+ Open data
Open data
- Basic information
Basic information
| Entry | 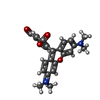 Database: PDB chemical components / ID: 323 Database: PDB chemical components / ID: 323 |
|---|---|
| Name | Name: Synonyms: 5-Carboxy-N,N'-tetramethyl rhodamine |
-Chemical information
| Composition |  | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: 323 / Ideal coordinates details: Corina / Model coordinates PDB-ID: 3D1F | ||||||||
| History |
| ||||||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  PubChem / PubChem /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 10.04 | [| CACTVS 3.341 | OpenEye OEToolkits 1.5.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.341 | | OpenEye OEToolkits 1.5.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 10.04 | | OpenEye OEToolkits 1.5.0 | |
|---|
-PDB entries
Showing all 4 items

PDB-3d1f: 
Crystal structure of E. coli sliding clamp (beta) bound to a polymerase III peptide

PDB-7v9b: 
Crystal Structure of 14-3-3 epsilon with FOXO3a peptide

PDB-8agc: 
Structure of yeast oligosaccharylransferase complex with lipid-linked oligosaccharide and non-acceptor peptide bound

PDB-8age: 
Structure of yeast oligosaccharylransferase complex with acceptor peptide bound
 Movie
Movie Controller
Controller



