+ Open data
Open data
- Basic information
Basic information
| Entry | 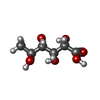 Database: PDB chemical components / ID: LFC Database: PDB chemical components / ID: LFC |
|---|---|
| Name | Name: |
-Chemical information
| Composition |  | ||||||
|---|---|---|---|---|---|---|---|
| Others | Type: L-saccharide / PDB classification: ATOMS / Three letter code: LFC / Model coordinates PDB-ID: 2HXU | ||||||
| History |
| ||||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  Brenda / Brenda /  ChEBI / ChEBI /  KEGG_Ligand / KEGG_Ligand /  Nikkaji / Nikkaji /  PubChem / PubChem /  SureChEMBL / SureChEMBL /  ZINC / ZINC /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 10.04 | | CACTVS 3.341 | OpenEye OEToolkits 1.5.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.341 | | OpenEye OEToolkits 1.5.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 10.04 | | OpenEye OEToolkits 1.5.0 | ( | |
|---|
-PDB entries
Showing all 2 items

PDB-2hxu: 
Crystal structure of K220A mutant of L-Fuconate Dehydratase from Xanthomonas campestris liganded with Mg++ and L-fuconate

PDB-4ovt: 
CRYSTAL STRUCTURE OF A TRAP PERIPLASMIC SOLUTE BINDING PROTEIN FROM OCHROBACTERIUM ANTHROPI (Oant_3902), TARGET EFI-510153, WITH BOUND L-FUCONATE
 Movie
Movie Controller
Controller



