[English] 日本語
 Yorodumi
Yorodumi- EMDB-50182: cryo-EM structure of human LST2 bound to human mTOR complex 1, fo... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | cryo-EM structure of human LST2 bound to human mTOR complex 1, focused on RAPTOR | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | MTOR / MTORC1 / LST2 / ZFYVE28 / EGFR / TOS / SIGNALING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of epidermal growth factor-activated receptor activity / positive regulation of pentose-phosphate shunt / TORC1 complex / positive regulation of odontoblast differentiation / TORC1 signaling / negative regulation of epidermal growth factor receptor signaling pathway / cellular response to L-leucine / MTOR signalling / Amino acids regulate mTORC1 / Energy dependent regulation of mTOR by LKB1-AMPK ...negative regulation of epidermal growth factor-activated receptor activity / positive regulation of pentose-phosphate shunt / TORC1 complex / positive regulation of odontoblast differentiation / TORC1 signaling / negative regulation of epidermal growth factor receptor signaling pathway / cellular response to L-leucine / MTOR signalling / Amino acids regulate mTORC1 / Energy dependent regulation of mTOR by LKB1-AMPK / phosphatidylinositol-3-phosphate binding / protein serine/threonine kinase inhibitor activity / positive regulation of osteoclast differentiation / cellular response to osmotic stress / enzyme-substrate adaptor activity / positive regulation of transcription by RNA polymerase III / regulation of cell size / positive regulation of G1/S transition of mitotic cell cycle / Macroautophagy / protein kinase activator activity / mTORC1-mediated signalling / social behavior / HSF1-dependent transactivation / TOR signaling / positive regulation of TOR signaling / cellular response to nutrient levels / positive regulation of lipid biosynthetic process / 14-3-3 protein binding / Regulation of PTEN gene transcription / positive regulation of endothelial cell proliferation / cellular response to starvation / positive regulation of glycolytic process / negative regulation of autophagy / cellular response to amino acid stimulus / positive regulation of peptidyl-threonine phosphorylation / regulation of autophagy / regulation of cell growth / TP53 Regulates Metabolic Genes / cellular response to glucose stimulus / small GTPase binding / cytoplasmic stress granule / positive regulation of peptidyl-serine phosphorylation / early endosome membrane / cellular response to hypoxia / positive regulation of cell growth / protein-macromolecule adaptor activity / lysosome / response to xenobiotic stimulus / lysosomal membrane / neuronal cell body / DNA damage response / dendrite / protein-containing complex binding / protein kinase binding / nucleoplasm / metal ion binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
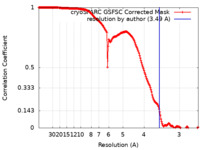
| Method | single particle reconstruction / cryo EM / Resolution: 3.49 Å | |||||||||
 Authors Authors | Craigie LM / Maier T | |||||||||
| Funding support |  Switzerland, European Union, 2 items Switzerland, European Union, 2 items
| |||||||||
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 2024 Journal: Proc.Natl.Acad.Sci.USA / Year: 2024Title: mTORC1 phosphorylates and stabilizes LST2 to negatively regulate EGFR Authors: Battaglioni S / Craigie LM / Filippini S / Maier T / Hall MN | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_50182.map.gz emd_50182.map.gz | 484 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-50182-v30.xml emd-50182-v30.xml emd-50182.xml emd-50182.xml | 18.4 KB 18.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_50182_fsc.xml emd_50182_fsc.xml | 16.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_50182.png emd_50182.png | 76.3 KB | ||
| Masks |  emd_50182_msk_1.map emd_50182_msk_1.map | 512 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-50182.cif.gz emd-50182.cif.gz | 7.1 KB | ||
| Others |  emd_50182_half_map_1.map.gz emd_50182_half_map_1.map.gz emd_50182_half_map_2.map.gz emd_50182_half_map_2.map.gz | 474.6 MB 474.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-50182 http://ftp.pdbj.org/pub/emdb/structures/EMD-50182 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-50182 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-50182 | HTTPS FTP |
-Validation report
| Summary document |  emd_50182_validation.pdf.gz emd_50182_validation.pdf.gz | 884.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_50182_full_validation.pdf.gz emd_50182_full_validation.pdf.gz | 884.1 KB | Display | |
| Data in XML |  emd_50182_validation.xml.gz emd_50182_validation.xml.gz | 25.6 KB | Display | |
| Data in CIF |  emd_50182_validation.cif.gz emd_50182_validation.cif.gz | 33.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-50182 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-50182 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-50182 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-50182 | HTTPS FTP |
-Related structure data
| Related structure data |  9f43MC  9f42C  9f44C  9f45C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_50182.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_50182.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.31 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_50182_msk_1.map emd_50182_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_50182_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_50182_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : mTORC1 in complex with LST2
| Entire | Name: mTORC1 in complex with LST2 |
|---|---|
| Components |
|
-Supramolecule #1: mTORC1 in complex with LST2
| Supramolecule | Name: mTORC1 in complex with LST2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Regulatory-associated protein of mTOR
| Macromolecule | Name: Regulatory-associated protein of mTOR / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 152.764656 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: HHHHHHHHHH EQKLISEEDL DYKDDDDKME SEMLQSPLLG LGEEDEADLT DWNLPLAFMK KRHCEKIEGS KSLAQSWRMK DRMKTVSVA LVLCLNVGVD PPDVVKTTPC ARLECWIDPL SMGPQKALET IGANLQKQYE NWQPRARYKQ SLDPTVDEVK K LCTSLRRN ...String: HHHHHHHHHH EQKLISEEDL DYKDDDDKME SEMLQSPLLG LGEEDEADLT DWNLPLAFMK KRHCEKIEGS KSLAQSWRMK DRMKTVSVA LVLCLNVGVD PPDVVKTTPC ARLECWIDPL SMGPQKALET IGANLQKQYE NWQPRARYKQ SLDPTVDEVK K LCTSLRRN AKEERVLFHY NGHGVPRPTV NGEVWVFNKN YTQYIPLSIY DLQTWMGSPS IFVYDCSNAG LIVKSFKQFA LQ REQELEV AAINPNHPLA QMPLPPSMKN CIQLAACEAT ELLPMIPDLP ADLFTSCLTT PIKIALRWFC MQKCVSLVPG VTL DLIEKI PGRLNDRRTP LGELNWIFTA ITDTIAWNVL PRDLFQKLFR QDLLVASLFR NFLLAERIMR SYNCTPVSSP RLPP TYMHA MWQAWDLAVD ICLSQLPTII EEGTAFRHSP FFAEQLTAFQ VWLTMGVENR NPPEQLPIVL QVLLSQVHRL RALDL LGRF LDLGPWAVSL ALSVGIFPYV LKLLQSSARE LRPLLVFIWA KILAVDSSCQ ADLVKDNGHK YFLSVLADPY MPAEHR TMT AFILAVIVNS YHTGQEACLQ GNLIAICLEQ LNDPHPLLRQ WVAICLGRIW QNFDSARWCG VRDSAHEKLY SLLSDPI PE VRCAAVFALG TFVGNSAERT DHSTTIDHNV AMMLAQLVSD GSPMVRKELV VALSHLVVQY ESNFCTVALQ FIEEEKNY A LPSPATTEGG SLTPVRDSPC TPRLRSVSSY GNIRAVATAR SLNKSLQNLS LTEESGGAVA FSPGNLSTSS SASSTLGSP ENEEHILSFE TIDKMRRASS YSSLNSLIGV SFNSVYTQIW RVLLHLAADP YPEVSDVAMK VLNSIAYKAT VNARPQRVLD TSSLTQSAP ASPTNKGVHI HQAGGSPPAS STSSSSLTND VAKQPVSRDL PSGRPGTTGP AGAQYTPHSH QFPRTRKMFD K GPEQTADD ADDAAGHKSF ISATVQTGFC DWSARYFAQP VMKIPEEHDL ESQIRKEREW RFLRNSRVRR QAQQVIQKGI TR LDDQIFL NRNPGVPSVV KFHPFTPCIA VADKDSICFW DWEKGEKLDY FHNGNPRYTR VTAMEYLNGQ DCSLLLTATD DGA IRVWKN FADLEKNPEM VTAWQGLSDM LPTTRGAGMV VDWEQETGLL MSSGDVRIVR IWDTDREMKV QDIPTGADSC VTSL SCDSH RSLIVAGLGD GSIRVYDRRM ALSECRVMTY REHTAWVVKA SLQKRPDGHI VSVSVNGDVR IFDPRMPESV NVLQI VKGL TALDIHPQAD LIACGSVNQF TAIYNSSGEL INNIKYYDGF MGQRVGAISC LAFHPHWPHL AVGSNDYYIS VYSVEK RVR UniProtKB: Regulatory-associated protein of mTOR |
-Macromolecule #2: Lateral signaling target protein 2 homolog
| Macromolecule | Name: Lateral signaling target protein 2 homolog / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 96.604477 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MMNRFRKWLY KPKRSDPQLL ARFYYADEEL NQVAAELDSL DGRKDPQRCT LLVSQFRSCQ DNVLNIINQI MDECIPQDRA PRDFCVKFP EEIRHDNLAG QLWFGAECLA AGSIIMNREL ESMAMRPLAK ELTRSLEDVR GALRDQALRD LNTYTEKMRE A LRHFDVLF ...String: MMNRFRKWLY KPKRSDPQLL ARFYYADEEL NQVAAELDSL DGRKDPQRCT LLVSQFRSCQ DNVLNIINQI MDECIPQDRA PRDFCVKFP EEIRHDNLAG QLWFGAECLA AGSIIMNREL ESMAMRPLAK ELTRSLEDVR GALRDQALRD LNTYTEKMRE A LRHFDVLF AEFELSYVSA MVPVKSPREY YVQQEVIVLF CETVERALDF GYLTQDMIDD YEPALMFSIP RLAIVCGLVV YA DGPLNLD RKVEDMSELF RPFHTLLRKI RDLLQTLTEE ELHTLERNLC ISQDVEFPIR ADVQGPAALA PALSAPLPPE GPL SAKAKD PDAELACSMQ YDDQELEQLS RMVHRAGDEM SSLLSPPIAC QSPAHRPGAE GSPGGEASPG RPRLRSGSDE EERV FFMDD VEGTAEALAR PESPAGPFGW AGSTWADPQE KGQGGPGGAA GISLPASEKE EDLSNNNLEA EGTDGASLAG TSSCS CLDS RLHLDGWEVG ADDAETAEMI AHRTGGMKLS ATVIFNPKSP TSLDSAVATQ EAASEPVAEG MDGGPHKLST GATNCL LHS CVCCGSCGDS REDVVERLRE KCSPGGVIGA SYAAGLAKAS DRAPERQEEA PPPSEDASNG REPKAPTSDK CLPHTSG SQ VDTASGLQGE AGVAGQQEPE ARELHAGSPS AHEAPQALSG SSSSTAGSCS SDKMGPEAAP AATHAAPQAT REKIRSRF H GSHDLIHRLF VCISGVADQL QTNYASDLRS ILKTLFEVMA TKPETDDKEK LRKVTQTLRS AALEDCALCQ ETLSSSELA AKTRDGDFED PPEWVPDEAC GFCTACKAPF TVIRRKHHCR SCGKIFCSRC SSHSAPLPRY GQVKPVRVCT HCYMFHVTPF YSDKAGL UniProtKB: Lateral signaling target protein 2 homolog |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil R2/1 / Material: GOLD / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 281 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 4719 / Average electron dose: 43.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z
Z Y
Y X
X