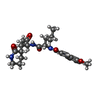
Entry Database : PDB / ID : 7mbiTitle Structure of SARS-CoV2 3CL protease covalently bound to peptidomimetic inhibitor 3C-like proteinase Keywords / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Method / / / Resolution : 2.15 Å Authors Khan, M.B. / Lu, J. / Young, H.S. / Lemieux, M.J. Funding support Organization Grant number Country Canadian Institutes of Health Research (CIHR) VR3-172655 Natural Sciences and Engineering Research Council (NSERC, Canada) 549297-2019
Journal : J.Med.Chem. / Year : 2022Title : Peptidomimetic alpha-Acyloxymethylketone Warheads with Six-Membered Lactam P1 Glutamine Mimic: SARS-CoV-2 3CL Protease Inhibition, Coronavirus Antiviral Activity, and in Vitro Biological Stability.Authors: Bai, B. / Belovodskiy, A. / Hena, M. / Kandadai, A.S. / Joyce, M.A. / Saffran, H.A. / Shields, J.A. / Khan, M.B. / Arutyunova, E. / Lu, J. / Bajwa, S.K. / Hockman, D. / Fischer, C. / Lamer, ... Authors : Bai, B. / Belovodskiy, A. / Hena, M. / Kandadai, A.S. / Joyce, M.A. / Saffran, H.A. / Shields, J.A. / Khan, M.B. / Arutyunova, E. / Lu, J. / Bajwa, S.K. / Hockman, D. / Fischer, C. / Lamer, T. / Vuong, W. / van Belkum, M.J. / Gu, Z. / Lin, F. / Du, Y. / Xu, J. / Rahim, M. / Young, H.S. / Vederas, J.C. / Tyrrell, D.L. / Lemieux, M.J. / Nieman, J.A. History Deposition Mar 31, 2021 Deposition site / Processing site Revision 1.0 Jul 21, 2021 Provider / Type Revision 1.1 Jul 28, 2021 Group / Category / Item / _citation.titleRevision 1.2 Mar 9, 2022 Group / Database referencesCategory / database_2 / pdbx_unobs_or_zero_occ_atomsItem _citation.journal_volume / _citation.page_first ... _citation.journal_volume / _citation.page_first / _citation.page_last / _citation.year / _database_2.pdbx_DOI / _database_2.pdbx_database_accession Revision 1.3 Oct 18, 2023 Group / Refinement descriptionCategory / chem_comp_bond / pdbx_initial_refinement_modelRevision 1.4 Nov 20, 2024 Group / Category / pdbx_modification_feature / Item
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.15 Å
MOLECULAR REPLACEMENT / Resolution: 2.15 Å  Authors
Authors Canada, 2items
Canada, 2items  Citation
Citation Journal: J.Med.Chem. / Year: 2022
Journal: J.Med.Chem. / Year: 2022 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 7mbi.cif.gz
7mbi.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb7mbi.ent.gz
pdb7mbi.ent.gz PDB format
PDB format 7mbi.json.gz
7mbi.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/mb/7mbi
https://data.pdbj.org/pub/pdb/validation_reports/mb/7mbi ftp://data.pdbj.org/pub/pdb/validation_reports/mb/7mbi
ftp://data.pdbj.org/pub/pdb/validation_reports/mb/7mbi
 Links
Links Assembly
Assembly


 Components
Components

 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  SSRL
SSRL  / Beamline: BL12-1 / Wavelength: 0.97946 Å
/ Beamline: BL12-1 / Wavelength: 0.97946 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller






















 PDBj
PDBj