR PENIQHLAGT AVRGNMAARA RGLGNISGNI YARSDRPNVG VNNMGGAFNL F UniProtKB: Chimallin |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.5 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.019 kPa / Details: 20 mA |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 289 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-44 / Number grids imaged: 1 / Number real images: 3921 / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)