+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7kzx | ||||||
|---|---|---|---|---|---|---|---|
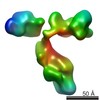
| Title | Cryo-EM structure of YiiP-Fab complex in Apo state | ||||||
 Components Components |
| ||||||
 Keywords Keywords | MEMBRANE PROTEIN / Zinc / transporter / cation diffusion facilitator / holo state / inward-facing state | ||||||
| Function / homology |  Function and homology information Function and homology informationzinc efflux antiporter activity / cadmium ion transmembrane transporter activity / ferrous iron transmembrane transporter activity / intracellular zinc ion homeostasis / metal ion binding / plasma membrane Similarity search - Function | ||||||
| Biological species |  Shewanella oneidensis (bacteria) Shewanella oneidensis (bacteria) Homo sapiens (human) Homo sapiens (human) | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 4 Å | ||||||
 Authors Authors | Lopez-Redondo, M.L. / Fan, S. / Koide, A. / Koide, S. / Beckstein, O. / Stokes, D.L. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: J Gen Physiol / Year: 2021 Journal: J Gen Physiol / Year: 2021Title: Zinc binding alters the conformational dynamics and drives the transport cycle of the cation diffusion facilitator YiiP. Authors: Maria Lopez-Redondo / Shujie Fan / Akiko Koide / Shohei Koide / Oliver Beckstein / David L Stokes /  Abstract: YiiP is a secondary transporter that couples Zn2+ transport to the proton motive force. Structural studies of YiiP from prokaryotes and Znt8 from humans have revealed three different Zn2+ sites and a ...YiiP is a secondary transporter that couples Zn2+ transport to the proton motive force. Structural studies of YiiP from prokaryotes and Znt8 from humans have revealed three different Zn2+ sites and a conserved homodimeric architecture. These structures define the inward-facing and outward-facing states that characterize the archetypal alternating access mechanism of transport. To study the effects of Zn2+ binding on the conformational transition, we use cryo-EM together with molecular dynamics simulation to compare structures of YiiP from Shewanella oneidensis in the presence and absence of Zn2+. To enable single-particle cryo-EM, we used a phage-display library to develop a Fab antibody fragment with high affinity for YiiP, thus producing a YiiP/Fab complex. To perform MD simulations, we developed a nonbonded dummy model for Zn2+ and validated its performance with known Zn2+-binding proteins. Using these tools, we find that, in the presence of Zn2+, YiiP adopts an inward-facing conformation consistent with that previously seen in tubular crystals. After removal of Zn2+ with high-affinity chelators, YiiP exhibits enhanced flexibility and adopts a novel conformation that appears to be intermediate between inward-facing and outward-facing states. This conformation involves closure of a hydrophobic gate that has been postulated to control access to the primary transport site. Comparison of several independent cryo-EM maps suggests that the transition from the inward-facing state is controlled by occupancy of a secondary Zn2+ site at the cytoplasmic membrane interface. This work enhances our understanding of individual Zn2+ binding sites and their role in the conformational dynamics that govern the transport cycle. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7kzx.cif.gz 7kzx.cif.gz | 244.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7kzx.ent.gz pdb7kzx.ent.gz | 191.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7kzx.json.gz 7kzx.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/kz/7kzx https://data.pdbj.org/pub/pdb/validation_reports/kz/7kzx ftp://data.pdbj.org/pub/pdb/validation_reports/kz/7kzx ftp://data.pdbj.org/pub/pdb/validation_reports/kz/7kzx | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  23092MC  7kzzC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 32485.211 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Shewanella oneidensis (bacteria) / Gene: fieF, SO_4475 / Production host: Shewanella oneidensis (bacteria) / Gene: fieF, SO_4475 / Production host:  #2: Antibody | Mass: 23580.242 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:  #3: Antibody | Mass: 25406.352 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:  Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component |
| |||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular weight |
| |||||||||||||||||||||||||||||||||||
| Source (natural) |
| |||||||||||||||||||||||||||||||||||
| Source (recombinant) |
| |||||||||||||||||||||||||||||||||||
| Buffer solution | pH: 7.5 | |||||||||||||||||||||||||||||||||||
| Buffer component |
| |||||||||||||||||||||||||||||||||||
| Specimen | Conc.: 3 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES Details: YiiP-Fab2R complex purified through SEC with DM, the same day as grid freezing. Main peak fraction was used for freezing. | |||||||||||||||||||||||||||||||||||
| Specimen support | Grid material: GOLD / Grid mesh size: 300 divisions/in. / Grid type: UltrAuFoil R1.2/1.3 | |||||||||||||||||||||||||||||||||||
| Vitrification | Instrument: FEI VITROBOT MARK IV / Cryogen name: ETHANE / Humidity: 100 % / Chamber temperature: 277 K Details: 3ul of protein mixture applied to grid. Blot time 4 seconds, blot force 0. |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 130000 X / Nominal defocus max: 3000 nm / Nominal defocus min: 1500 nm / Cs: 2.7 mm / C2 aperture diameter: 70 µm / Alignment procedure: COMA FREE |
| Specimen holder | Cryogen: NITROGEN / Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Temperature (max): 77 K / Temperature (min): 77 K |
| Image recording | Average exposure time: 10 sec. / Electron dose: 65 e/Å2 / Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Num. of grids imaged: 1 / Num. of real images: 1945 |
| EM imaging optics | Energyfilter name: GIF Bioquantum / Energyfilter slit width: 15 eV |
| Image scans | Sampling size: 14 µm / Width: 3710 / Height: 3838 / Movie frames/image: 50 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EM software |
| |||||||||||||||||||||||||||||||||||||||||||||
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | |||||||||||||||||||||||||||||||||||||||||||||
| Particle selection | Num. of particles selected: 848576 | |||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Point symmetry: C1 (asymmetric) | |||||||||||||||||||||||||||||||||||||||||||||
| 3D reconstruction | Resolution: 4 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 140565 / Algorithm: FOURIER SPACE / Num. of class averages: 2 / Symmetry type: POINT | |||||||||||||||||||||||||||||||||||||||||||||
| Atomic model building | B value: 101 / Protocol: BACKBONE TRACE / Space: REAL / Target criteria: cross-correlation | |||||||||||||||||||||||||||||||||||||||||||||
| Atomic model building | PDB-ID: 5VRF Accession code: 5VRF / Source name: PDB / Type: experimental model | |||||||||||||||||||||||||||||||||||||||||||||
| Refinement | Cross valid method: NONE Stereochemistry target values: GeoStd + Monomer Library + CDL v1.2 | |||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 101.64 Å2 | |||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller









 PDBj
PDBj


