| Entry | Database: PDB / ID: 5ugu
|
|---|
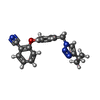
| Title | Crystal structure of M. tuberculosis InhA inhibited by PT506 |
|---|
 Components Components | Enoyl-[acyl-carrier-protein] reductase [NADH] |
|---|
 Keywords Keywords | OXIDOREDUCTASE / bacterial enoyl-ACP reductase / diphenylether / residence time |
|---|
| Function / homology |  Function and homology information Function and homology information
trans-2-enoyl-CoA reductase (NADH) activity / mycolic acid biosynthetic process / fatty acid elongation / enoyl-[acyl-carrier-protein] reductase (NADH) / enoyl-[acyl-carrier-protein] reductase (NADH) activity / NAD+ binding / peptidoglycan-based cell wall / fatty acid binding / response to antibiotic / plasma membraneSimilarity search - Function : / Enoyl-[acyl-carrier-protein] reductase (NADH) / Enoyl-(Acyl carrier protein) reductase / NAD(P)-binding Rossmann-like Domain / NAD(P)-binding domain superfamily / Rossmann fold / 3-Layer(aba) Sandwich / Alpha BetaSimilarity search - Domain/homology |
|---|
| Biological species |   Mycobacterium tuberculosis (bacteria) Mycobacterium tuberculosis (bacteria) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.95 Å MOLECULAR REPLACEMENT / Resolution: 1.95 Å |
|---|
 Authors Authors | Eltschkner, S. / Pschibul, A. / Spagnuolo, L.A. / Yu, W. / Tonge, P.J. / Kisker, C. |
|---|
| Funding support |  Germany, 1items Germany, 1items | Organization | Grant number | Country |
|---|
| German Research Foundation | SFB 630 |  Germany Germany |
|
|---|
 Citation Citation |  Journal: J. Am. Chem. Soc. / Year: 2017 Journal: J. Am. Chem. Soc. / Year: 2017
Title: Evaluating the Contribution of Transition-State Destabilization to Changes in the Residence Time of Triazole-Based InhA Inhibitors.
Authors: Spagnuolo, L.A. / Eltschkner, S. / Yu, W. / Daryaee, F. / Davoodi, S. / Knudson, S.E. / Allen, E.K. / Merino, J. / Pschibul, A. / Moree, B. / Thivalapill, N. / Truglio, J.J. / Salafsky, J. / ...Authors: Spagnuolo, L.A. / Eltschkner, S. / Yu, W. / Daryaee, F. / Davoodi, S. / Knudson, S.E. / Allen, E.K. / Merino, J. / Pschibul, A. / Moree, B. / Thivalapill, N. / Truglio, J.J. / Salafsky, J. / Slayden, R.A. / Kisker, C. / Tonge, P.J. |
|---|
| History | | Deposition | Jan 10, 2017 | Deposition site: RCSB / Processing site: PDBE |
|---|
| Revision 1.0 | Feb 15, 2017 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Mar 22, 2017 | Group: Database references |
|---|
| Revision 1.2 | Sep 6, 2017 | Group: Author supporting evidence / Category: pdbx_audit_support / Item: _pdbx_audit_support.funding_organization |
|---|
| Revision 1.3 | Jan 17, 2024 | Group: Data collection / Database references / Refinement description
Category: chem_comp_atom / chem_comp_bond ...chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model
Item: _database_2.pdbx_DOI / _database_2.pdbx_database_accession |
|---|
|
|---|
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.95 Å
MOLECULAR REPLACEMENT / Resolution: 1.95 Å  Authors
Authors Germany, 1items
Germany, 1items  Citation
Citation Journal: J. Am. Chem. Soc. / Year: 2017
Journal: J. Am. Chem. Soc. / Year: 2017 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 5ugu.cif.gz
5ugu.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb5ugu.ent.gz
pdb5ugu.ent.gz PDB format
PDB format 5ugu.json.gz
5ugu.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads 5ugu_validation.pdf.gz
5ugu_validation.pdf.gz wwPDB validaton report
wwPDB validaton report 5ugu_full_validation.pdf.gz
5ugu_full_validation.pdf.gz 5ugu_validation.xml.gz
5ugu_validation.xml.gz 5ugu_validation.cif.gz
5ugu_validation.cif.gz https://data.pdbj.org/pub/pdb/validation_reports/ug/5ugu
https://data.pdbj.org/pub/pdb/validation_reports/ug/5ugu ftp://data.pdbj.org/pub/pdb/validation_reports/ug/5ugu
ftp://data.pdbj.org/pub/pdb/validation_reports/ug/5ugu





 Links
Links Assembly
Assembly

 Components
Components

 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  BESSY
BESSY  / Beamline: 14.1 / Wavelength: 0.918409 Å
/ Beamline: 14.1 / Wavelength: 0.918409 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller













 PDBj
PDBj