+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6kk3 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
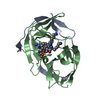
| Title | Crystal structure of Zika NS2B-NS3 protease with compound 4 | ||||||||||||
 Components Components | (Genome polyprotein) x 2 | ||||||||||||
 Keywords Keywords | VIRAL PROTEIN / Viral protease / Protease inhibitor complex | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated suppression of host JAK-STAT cascade via inhibition of host TYK2 activity / flavivirin / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT2 activity / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT1 activity / ribonucleoside triphosphate phosphatase activity / viral capsid / double-stranded RNA binding / nucleoside-triphosphate phosphatase / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of TBK1 activity / symbiont-mediated suppression of host toll-like receptor signaling pathway ...symbiont-mediated suppression of host JAK-STAT cascade via inhibition of host TYK2 activity / flavivirin / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT2 activity / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT1 activity / ribonucleoside triphosphate phosphatase activity / viral capsid / double-stranded RNA binding / nucleoside-triphosphate phosphatase / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of TBK1 activity / symbiont-mediated suppression of host toll-like receptor signaling pathway / mRNA (guanine-N7)-methyltransferase / methyltransferase cap1 / clathrin-dependent endocytosis of virus by host cell / mRNA (nucleoside-2'-O-)-methyltransferase activity / mRNA 5'-cap (guanine-N7-)-methyltransferase activity / RNA helicase activity / host cell perinuclear region of cytoplasm / host cell endoplasmic reticulum membrane / protein dimerization activity / symbiont-mediated suppression of host innate immune response / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / RNA helicase / symbiont entry into host cell / induction by virus of host autophagy / serine-type endopeptidase activity / RNA-directed RNA polymerase / viral RNA genome replication / virus-mediated perturbation of host defense response / RNA-dependent RNA polymerase activity / fusion of virus membrane with host endosome membrane / lipid binding / viral envelope / host cell nucleus / virion attachment to host cell / GTP binding / structural molecule activity / virion membrane / ATP hydrolysis activity / proteolysis / extracellular region / ATP binding / membrane / metal ion binding Similarity search - Function | ||||||||||||
| Biological species |   Zika virus Zika virus | ||||||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.05 Å MOLECULAR REPLACEMENT / Resolution: 2.05 Å | ||||||||||||
 Authors Authors | Quek, J.P. | ||||||||||||
| Funding support |  Singapore, 3items Singapore, 3items
| ||||||||||||
 Citation Citation |  Journal: Chemmedchem / Year: 2020 Journal: Chemmedchem / Year: 2020Title: Structure-Based Macrocyclization of Substrate Analogue NS2B-NS3 Protease Inhibitors of Zika, West Nile and Dengue viruses. Authors: Braun, N.J. / Quek, J.P. / Huber, S. / Kouretova, J. / Rogge, D. / Lang-Henkel, H. / Cheong, E.Z.K. / Chew, B.L.A. / Heine, A. / Luo, D. / Steinmetzer, T. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6kk3.cif.gz 6kk3.cif.gz | 93.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6kk3.ent.gz pdb6kk3.ent.gz | 68.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6kk3.json.gz 6kk3.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  6kk3_validation.pdf.gz 6kk3_validation.pdf.gz | 446.5 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  6kk3_full_validation.pdf.gz 6kk3_full_validation.pdf.gz | 447 KB | Display | |
| Data in XML |  6kk3_validation.xml.gz 6kk3_validation.xml.gz | 10 KB | Display | |
| Data in CIF |  6kk3_validation.cif.gz 6kk3_validation.cif.gz | 13.3 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/kk/6kk3 https://data.pdbj.org/pub/pdb/validation_reports/kk/6kk3 ftp://data.pdbj.org/pub/pdb/validation_reports/kk/6kk3 ftp://data.pdbj.org/pub/pdb/validation_reports/kk/6kk3 | HTTPS FTP |
-Related structure data
| Related structure data |  6kk2C  6kk4C  6kk5C  6kk6C  6kpqC  6y3bC  5gpiS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||||||||
| Unit cell |
| |||||||||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein/peptide | Mass: 4218.545 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Zika virus / Production host: Zika virus / Production host:  References: UniProt: Q32ZE1, flavivirin, nucleoside-triphosphate phosphatase, RNA helicase, mRNA (guanine-N7)-methyltransferase, methyltransferase cap1, RNA-directed RNA polymerase |
|---|---|
| #2: Protein | Mass: 16499.791 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Zika virus / Gene: GP1, A2G93_63394gpGP1 / Production host: Zika virus / Gene: GP1, A2G93_63394gpGP1 / Production host:  |
| #3: Chemical | ChemComp-DUU / |
| #4: Water | ChemComp-HOH / |
| Has ligand of interest | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.34 Å3/Da / Density % sol: 47.44 % |
|---|---|
| Crystal grow | Temperature: 291 K / Method: vapor diffusion, hanging drop Details: 0.2M Ammonium Sulfate, 0.1M Sodium Acetate Trihydrate pH 4.6, 30% Peg 2000 |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSRRC NSRRC  / Beamline: TPS 05A / Wavelength: 1 Å / Beamline: TPS 05A / Wavelength: 1 Å |
| Detector | Type: RAYONIX MX300-HS / Detector: CCD / Date: Nov 15, 2018 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 2.05→53.815 Å / Num. obs: 13375 / % possible obs: 99.9 % / Redundancy: 12 % / CC1/2: 1 / Rmerge(I) obs: 0.155 / Net I/σ(I): 10.6 |
| Reflection shell | Resolution: 2.05→2.12 Å / Redundancy: 12.4 % / Rmerge(I) obs: 1.436 / Num. unique obs: 1289 / CC1/2: 0.72 / % possible all: 100 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 5gpi Resolution: 2.05→53.815 Å / Cross valid method: NONE
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 97.02 Å2 / Biso min: 19.08 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 2.05→53.815 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.05→2.12 Å /
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj





