+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of mouse heavy-chain apoferritin | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | apoferritin / mouse / heavy-chain / high resolution / METAL BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationIron uptake and transport / Golgi Associated Vesicle Biogenesis / negative regulation of ferroptosis / iron ion sequestering activity / autolysosome / ferroxidase / intracellular sequestering of iron ion / ferroxidase activity / negative regulation of fibroblast proliferation / endocytic vesicle lumen ...Iron uptake and transport / Golgi Associated Vesicle Biogenesis / negative regulation of ferroptosis / iron ion sequestering activity / autolysosome / ferroxidase / intracellular sequestering of iron ion / ferroxidase activity / negative regulation of fibroblast proliferation / endocytic vesicle lumen / Neutrophil degranulation / ferric iron binding / ferrous iron binding / iron ion transport / iron ion binding / immune response / negative regulation of cell population proliferation / mitochondrion / extracellular region / identical protein binding / membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
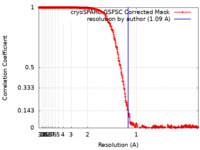
| Method | single particle reconstruction / cryo EM / Resolution: 1.09 Å | |||||||||
 Authors Authors | Nazarov S / Myasnikov A / Stahlberg H | |||||||||
| Funding support |  Switzerland, 1 items Switzerland, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM structure of mouse heavy-chain apoferritin Authors: Nazarov S / Mohammed I / Kucukoglu B / Myasnikov A / Stahlberg H | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_19436.map.gz emd_19436.map.gz | 431.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-19436-v30.xml emd-19436-v30.xml emd-19436.xml emd-19436.xml | 17.1 KB 17.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_19436_fsc.xml emd_19436_fsc.xml | 26.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_19436.png emd_19436.png | 103.1 KB | ||
| Masks |  emd_19436_msk_1.map emd_19436_msk_1.map | 465.5 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-19436.cif.gz emd-19436.cif.gz | 5.6 KB | ||
| Others |  emd_19436_additional_1.map.gz emd_19436_additional_1.map.gz emd_19436_half_map_1.map.gz emd_19436_half_map_1.map.gz emd_19436_half_map_2.map.gz emd_19436_half_map_2.map.gz | 426.9 MB 431.5 MB 430.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-19436 http://ftp.pdbj.org/pub/emdb/structures/EMD-19436 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19436 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-19436 | HTTPS FTP |
-Validation report
| Summary document |  emd_19436_validation.pdf.gz emd_19436_validation.pdf.gz | 1.5 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_19436_full_validation.pdf.gz emd_19436_full_validation.pdf.gz | 1.5 MB | Display | |
| Data in XML |  emd_19436_validation.xml.gz emd_19436_validation.xml.gz | 30.1 KB | Display | |
| Data in CIF |  emd_19436_validation.cif.gz emd_19436_validation.cif.gz | 40.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19436 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19436 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19436 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-19436 | HTTPS FTP |
-Related structure data
| Related structure data |  8rqbMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_19436.map.gz / Format: CCP4 / Size: 465.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_19436.map.gz / Format: CCP4 / Size: 465.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.30528 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_19436_msk_1.map emd_19436_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_19436_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_19436_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_19436_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Apoferritin from Mus musculus
| Entire | Name: Apoferritin from Mus musculus |
|---|---|
| Components |
|
-Supramolecule #1: Apoferritin from Mus musculus
| Supramolecule | Name: Apoferritin from Mus musculus / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Ferritin heavy chain, N-terminally processed
| Macromolecule | Name: Ferritin heavy chain, N-terminally processed / type: protein_or_peptide / ID: 1 / Number of copies: 24 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 20.079594 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: PSQVRQNYHQ DAEAAINRQI NLELYASYVY LSMSCYFDRD DVALKNFAKY FLHQSHEERE HAEKLMKLQN QRGGRIFLQD IKKPDRDDW ESGLNAMECA LHLEKSVNQS LLELHKLATD KNDPHLCDFI ETYYLSEQVK SIKELGDHVT NLRKMGAPEA G MAEYLFDK HTLG UniProtKB: Ferritin heavy chain |
-Macromolecule #2: FE (III) ION
| Macromolecule | Name: FE (III) ION / type: ligand / ID: 2 / Number of copies: 6 / Formula: FE |
|---|---|
| Molecular weight | Theoretical: 55.845 Da |
-Macromolecule #3: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 3 / Number of copies: 24 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #4: water
| Macromolecule | Name: water / type: ligand / ID: 4 / Number of copies: 2622 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 8 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 0.9 µm / Nominal defocus min: 0.3 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z
Z Y
Y X
X