+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6803 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Human DNA-PK Holoenzyme | |||||||||||||||
 Map data Map data | None | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Cryo-EM structure / DNA-PK / DNAPKcs / activation / NHEJ / DNA BINDING PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of lymphocyte differentiation / small-subunit processome assembly / Ku70:Ku80 complex / DNA-dependent protein kinase complex / negative regulation of t-circle formation / DNA end binding / DNA-dependent protein kinase-DNA ligase 4 complex / MHC class II antigen presentation / nonhomologous end joining complex / cellular response to X-ray ...positive regulation of lymphocyte differentiation / small-subunit processome assembly / Ku70:Ku80 complex / DNA-dependent protein kinase complex / negative regulation of t-circle formation / DNA end binding / DNA-dependent protein kinase-DNA ligase 4 complex / MHC class II antigen presentation / nonhomologous end joining complex / cellular response to X-ray / regulation of smooth muscle cell proliferation / Cytosolic sensors of pathogen-associated DNA / Neutrophil degranulation / DNA ligation / IRF3-mediated induction of type I IFN / nuclear telomere cap complex / double-strand break repair via classical nonhomologous end joining / entry into host cell by a symbiont-containing vacuole / positive regulation of catalytic activity / U3 snoRNA binding / recombinational repair / regulation of telomere maintenance / protein localization to chromosome, telomeric region / positive regulation of neurogenesis / cellular response to fatty acid / cellular hyperosmotic salinity response / hematopoietic stem cell proliferation / Hydrolases; Acting on peptide bonds (peptidases); Aspartic endopeptidases / telomeric DNA binding / 2-LTR circle formation / : / site of DNA damage / protein autoprocessing / Lyases; Carbon-oxygen lyases; Other carbon-oxygen lyases / 5'-deoxyribose-5-phosphate lyase activity / hematopoietic stem cell differentiation / positive regulation of protein kinase activity / ATP-dependent activity, acting on DNA / activation of innate immune response / enzyme activator activity / transport vesicle / positive regulation of telomere maintenance via telomerase / telomere maintenance / DNA helicase activity / neurogenesis / cyclin binding / protein-DNA complex / cellular response to leukemia inhibitory factor / small-subunit processome / Nonhomologous End-Joining (NHEJ) / Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement / cellular response to gamma radiation / protein processing / double-strand break repair via nonhomologous end joining / double-strand break repair / double-stranded DNA binding / scaffold protein binding / secretory granule lumen / DNA recombination / transcription regulator complex / ficolin-1-rich granule lumen / damaged DNA binding / chromosome, telomeric region / aspartic-type endopeptidase activity / transcription cis-regulatory region binding / response to xenobiotic stimulus / ribonucleoprotein complex / innate immune response / negative regulation of DNA-templated transcription / DNA damage response / ubiquitin protein ligase binding / Neutrophil degranulation / protein-containing complex binding / nucleolus / positive regulation of DNA-templated transcription / positive regulation of transcription by RNA polymerase II / ATP hydrolysis activity / protein-containing complex / proteolysis / DNA binding / RNA binding / extracellular region / nucleoplasm / ATP binding / membrane / nucleus / plasma membrane / cytosol Similarity search - Function | |||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
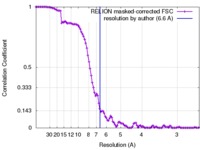
| Method | single particle reconstruction / cryo EM / Resolution: 6.6 Å | |||||||||||||||
 Authors Authors | Yin X / Liu M | |||||||||||||||
| Funding support |  China, 4 items China, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Cell Res / Year: 2017 Journal: Cell Res / Year: 2017Title: Cryo-EM structure of human DNA-PK holoenzyme. Authors: Xiaotong Yin / Mengjie Liu / Yuan Tian / Jiawei Wang / Yanhui Xu /  Abstract: DNA-dependent protein kinase (DNA-PK) is a serine/threonine protein kinase complex composed of a catalytic subunit (DNA-PKcs) and KU70/80 heterodimer bound to DNA. DNA-PK holoenzyme plays a critical ...DNA-dependent protein kinase (DNA-PK) is a serine/threonine protein kinase complex composed of a catalytic subunit (DNA-PKcs) and KU70/80 heterodimer bound to DNA. DNA-PK holoenzyme plays a critical role in non-homologous end joining (NHEJ), the major DNA repair pathway. Here, we determined cryo-electron microscopy structure of human DNA-PK holoenzyme at 6.6 Å resolution. In the complex structure, DNA-PKcs, KU70, KU80 and DNA duplex form a 650-kDa heterotetramer with 1:1:1:1 stoichiometry. The N-terminal α-solenoid (∼2 800 residues) of DNA-PKcs adopts a double-ring fold and connects the catalytic core domain of DNA-PKcs and KU70/80-DNA. DNA-PKcs and KU70/80 together form a DNA-binding tunnel, which cradles ∼30-bp DNA and prevents sliding inward of DNA-PKcs along with DNA duplex, suggesting a mechanism by which the broken DNA end is protected from unnecessary processing. Structural and biochemical analyses indicate that KU70/80 and DNA coordinately induce conformational changes of DNA-PKcs and allosterically stimulate its kinase activity. We propose a model for activation of DNA-PKcs in which allosteric signals are generated upon DNA-PK holoenzyme formation and transmitted to the kinase domain through N-terminal HEAT repeats and FAT domain of DNA-PKcs. Our studies suggest a mechanism for recognition and protection of broken DNA ends and provide a structural basis for understanding the activation of DNA-PKcs and DNA-PK-mediated NHEJ pathway. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6803.map.gz emd_6803.map.gz | 6.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6803-v30.xml emd-6803-v30.xml emd-6803.xml emd-6803.xml | 21.2 KB 21.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_6803_fsc.xml emd_6803_fsc.xml | 10.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_6803.png emd_6803.png | 116.9 KB | ||
| Filedesc metadata |  emd-6803.cif.gz emd-6803.cif.gz | 9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6803 http://ftp.pdbj.org/pub/emdb/structures/EMD-6803 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6803 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6803 | HTTPS FTP |
-Validation report
| Summary document |  emd_6803_validation.pdf.gz emd_6803_validation.pdf.gz | 382.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_6803_full_validation.pdf.gz emd_6803_full_validation.pdf.gz | 381.6 KB | Display | |
| Data in XML |  emd_6803_validation.xml.gz emd_6803_validation.xml.gz | 11.5 KB | Display | |
| Data in CIF |  emd_6803_validation.cif.gz emd_6803_validation.cif.gz | 15.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6803 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6803 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6803 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6803 | HTTPS FTP |
-Related structure data
| Related structure data |  5y3rMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6803.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6803.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.3 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
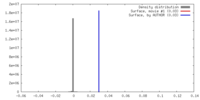
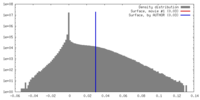
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : PRKDC-Helix
| Entire | Name: PRKDC-Helix |
|---|---|
| Components |
|
-Supramolecule #1: PRKDC-Helix
| Supramolecule | Name: PRKDC-Helix / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: X-ray repair cross-complementing protein 6
| Macromolecule | Name: X-ray repair cross-complementing protein 6 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO EC number: Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 57.619352 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GRDSLIFLVD ASKAMFESQS EDELTPFDMS IQCIQSVYIS KIISSDRDLL AVVFYGTEKD KNSVNFKNIY VLQELDNPGA KRILELDQF KGQQGQKRFQ DMMGHGSDYS LSEVLWVCAN LFSDVQFKMS HKRIMLFTNE DNPHGNDSAK ASRARTKAGD L RDTGIFLD ...String: GRDSLIFLVD ASKAMFESQS EDELTPFDMS IQCIQSVYIS KIISSDRDLL AVVFYGTEKD KNSVNFKNIY VLQELDNPGA KRILELDQF KGQQGQKRFQ DMMGHGSDYS LSEVLWVCAN LFSDVQFKMS HKRIMLFTNE DNPHGNDSAK ASRARTKAGD L RDTGIFLD LMHLKKPGGF DISLFYRDII SIAEDEDLRV HFEESSKLED LLRKVRAKET RKRALSRLKL KLNKDIVISV GI YNLVQKA LKPPPIKLYR ETNEPVKTKT RTFNTSTGGL LLPSDTKRSQ IYGSRQIILE KEETEELKRF DDPGLMLMGF KPL VLLKKH HYLRPSLFVY PEESLVIGSS TLFSALLIKC LEKEVAALCR YTPRRNIPPY FVALVPQEEE LDDQKIQVTP PGFQ LVFLP FADDKRKMPF TEKIMATPEQ VGKMKAIVEK LRFTYRSDSF ENPVLQQHFR NLEALALDLM EPEQAVDLTL PKVEA MNKR LGSLVDEFKE LVYPPDY UniProtKB: X-ray repair cross-complementing protein 6 |
-Macromolecule #2: X-ray repair cross-complementing protein 5
| Macromolecule | Name: X-ray repair cross-complementing protein 5 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO EC number: Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 60.972969 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: NKAAVVLCMD VGFTMSNSIP GIESPFEQAK KVITMFVQRQ VFAENKDEIA LVLFGTDGTD NPLSGGDQYQ NITVHRHLML PDFDLLEDI ESKIQPGSQQ ADFLDALIVS MDVIQHETIG KKFEKRHIEI FTDLSSRFSK SQLDIIIHSL KKCDISLQFF L PFSLGKED ...String: NKAAVVLCMD VGFTMSNSIP GIESPFEQAK KVITMFVQRQ VFAENKDEIA LVLFGTDGTD NPLSGGDQYQ NITVHRHLML PDFDLLEDI ESKIQPGSQQ ADFLDALIVS MDVIQHETIG KKFEKRHIEI FTDLSSRFSK SQLDIIIHSL KKCDISLQFF L PFSLGKED GSGDRGDGPF RLGGHGPSFP LKGITEQQKE GLEIVKMVMI SLEGEDGLDE IYSFSESLRK LCVFKKIERH SI HWPCRLT IGSNLSIRIA AYKSILQERV KKTWTVVDAK TLKKEDIQKE TVYCLNDDDE TEVLKEDIIQ GFRYGSDIVP FSK VDEEQM KYKSEGKCFS VLGFCKSSQV QRRFFMGNQV LKVFAARDDE AAAVALSSLI HALDDLDMVA IVRYAYDKRA NPQV GVAFP HIKHNYECLV YVQLPFMEDL RQYMFSSLKN SKKYAPTEAQ LNAVDALIDS MSLAKKDEKT DTLEDLFPTT KIPNP RFQR LFQCLLHRAL HPREPLPPIQ QHIWNMLNPP AEVTTKSQIP LSKIKTLFPL IE UniProtKB: X-ray repair cross-complementing protein 5 |
-Macromolecule #3: PRKDC-Helix
| Macromolecule | Name: PRKDC-Helix / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 1.084182 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AAAAAAAAAA AAAAA |
-Macromolecule #6: DNA-dependent protein kinase catalytic subunit
| Macromolecule | Name: DNA-dependent protein kinase catalytic subunit / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO / EC number: non-specific serine/threonine protein kinase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 468.885344 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: CSLLRLQETL SAADRCGAAL AGHQLIRGLG QECVLSSSPA VLALQTSLVF SRDFGLLVFV RKSLNSIEFR ECREEILKFL CIFLEKMGQ KIAPYSVEIK NTCTSVYTKD RAAKCKIPAL DLLIKLLQTF RSSRLMDEFK IGELFSKFYG ELALKKKIPD T VLEKVYEL ...String: CSLLRLQETL SAADRCGAAL AGHQLIRGLG QECVLSSSPA VLALQTSLVF SRDFGLLVFV RKSLNSIEFR ECREEILKFL CIFLEKMGQ KIAPYSVEIK NTCTSVYTKD RAAKCKIPAL DLLIKLLQTF RSSRLMDEFK IGELFSKFYG ELALKKKIPD T VLEKVYEL LGLLGEVHPS EMINNAENLF RAFLGELKTQ MTSAVREPKL PVLAGCLKGL SSLLCNFTKS MEEDPQTSRE IF NFVLKAI RPQIDLKRYA VPSAGLRLFA LHASQFSTCL LDNYVSLFEV LLKWCAHTNV ELKKAALSAL ESFLKQVSNM VAK NAEMHK NKLQYFMEQF YGIIRNVDSN NKELSIAIRG YGLFAGPCKV INAKDVDFMY VELIQRCKQM FLTQTDTGDD RVYQ MPSFL QSVASVLLYL DTVPEVYTPV LEHLVVMQID SFPQYSPKMQ LVCCRAIVKV FLALAAKGPV LRNCISTVVH QGLIR ICSK PVVLPKGPES ESEDHRASGE VRTGKWKVPT YKDYVDLFRH LLSSDQMMDS ILADEAFFSV NSSSESLNHL LYDEFV KSV LKIVEKLDLT LEIQTVGEQE NGDEAPGVWM IPTSDPAANL HPAKPKDFSA FINLVEFCRE ILPEKQAEFF EPWVYSF SY ELILQSTRLP LISGFYKLLS ITVRNAKKIK YFEGVSPKSL KHSPEDPEKY SCFALFVKFG KEVAVKMKQY KDELLASC L TFLLSLPHNI IELDVRAYVP ALQMAFKLGL SYTPLAEVGL NALEEWSIYI DRHVMQPYYK DILPCLDGYL KTSALSDET KNNWEVSALS RAAQKGFNKV VLKHLKKTKN LSSNEAISLE EIRIRVVQML GSLGGQINKN LLTVTSSDEM MKSYVAWDRE KRLSFAVPF REMKPVIFLD VFLPRVTELA LTASDRQTKV AACELLHSMV MFMLGKATQM PEGGQGAPPM YQLYKRTFPV L LRLACDVD QVTRQLYEPL VMQLIHWFTN NKKFESQDTV ALLEAILDGI VDPVDSTLRD FCGRCIREFL KWSIKQITPQ QQ EKSPVNT KSLFKRLYSL ALHPNAFKRL GASLAFNNIY REFREEESLV EQFVFEALVI YMESLALAHA DEKSLGTIQQ CCD AIDHLC RIIEKKHVSL NKAKKRRLPR GFPPSASLCL LDLVKWLLAH CGRPQTECRH KSIELFYKFV PLLPGNRSPN LWLK DVLKE EGVSFLINTF EGGGCGQPSG ILAQPTLLYL RGPFSLQATL CWLDLLLAAL ECYNTFIGER TVGALQVLGT EAQSS LLKA VAFFLESIAM HDIIAAEKCF GTGAAGNRTS PQEGERYNYS KCTVVVRIME FTTTLLNTSP EGWKLLKKDL CNTHLM RVL VQTLCEPASI GFNIGDVQVM AHLPDVCVNL MKALKMSPYK DILETHLREK ITAQSIEELC AVNLYGPDAQ VDRSRLA AV VSACKQLHRA GLLHNILPSQ STDLHHSVGT ELLSLVYKGI APGDERQCLP SLDLSCKQLA SGLLELAFAF GGLCERLV S LLLNPAVLST ASLGSSQGSV IHFSHGEYFY SLFSETINTE LLKNLDLAVL ELMQSSVDNT KMVSAVLNGM LDQSFRERA NQKHQGLKLA TTILQHWKKC DSWWAKDSPL ETKMAVLALL AKILQIDSSV SFNTSHGSFP EVFTTYISLL ADTKLDLHLK GQAVTLLPF FTSLTGGSLE ELRRVLEQLI VAHFPMQSRE FPPGTPRFNN YVDCMKKFLD ALELSQSPML LELMTEVLCR E QQHVMEEL FQSSFRRIAR RGSCVTQVGL LESVYEMFRK DDPRLSFTRQ SFVDRSLLTL LWHCSLDALR EFFSTIVVDA ID VLKSRFT KLNESTFDTQ ITKKMGYYKI LDVMYSRLPK DDVHAKESKI NQVFHGSCIT EGNELTKTLI KLCYDAFTEN MAG ENQLLE RRRLYHCAAY NCAISVICCV FNELKFYQGF LFSEKPEKNL LIFENLIDLK RRYNFPVEVE VPMERKKKYI EIRK EAREA ANGDSDGPSY MSSLSYLADS TLSEEMSQFD FSTGVQSYSY SSQDPRPATG RFRRREQRDP TVHDDVLELE MDELN RHEC MAPLTALVKH MHRSLGPPQG EEDSVPRDLP SWMKFLHGKL GNPIVPLNIR LFLAKLVINT EEVFRPYAKH WLSPLL QLA ASENNGGEGI HYMVVEIVAT ILSWTGLATP TGVPKDEVLA NRLLNFLMKH VFHPKRAVFR HNLEIIKTLV ECWKDCL SI PYRLIFEKFS GKDPNSKDNS VGIQLLGIVM ANDLPPYDPQ CGIQSSEYFQ ALVNNMSFVR YKEVYAAAAE VLGLILRY V MERKNILEES LCELVAKQLK QHQNTMEDKF IVCLNKVTKS FPPLADRFMN AVFFLLPKFH GVLKTLCLEV VLCRVEGMT ELYFQLKSKD FVQVMRHRDD ERQKVCLDII YKMMPKLKPV ELRELLNPVV EFVSHPSTTC REQMYNILMW IHDNYRDPES ETDNDSQEI FKLAKDVLIQ GLIDENPGLQ LIIRNFWSHE TRLPSNTLDR LLALNSLYSP KIEVHFLSLA TNFLLEMTSM S PDYPNPMF EHPLSECEFQ EYTIDSDWRF RSTVLTPMFV ETQASQGTLQ TRTQEGSLSA RWPVAGQIRA TQQQHDFTLT QT ADGRSSF DWLTGSSTDP LVDHTSPSSD SLLFAHKRSE RLQRAPLKSV GPDFGKKRLG LPGDEVDNKV KGAAGRTDLL RLR RRFMRD QEKLSLMYAR KGVAEQKREK EIKSELKMKQ DAQVVLYRSY RHGDLPDIQI KHSSLITPLQ AVAQRDPIIA KQLF SSLFS GILKEMDKFK TLSEKNNITQ KLLQDFNRFL NTTFSFFPPF VSCIQDISCQ HAALLSLDPA AVSAGCLASL QQPVG IRLL EEALLRLLPA ELPAKRVRGK ARLPPDVLRW VELAKLYRSI GEYDVLRGIF TSEIGTKQIT QSALLAEARS DYSEAA KQY DEALNKQDWV DGEPTEAEKD FWELASLDCY NHLAEWKSLE YCSTASIDSE NPPDLNKIWS EPFYQETYLP YMIRSKL KL LLQGEADQSL LTFIDKAMHG ELQKAILELH YSQELSLLYL LQDDVDRAKY YIQNGIQSFM QNYSSIDVLL HQSRLTKL Q SVQALTEIQE FISFISKQGN LSSQVPLKRL LNTWTNRYPD AKMDPMNIWD DIITNRCFFL SKIEEKLTPL PEDNSMNVD QDGDPSDRME VQEQEEDISS LIRSCKFSMK MKMIDSARKQ NNFSLAMKLL KELHKESKTR DDWLVSWVQS YCRLSHCRSR SQGCSEQVL TVLKTVSLLD ENNVSSYLSK NILAFRDQNI LLGTTYRIIA NALSSEPACL AEIEEDKARR ILELSGSSSE D SEKVIAGL YQRAFQHLSE AVQAAEEEAQ PPSWSCGPAA GVIDAYMTLA DFCDQQLRKE EENASVIDSA ELQAYPALVV EK MLKALKL NSNEARLKFP RLLQIIERYP EETLSLMTKE ISSVPCWQFI SWISHMVALL DKDQAVAVQH SVEEITDNYP QAI VYPFII SSESYSFKDT STGHKNKEFV ARIKSKLDQG GVIQDFINAL DQLSNPELLF KDWSNDVRAE LAKTPVNKKN IEKM YERMY AALGDPKAPG LGAFRRKFIQ TFGKEFDKHF GKGGSKLLRM KLSDFNDITN MLLLKMNKDS KPPGNLKECS PWMSD FKVE FLRNELEIPG QYDGRGKPLP EYHVRIAGFD ERVTVMASLR RPKRIIIRGH DEREHPFLVK GGEDLRQDQR VEQLFQ VMN GILAQDSACS QRALQLRTYS VVPMTSRLGL IEWLENTVTL KDLLLNTMSQ EEKAAYLSDP RAPPCEYKDW LTKMSGK HD VGAYMLMYKG ANRTETVTSF RKRESKVPAD LLKRAFVRMS TSPEAFLALR SHFASSHALI CISHWILGIG DRHLNNFM V AMETGGVIGI DFGHAFGSAT QFLPVPELMP FRLTRQFINL MLPMKETGLM YSIMVHALRA FRSDPGLLTN TMDVFVKEP SFDWKNFEQK MLKKGGSWIQ EINVAEKNWY PRQKICYAKR KLAGANPAVI TCDELLLGHE KAPAFRDYVA VARGSKDHNI RAQEPESGL SEETQVKCLM DQATDPNILG RTWEGWEPWM UniProtKB: Plasmepsin X |
-Macromolecule #4: DNA (34-MER)
| Macromolecule | Name: DNA (34-MER) / type: dna / ID: 4 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 10.502785 KDa |
| Sequence | String: (DT)(DA)(DA)(DA)(DA)(DA)(DC)(DT)(DA)(DT) (DT)(DA)(DT)(DT)(DA)(DT)(DG)(DG)(DT)(DA) (DT)(DT)(DA)(DT)(DG)(DG)(DC)(DC)(DT) (DT)(DG)(DG)(DG)(DC) |
-Macromolecule #5: DNA (36-MER)
| Macromolecule | Name: DNA (36-MER) / type: dna / ID: 5 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.042164 KDa |
| Sequence | String: (DC)(DA)(DG)(DC)(DT)(DA)(DA)(DT)(DG)(DG) (DC)(DC)(DA)(DT)(DA)(DA)(DT)(DA)(DC)(DC) (DA)(DT)(DA)(DA)(DT)(DA)(DA)(DT)(DA) (DG)(DT)(DT)(DT)(DT)(DT)(DA) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.6 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 40 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 1-32 / Number real images: 3496 / Average exposure time: 8.0 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.7 µm / Nominal defocus min: 1.7 µm / Nominal magnification: 18000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



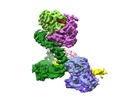


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)