[English] 日本語
 Yorodumi
Yorodumi- EMDB-7546: Monomer yeast ATP synthase (F1Fo) reconstituted in nanodisc with ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7546 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Monomer yeast ATP synthase (F1Fo) reconstituted in nanodisc with inhibitor of oligomycin bound. | |||||||||
 Map data Map data | Monomer yeast ATP synthase (F1Fo) reconstituted in nanodisc with inhibitor of oligomycin bound | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ATP synthase / BIOSYNTHETIC PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcristae formation / Mitochondrial protein degradation / mitochondrial proton-transporting ATP synthase complex assembly / proton transmembrane transporter activity / proton motive force-driven ATP synthesis / proton-transporting two-sector ATPase complex, proton-transporting domain / mitochondrial nucleoid / proton motive force-driven mitochondrial ATP synthesis / proton-transporting ATPase activity, rotational mechanism / H+-transporting two-sector ATPase ...cristae formation / Mitochondrial protein degradation / mitochondrial proton-transporting ATP synthase complex assembly / proton transmembrane transporter activity / proton motive force-driven ATP synthesis / proton-transporting two-sector ATPase complex, proton-transporting domain / mitochondrial nucleoid / proton motive force-driven mitochondrial ATP synthesis / proton-transporting ATPase activity, rotational mechanism / H+-transporting two-sector ATPase / proton-transporting ATP synthase complex / proton-transporting ATP synthase activity, rotational mechanism / proton transmembrane transport / ADP binding / mitochondrial intermembrane space / protein-containing complex assembly / mitochondrial inner membrane / lipid binding / mitochondrion / ATP binding / identical protein binding / cytosol Similarity search - Function | |||||||||
| Biological species |   | |||||||||
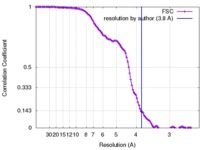
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Srivastava AP / Luo M | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2018 Journal: Science / Year: 2018Title: High-resolution cryo-EM analysis of the yeast ATP synthase in a lipid membrane. Authors: Anurag P Srivastava / Min Luo / Wenchang Zhou / Jindrich Symersky / Dongyang Bai / Melissa G Chambers / José D Faraldo-Gómez / Maofu Liao / David M Mueller /  Abstract: Mitochondrial adenosine triphosphate (ATP) synthase comprises a membrane embedded F motor that rotates to drive ATP synthesis in the F subunit. We used single-particle cryo-electron microscopy (cryo- ...Mitochondrial adenosine triphosphate (ATP) synthase comprises a membrane embedded F motor that rotates to drive ATP synthesis in the F subunit. We used single-particle cryo-electron microscopy (cryo-EM) to obtain structures of the full complex in a lipid bilayer in the absence or presence of the inhibitor oligomycin at 3.6- and 3.8-angstrom resolution, respectively. To limit conformational heterogeneity, we locked the rotor in a single conformation by fusing the F6 subunit of the stator with the δ subunit of the rotor. Assembly of the enzyme with the F6-δ fusion caused a twisting of the rotor and a 9° rotation of the F c-ring in the direction of ATP synthesis, relative to the structure of isolated F Our cryo-EM structures show how F and F are coupled, give insight into the proton translocation pathway, and show how oligomycin blocks ATP synthesis. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7546.map.gz emd_7546.map.gz | 116.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7546-v30.xml emd-7546-v30.xml emd-7546.xml emd-7546.xml | 29.6 KB 29.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_7546_fsc.xml emd_7546_fsc.xml | 13.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_7546.png emd_7546.png | 57.3 KB | ||
| Filedesc metadata |  emd-7546.cif.gz emd-7546.cif.gz | 7.6 KB | ||
| Others |  emd_7546_additional.map.gz emd_7546_additional.map.gz | 98.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7546 http://ftp.pdbj.org/pub/emdb/structures/EMD-7546 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7546 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7546 | HTTPS FTP |
-Related structure data
| Related structure data |  6cp3MC  7547C  7548C  7549C  6cp5C  6cp6C  6cp7C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_7546.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7546.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Monomer yeast ATP synthase (F1Fo) reconstituted in nanodisc with inhibitor of oligomycin bound | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
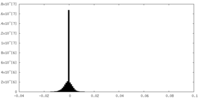
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.23 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Additional map
| File | emd_7546_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Additional map | ||||||||||||
| Projections & Slices |
| ||||||||||||
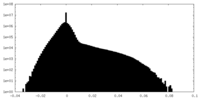
| Density Histograms |
- Sample components
Sample components
+Entire : Monomer yeast ATP synthase (F1Fo) reconstituted in nanodisc with ...
+Supramolecule #1: Monomer yeast ATP synthase (F1Fo) reconstituted in nanodisc with ...
+Macromolecule #1: ATP synthase subunit 9, mitochondrial
+Macromolecule #2: ATP synthase subunit 5, mitochondrial
+Macromolecule #3: ATP synthase subunit alpha, mitochondrial
+Macromolecule #4: ATP synthase subunit beta, mitochondrial
+Macromolecule #5: ATP synthase subunit gamma, mitochondrial
+Macromolecule #6: ATP synthase subunit delta, mitochondrial
+Macromolecule #7: ATP synthase subunit epsilon, mitochondrial
+Macromolecule #8: ATP synthase subunit 4, mitochondrial
+Macromolecule #9: ATP synthase subunit d, mitochondrial
+Macromolecule #10: ATP synthase subunit H, mitochondrial
+Macromolecule #11: ATP synthase subunit f, mitochondrial
+Macromolecule #12: ATP synthase protein 8
+Macromolecule #13: ATP synthase subunit a
+Macromolecule #14: ATP synthase subunit J, mitochondrial
+Macromolecule #15: ADENOSINE-5'-TRIPHOSPHATE
+Macromolecule #16: ADENOSINE-5'-DIPHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1. mg/mL |
|---|---|
| Buffer | pH: 8 / Details: 20 mM Tris-HCl, 150 mM NaCl, pH 8.0 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 91 % |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Temperature | Min: 80.0 K / Max: 105.0 K |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number real images: 2896 / Average electron dose: 8.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 2.7 µm / Calibrated defocus min: 1.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal magnification: 31000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-6cp3: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)