+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ar0 | ||||||
|---|---|---|---|---|---|---|---|
| Title | NUCLEAR TRANSPORT FACTOR 2 (NTF2) E42K MUTANT | ||||||
 Components Components | NUCLEAR TRANSPORT FACTOR 2 | ||||||
 Keywords Keywords | TRANSPORT / NUCLEAR TRANSPORT PROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of vascular endothelial growth factor production / protein localization to nuclear pore / nuclear pore central transport channel / structural constituent of nuclear pore / nuclear inner membrane / nuclear import signal receptor activity / nuclear outer membrane / mRNA transport / protein export from nucleus / positive regulation of protein import into nucleus ...negative regulation of vascular endothelial growth factor production / protein localization to nuclear pore / nuclear pore central transport channel / structural constituent of nuclear pore / nuclear inner membrane / nuclear import signal receptor activity / nuclear outer membrane / mRNA transport / protein export from nucleus / positive regulation of protein import into nucleus / small GTPase binding / protein import into nucleus / nuclear membrane / nucleoplasm / identical protein binding / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.3 Å MOLECULAR REPLACEMENT / Resolution: 2.3 Å | ||||||
 Authors Authors | Mccoy, A.J. / Stewart, M.J. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1997 Journal: J.Mol.Biol. / Year: 1997Title: Nuclear protein import is decreased by engineered mutants of nuclear transport factor 2 (NTF2) that do not bind GDP-Ran. Authors: Clarkson, W.D. / Corbett, A.H. / Paschal, B.M. / Kent, H.M. / McCoy, A.J. / Gerace, L. / Silver, P.A. / Stewart, M. #1:  Journal: J.Mol.Biol. / Year: 1996 Journal: J.Mol.Biol. / Year: 1996Title: The 1.6 Angstroms Resolution Crystal Structure of Nuclear Transport Factor 2 (Ntf2) Authors: Bullock, T.L. / Clarkson, W.D. / Kent, H.M. / Stewart, M. #2:  Journal: Structure / Year: 1994 Journal: Structure / Year: 1994Title: Crystal Structure of Scytalone Dehydratase--A Disease Determinant of the Rice Pathogen, Magnaporthe Grisea Authors: Lundqvist, T. / Rice, J. / Hodge, C.N. / Basarab, G.S. / Pierce, J. / Lindqvist, Y. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ar0.cif.gz 1ar0.cif.gz | 60.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ar0.ent.gz pdb1ar0.ent.gz | 46.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ar0.json.gz 1ar0.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ar/1ar0 https://data.pdbj.org/pub/pdb/validation_reports/ar/1ar0 ftp://data.pdbj.org/pub/pdb/validation_reports/ar/1ar0 ftp://data.pdbj.org/pub/pdb/validation_reports/ar/1ar0 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  1askC 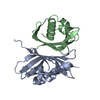 1ounS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 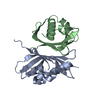
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (-0.914096, 0.124306, 0.385975), Vector: |
- Components
Components
| #1: Protein | Mass: 14491.491 Da / Num. of mol.: 2 / Mutation: E42K Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.47 Å3/Da / Density % sol: 47 % | |||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 4.5 / Details: pH 4.5 | |||||||||||||||||||||||||||||||||||||||||||||
| Crystal | *PLUS | |||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion / Details: Kent, H.M., (1996) J. Struct. Biol., 116, 325. | |||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 293 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: ELLIOTT GX-13 / Wavelength: 1.5418 ROTATING ANODE / Type: ELLIOTT GX-13 / Wavelength: 1.5418 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Nov 1, 1996 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.3→100 Å / Num. obs: 13342 / % possible obs: 99.5 % / Observed criterion σ(I): 0 / Redundancy: 4.2 % / Rmerge(I) obs: 0.077 / Net I/σ(I): 6.5 |
| Reflection shell | Resolution: 2.3→2.42 Å / Redundancy: 4 % / Rmerge(I) obs: 0.217 / Mean I/σ(I) obs: 3.4 / % possible all: 98.1 |
| Reflection shell | *PLUS % possible obs: 98.1 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: NTF2 (PDB ENTRY 1OUN) Resolution: 2.3→20 Å / Cross valid method: THROUGHOUT / σ(F): 0 Details: ASP 92 IN BOTH CHAINS WAS DISTORTED BY BEING IN AN EXTREMELY TIGHT SURFACE LOOP AND, ALTHOUGH IT HAD UNFAVORABLE PHI/PSI ANGLES, THE MODEL FITTED THE ELECTRON DENSITY CLOSELY.
| ||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.3→20 Å
| ||||||||||||||||||||
| Software | *PLUS Name: REFMAC / Classification: refinement | ||||||||||||||||||||
| Refinement | *PLUS Rfactor Rfree: 0.228 / Rfactor Rwork: 0.2 | ||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller












 PDBj
PDBj

