+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6425 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Retinoschisin, back-to-back octameric rings | |||||||||
 Map data Map data | Retinoschisin double octamer rings | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | X-linked retinoschisis / discoidin domain / retina / macular degeneration / vision | |||||||||
| Function / homology |  Function and homology information Function and homology informationneuron to neuron synapse / retina layer formation / eye development / phosphatidylinositol-3,4-bisphosphate binding / phosphatidylserine binding / visual perception / photoreceptor inner segment / protein homooligomerization / cell adhesion / external side of plasma membrane ...neuron to neuron synapse / retina layer formation / eye development / phosphatidylinositol-3,4-bisphosphate binding / phosphatidylserine binding / visual perception / photoreceptor inner segment / protein homooligomerization / cell adhesion / external side of plasma membrane / protein-containing complex binding / protein-containing complex / extracellular space Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | Heymann JB / Tolun G / Vijayasarathy C / Huang R / Zeng Y / Li Y / Steven AC / Sieving PA | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2016 Journal: Proc Natl Acad Sci U S A / Year: 2016Title: Paired octamer rings of retinoschisin suggest a junctional model for cell-cell adhesion in the retina. Authors: Gökhan Tolun / Camasamudram Vijayasarathy / Rick Huang / Yong Zeng / Yan Li / Alasdair C Steven / Paul A Sieving / J Bernard Heymann /  Abstract: Retinoschisin (RS1) is involved in cell-cell junctions in the retina, but is unique among known cell-adhesion proteins in that it is a soluble secreted protein. Loss-of-function mutations in RS1 lead ...Retinoschisin (RS1) is involved in cell-cell junctions in the retina, but is unique among known cell-adhesion proteins in that it is a soluble secreted protein. Loss-of-function mutations in RS1 lead to early vision impairment in young males, called X-linked retinoschisis. The disease is characterized by separation of inner retinal layers and disruption of synaptic signaling. Using cryo-electron microscopy, we report the structure at 4.1 Å, revealing double octamer rings not observed before. Each subunit is composed of a discoidin domain and a small N-terminal (RS1) domain. The RS1 domains occupy the centers of the rings, but are not required for ring formation and are less clearly defined, suggesting mobility. We determined the structure of the discoidin rings, consistent with known intramolecular and intermolecular disulfides. The interfaces internal to and between rings feature residues implicated in X-linked retinoschisis, indicating the importance of correct assembly. Based on this structure, we propose that RS1 couples neighboring membranes together through octamer-octamer contacts, perhaps modulated by interactions with other membrane components. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6425.map.gz emd_6425.map.gz | 14.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6425-v30.xml emd-6425-v30.xml emd-6425.xml emd-6425.xml | 11.1 KB 11.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_6425_fsc.xml emd_6425_fsc.xml | 5.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_6425.png emd_6425.png | 321.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6425 http://ftp.pdbj.org/pub/emdb/structures/EMD-6425 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6425 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6425 | HTTPS FTP |
-Validation report
| Summary document |  emd_6425_validation.pdf.gz emd_6425_validation.pdf.gz | 399.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_6425_full_validation.pdf.gz emd_6425_full_validation.pdf.gz | 399.4 KB | Display | |
| Data in XML |  emd_6425_validation.xml.gz emd_6425_validation.xml.gz | 9.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6425 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6425 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6425 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6425 | HTTPS FTP |
-Related structure data
| Related structure data |  3jd6MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6425.map.gz / Format: CCP4 / Size: 15.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6425.map.gz / Format: CCP4 / Size: 15.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Retinoschisin double octamer rings | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.03 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
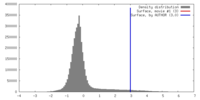
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Mature retinoschisin with a C-terminal hexahistidine tag
| Entire | Name: Mature retinoschisin with a C-terminal hexahistidine tag |
|---|---|
| Components |
|
-Supramolecule #1000: Mature retinoschisin with a C-terminal hexahistidine tag
| Supramolecule | Name: Mature retinoschisin with a C-terminal hexahistidine tag type: sample / ID: 1000 / Oligomeric state: 16-mer / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 400 KDa / Theoretical: 382 KDa / Method: Blue native gel electrophoresis |
-Macromolecule #1: Retinoschisin 1
| Macromolecule | Name: Retinoschisin 1 / type: protein_or_peptide / ID: 1 / Name.synonym: RS1 Details: The mature RS1 protein (no signal sequence) with a C-terminal six-histidine tag Number of copies: 16 / Oligomeric state: 16-mer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human / Tissue: Retina / Cell: Photoreceptors and bipolar cells / Organelle: Extracellular space / Location in cell: Forming a junction between cells Homo sapiens (human) / synonym: Human / Tissue: Retina / Cell: Photoreceptors and bipolar cells / Organelle: Extracellular space / Location in cell: Forming a junction between cells |
| Molecular weight | Theoretical: 24 KDa |
| Recombinant expression | Organism:  |
| Sequence | UniProtKB: Retinoschisin |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 20 mM Tris-HCl, 150 mM NaCl |
| Grid | Details: C-flat holey carbon grids (2.0 um holes, 2.0 um spaces) |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Instrument: LEICA EM GP / Details: Plunged at room temperature. / Method: Blotted from the copper side. |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Date | Jun 11, 2014 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 (4k x 4k) / Digitization - Sampling interval: 5 µm / Number real images: 19 / Average electron dose: 30 e/Å2 Details: Each image is the average of 10 frames taken over 3 seconds. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal magnification: 39000 |
| Sample stage | Specimen holder model: OTHER |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)