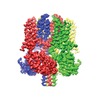
登録情報 データベース : EMDB / ID : EMD-8712タイトル CryoEM structure of Xenopus KCNQ1 channel Xenopus KCNQ1 channel, unsharpened map 複合体 : Complex of Xenopus KCNQ1 and CaMタンパク質・ペプチド : Potassium voltage-gated channel subfamily KQT member 1タンパク質・ペプチド : Calmodulinリガンド : CALCIUM ION / / / / 機能・相同性 分子機能 ドメイン・相同性 構成要素
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / 生物種 Xenopus laevis (アフリカツメガエル) / Homo sapiens (ヒト)手法 / / 解像度 : 3.7 Å Mackinnon R / Sun J 資金援助 Organization Grant number 国 Howard Hughes Medical Institute (HHMI) American Heart Association 17POST32260003
ジャーナル : Cell / 年 : 2017タイトル : Cryo-EM Structure of a KCNQ1/CaM Complex Reveals Insights into Congenital Long QT Syndrome.著者 : Ji Sun / Roderick MacKinnon / 要旨 : KCNQ1 is the pore-forming subunit of cardiac slow-delayed rectifier potassium (I) channels. Mutations in the kcnq1 gene are the leading cause of congenital long QT syndrome (LQTS). Here, we present ... KCNQ1 is the pore-forming subunit of cardiac slow-delayed rectifier potassium (I) channels. Mutations in the kcnq1 gene are the leading cause of congenital long QT syndrome (LQTS). Here, we present the cryoelectron microscopy (cryo-EM) structure of a KCNQ1/calmodulin (CaM) complex. The conformation corresponds to an "uncoupled," PIP-free state of KCNQ1, with activated voltage sensors and a closed pore. Unique structural features within the S4-S5 linker permit uncoupling of the voltage sensor from the pore in the absence of PIP. CaM contacts the KCNQ1 voltage sensor through a specific interface involving a residue on CaM that is mutated in a form of inherited LQTS. Using an electrophysiological assay, we find that this mutation on CaM shifts the KCNQ1 voltage-activation curve. This study describes one physiological form of KCNQ1, depolarized voltage sensors with a closed pore in the absence of PIP, and reveals a regulatory interaction between CaM and KCNQ1 that may explain CaM-mediated LQTS. 履歴 登録 2017年4月28日 - ヘッダ(付随情報) 公開 2017年5月24日 - マップ公開 2017年6月7日 - 更新 2024年3月13日 - 現状 2024年3月13日 処理サイト : RCSB / 状態 : 公開
すべて表示 表示を減らす
 データを開く
データを開く 基本情報
基本情報 マップデータ
マップデータ 試料
試料 キーワード
キーワード 機能・相同性情報
機能・相同性情報 Homo sapiens (ヒト)
Homo sapiens (ヒト) データ登録者
データ登録者 米国, 2件
米国, 2件  引用
引用 ジャーナル: Cell / 年: 2017
ジャーナル: Cell / 年: 2017
 構造の表示
構造の表示 ムービービューア
ムービービューア SurfView
SurfView Molmil
Molmil Jmol/JSmol
Jmol/JSmol ダウンロードとリンク
ダウンロードとリンク emd_8712.map.gz
emd_8712.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-8712-v30.xml
emd-8712-v30.xml emd-8712.xml
emd-8712.xml EMDBヘッダ
EMDBヘッダ emd_8712.png
emd_8712.png emd-8712.cif.gz
emd-8712.cif.gz emd_8712_additional.map.gz
emd_8712_additional.map.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-8712
http://ftp.pdbj.org/pub/emdb/structures/EMD-8712 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8712
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8712 emd_8712_validation.pdf.gz
emd_8712_validation.pdf.gz EMDB検証レポート
EMDB検証レポート emd_8712_full_validation.pdf.gz
emd_8712_full_validation.pdf.gz emd_8712_validation.xml.gz
emd_8712_validation.xml.gz emd_8712_validation.cif.gz
emd_8712_validation.cif.gz https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8712
https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8712 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8712
ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8712 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_8712.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_8712.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) 試料の構成要素
試料の構成要素 Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN
 画像解析
画像解析 ムービー
ムービー コントローラー
コントローラー

































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)