+ Open data
Open data
- Basic information
Basic information
| Entry | 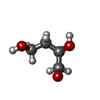 Database: PDB chemical components / ID: BGQ Database: PDB chemical components / ID: BGQ |
|---|---|
| Name | Name: |
-Chemical information
| Composition |  | ||||
|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: BGQ / Ideal coordinates details: Corina / Model coordinates PDB-ID: 4V1K | ||||
| History |
| ||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  Brenda / Brenda /  CompTox / CompTox /  PubChem / PubChem /  SureChEMBL / SureChEMBL /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 12.01 | | CACTVS 3.385 | [ | OpenEye OEToolkits 1.7.6 | |
|---|
-SMILES CANONICAL
| CACTVS 3.385 | [| OpenEye OEToolkits 1.7.6 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 12.01 | (| OpenEye OEToolkits 1.7.6 | ( | |
|---|
-PDB entries
Showing all 6 items

PDB-4v1k: 
SeMet structure of a novel carbohydrate binding module from glycoside hydrolase family 9 (Cel9A) from Ruminococcus flavefaciens FD-1

PDB-7c0k: 
Crystal structure of a dinucleotide-binding protein of ABC transporter endogenously bound to uridylyl-3'-5'-phospho-guanosine (Form II)

PDB-7c63: 
Crystal structure of beta-glycosides-binding protein of ABC transporter in an open state (Form I)

PDB-7f6u: 
Crystal structure of metal-citrate-binding mutant (Y221A) protein (MctA) of ABC transporter in apo state

PDB-8pha: 
O(S)-methyltransferase from Pleurotus sapidus

PDB-8v9i: 
1-deoxy-D-xylulose 5-phosphate synthase (DXPS) from Deinococcus radiodurans with D-phenylalanine-derived triazole acetylphosphonate (D-PheTrAP) bound
 Movie
Movie Controller
Controller



