+ Open data
Open data
- Basic information
Basic information
| Entry |  Database: PDB chemical components / ID: A3D Database: PDB chemical components / ID: A3D |
|---|---|
| Name | Name: |
-Chemical information
| Composition |  | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: A3D / Ideal coordinates details: Corina / Model coordinates PDB-ID: 1PZG / Replaces: NAC | ||||||||
| History |
| ||||||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  Brenda Brenda              / /  ChEBI / ChEBI /  DrugBank / DrugBank /  PubChem PubChem  / /  Wikipedia search / Wikipedia search /  Google search Google search |
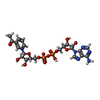
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| CACTVS 3.385 | | OpenEye OEToolkits 1.7.5 | |
|---|
-SMILES CANONICAL
| CACTVS 3.385 | | OpenEye OEToolkits 1.7.5 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| OpenEye OEToolkits 1.5.0 | [[( |
|---|
-PDB entries
Showing all 6 items

PDB-1drv: 
ESCHERICHIA COLI DHPR/ACNADH COMPLEX

PDB-1pzf: 
T.gondii LDH1 ternary complex with APAD+ and oxalate

PDB-1pzg: 
T.gondii LDH1 complexed with APAD and sulfate at 1.6 Angstroms

PDB-4nd2: 
Crystal structure of the lactate dehydrogenase from cryptosporidium parvum complexed with substrate (pyruvic acid) and cofactor analog (3-acetylpyridine adenine dinucleotide)

PDB-8qgw: 
Crystal structure of oxidized respiratory Complex I subunits NuoEF from Aquifex aeolicus bound to oxidized 3-acetylpyridine adenine dinucleotide

PDB-8qh4: 
Crystal structure of reduced respiratory Complex I subunits NuoEF from Aquifex aeolicus bound to oxidized 3-acetylpyridine adenine dinucleotide
 Movie
Movie Controller
Controller