+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Caenorhabditis elegans ACR-23 in apo state | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ion channel / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationHighly calcium permeable postsynaptic nicotinic acetylcholine receptors / Neurotransmitter receptors and postsynaptic signal transmission / excitatory extracellular ligand-gated monoatomic ion channel activity / transmembrane transporter complex / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / electron transport chain / transmembrane signaling receptor activity / postsynapse / periplasmic space / electron transfer activity ...Highly calcium permeable postsynaptic nicotinic acetylcholine receptors / Neurotransmitter receptors and postsynaptic signal transmission / excitatory extracellular ligand-gated monoatomic ion channel activity / transmembrane transporter complex / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / electron transport chain / transmembrane signaling receptor activity / postsynapse / periplasmic space / electron transfer activity / neuron projection / iron ion binding / heme binding / synapse / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.61 Å | |||||||||
 Authors Authors | Chen QF / Liu FL / Li TY / Gong HH / Guo F / Liu S | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structural basis of acetylcholine receptor like-23 (ACR-23) activation by anthelmintics monepantel and betaine Authors: Liu FL / Li TY / Gong HH / Guo F / Liu S / Chen QF | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_60064.map.gz emd_60064.map.gz | 306.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-60064-v30.xml emd-60064-v30.xml emd-60064.xml emd-60064.xml | 15.3 KB 15.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_60064.png emd_60064.png | 71.1 KB | ||
| Filedesc metadata |  emd-60064.cif.gz emd-60064.cif.gz | 5.8 KB | ||
| Others |  emd_60064_half_map_1.map.gz emd_60064_half_map_1.map.gz emd_60064_half_map_2.map.gz emd_60064_half_map_2.map.gz | 301.4 MB 301.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-60064 http://ftp.pdbj.org/pub/emdb/structures/EMD-60064 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60064 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-60064 | HTTPS FTP |
-Validation report
| Summary document |  emd_60064_validation.pdf.gz emd_60064_validation.pdf.gz | 808.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_60064_full_validation.pdf.gz emd_60064_full_validation.pdf.gz | 807.7 KB | Display | |
| Data in XML |  emd_60064_validation.xml.gz emd_60064_validation.xml.gz | 17.2 KB | Display | |
| Data in CIF |  emd_60064_validation.cif.gz emd_60064_validation.cif.gz | 20.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-60064 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-60064 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-60064 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-60064 | HTTPS FTP |
-Related structure data
| Related structure data |  8zflMC  8zfkC  8zfmC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_60064.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_60064.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.856 Å | ||||||||||||||||||||||||||||||||||||
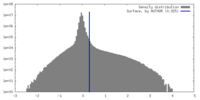
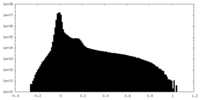
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_60064_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
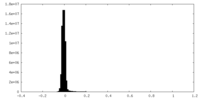
| Density Histograms |
-Half map: #1
| File | emd_60064_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
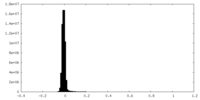
| Density Histograms |
- Sample components
Sample components
-Entire : acetylcholine receptor like-23(ACR-23)
| Entire | Name: acetylcholine receptor like-23(ACR-23) |
|---|---|
| Components |
|
-Supramolecule #1: acetylcholine receptor like-23(ACR-23)
| Supramolecule | Name: acetylcholine receptor like-23(ACR-23) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 67 KDa |
-Macromolecule #1: Betaine receptor acr-23,Soluble cytochrome b562
| Macromolecule | Name: Betaine receptor acr-23,Soluble cytochrome b562 / type: protein_or_peptide / ID: 1 Details: Residues 387-500 correspond to the bril expression tag. Residues 599-603 and 610-612 correspond to flexible linkers, residues 604-609 correspond to the thrombin cleavage site, and residues ...Details: Residues 387-500 correspond to the bril expression tag. Residues 599-603 and 610-612 correspond to flexible linkers, residues 604-609 correspond to the thrombin cleavage site, and residues 613-620 correspond to the strep ll tag. Number of copies: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 70.582281 KDa |
| Recombinant expression | Organism:  Spodoptera aff. frugiperda 1 BOLD-2017 (butterflies/moths) Spodoptera aff. frugiperda 1 BOLD-2017 (butterflies/moths) |
| Sequence | String: MHRIYTFLIF ISQLALGLSN NPDIPIQYEL ANNIMENYQK GLIPKVRKGS PINVTLSLQL YQIIQVNEPQ QYLLLNAWAV ERWVDQMLG WDPSEFDNET EIMARHDDIW LPDTTLYNSL EMDDSASKKL THVKLTTLGK NQGAMVELLY PTIYKISCLL N LKYFPFDT ...String: MHRIYTFLIF ISQLALGLSN NPDIPIQYEL ANNIMENYQK GLIPKVRKGS PINVTLSLQL YQIIQVNEPQ QYLLLNAWAV ERWVDQMLG WDPSEFDNET EIMARHDDIW LPDTTLYNSL EMDDSASKKL THVKLTTLGK NQGAMVELLY PTIYKISCLL N LKYFPFDT QTCRMTFGSW SFDNSLIDYF PRTFTNGPIG LANFLENDAW SVLGTKVNRE EKKYTCCPVN YTLLHYDVVI QR KPLYYVL NLIAPTAVIT FISIIGFFTS SSVHDLRQEK ITLGITTLLS MSIMIFMVSD KMPSTSTCVP LIALFYTLMI TII SVGTLA ASSVIFVQKL GSIGNPPASK TMKWTHRIAP FVLIQMPLVM KQAYAKRAKE EKHRKRMSRK MAEAGAMADL EDNW ETLND NLKVIEKADN AAQVKDALTK MRAAALDAQK ATPPKLEDKS PDSPEMKDFR HGFDILVGQI DDALKLANEG KVKEA QAAA EQLKTTRNAY IQKYLSQTSE TFAAPMDTSF TESLHIPELN RVASSNSIQS VLKPTEIQLT PYCTRNIVEL EWDWVA AVL ERVFLIFFTI CFLFSAIGIN LYGWYIWYTE NHFLFLEGTK LVPRGSSSGW SHPQFEK UniProtKB: Betaine receptor acr-23, Soluble cytochrome b562, Betaine receptor acr-23 |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 5 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 4.00 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | #0 - Image recording ID: 1 / #0 - Film or detector model: GATAN K2 BASE (4k x 4k) / #0 - Average electron dose: 50.0 e/Å2 / #1 - Image recording ID: 2 / #1 - Film or detector model: GATAN K2 BASE (4k x 4k) / #1 - Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing #1
Image processing #1
- Image processing #2
Image processing #2
| Image processing ID | 2 |
|---|---|
| Image recording ID | 2 |
| Startup model | Type of model: NONE |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)