+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4udv | ||||||
|---|---|---|---|---|---|---|---|
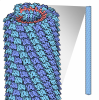
| Title | Cryo-EM structure of TMV at 3.35 A resolution | ||||||
 Components Components |
| ||||||
 Keywords Keywords | VIRAL PROTEIN / DIRECT ELECTRON DETECTORS / SINGLE PARTICLE HELICAL RECONSTRUCTION / HIGH RESOLUTION | ||||||
| Function / homology |  Function and homology information Function and homology informationhelical viral capsid / structural molecule activity / identical protein binding Similarity search - Function | ||||||
| Biological species |   TOBACCO MOSAIC VIRUS TOBACCO MOSAIC VIRUS | ||||||
| Method | ELECTRON MICROSCOPY / helical reconstruction / cryo EM / Resolution: 3.35 Å | ||||||
 Authors Authors | Fromm, S.A. / Bharat, T.A.M. / Jakobi, A.J. / Hagen, W.J.H. / Sachse, C. | ||||||
 Citation Citation |  Journal: J Struct Biol / Year: 2015 Journal: J Struct Biol / Year: 2015Title: Seeing tobacco mosaic virus through direct electron detectors. Authors: Simon A Fromm / Tanmay A M Bharat / Arjen J Jakobi / Wim J H Hagen / Carsten Sachse /   Abstract: With the introduction of direct electron detectors (DED) to the field of electron cryo-microscopy, a wave of atomic-resolution structures has become available. As the new detectors still require ...With the introduction of direct electron detectors (DED) to the field of electron cryo-microscopy, a wave of atomic-resolution structures has become available. As the new detectors still require comparative characterization, we have used tobacco mosaic virus (TMV) as a test specimen to study the quality of 3D image reconstructions from data recorded on the two direct electron detector cameras, K2 Summit and Falcon II. Using DED movie frames, we explored related image-processing aspects and compared the performance of micrograph-based and segment-based motion correction approaches. In addition, we investigated the effect of dose deposition on the atomic-resolution structure of TMV and show that radiation damage affects negative carboxyl chains first in a side-chain specific manner. Finally, using 450,000 asymmetric units and limiting the effects of radiation damage, we determined a high-resolution cryo-EM map at 3.35Å resolution. Here, we provide a comparative case study of highly ordered TMV recorded on different direct electron detectors to establish recording and processing conditions that enable structure determination up to 3.2Å in resolution using cryo-EM. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4udv.cif.gz 4udv.cif.gz | 40.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4udv.ent.gz pdb4udv.ent.gz | 26.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4udv.json.gz 4udv.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ud/4udv https://data.pdbj.org/pub/pdb/validation_reports/ud/4udv ftp://data.pdbj.org/pub/pdb/validation_reports/ud/4udv ftp://data.pdbj.org/pub/pdb/validation_reports/ud/4udv | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2842MC  2833C  2834C  2835C  2836C  2837C  2838C  2839C  2840C  2841C M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10022 (Title: Tobacco Mosaic Virus Falcon II dataset including manually boxed helix coordinates EMPIAR-10022 (Title: Tobacco Mosaic Virus Falcon II dataset including manually boxed helix coordinatesData size: 47.7 Data #1: Tobacco Mosaic Virus micrographs [micrographs - multiframe]) |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 | x 49
|
| 2 |
|
| 3 | 
|
| Symmetry | Helical symmetry: (Circular symmetry: 1 / Dyad axis: no / N subunits divisor: 1 / Num. of operations: 49 / Rise per n subunits: 1.408 Å / Rotation per n subunits: 22.03 °) |
- Components
Components
| #1: Protein | Mass: 17505.426 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   TOBACCO MOSAIC VIRUS / Strain: VULGARE / References: UniProt: P69687 TOBACCO MOSAIC VIRUS / Strain: VULGARE / References: UniProt: P69687 |
|---|---|
| #2: RNA chain | Mass: 958.660 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   TOBACCO MOSAIC VIRUS / Strain: VULGARE TOBACCO MOSAIC VIRUS / Strain: VULGARE |
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: FILAMENT / 3D reconstruction method: helical reconstruction |
- Sample preparation
Sample preparation
| Component | Name: TOBACCO MOSAIC VIRUS / Type: VIRUS |
|---|---|
| Buffer solution | Name: 50 MM NAPO4 / pH: 7 / Details: 50 MM NAPO4 |
| Specimen | Conc.: 11 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Specimen support | Details: HOLEY CARBON |
| Vitrification | Instrument: FEI VITROBOT MARK III / Cryogen name: ETHANE Details: CRYOGEN - ETHANE, HUMIDITY - 90 PERCENT, TEMPERATURE - 95 K, INSTRUMENT - FEI VITROBOT MARK III PROCEDURE - BLOT FOR 8 SECONDS WITH AN OFFSET OF -2 MM CA 30-45 SECONDS AFTER SAMPLE APPLICATION |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS / Date: Oct 16, 2013 Details: NANOPROBE MODE, DOSE RATE CA 41 E- PX S ON THE CAMERA LEVEL, FULLY AUTOMATED ACQUISITION USING FEI EPU SOFTWARE |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 75000 X / Calibrated magnification: 131827 X / Nominal defocus max: 4500 nm / Nominal defocus min: 1000 nm / Cs: 2.7 mm |
| Image recording | Electron dose: 30.7 e/Å2 / Film or detector model: FEI FALCON II (4k x 4k) |
| Image scans | Num. digital images: 109 |
- Processing
Processing
| EM software | Name: SPRING / Category: 3D reconstruction | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CTF correction | Details: CTFTILT (SPECIFIC FOR EACH SEGMENT) | ||||||||||||
| 3D reconstruction | Method: PROJECTION MATCHING / Resolution: 3.35 Å / Num. of particles: 450000 / Nominal pixel size: 1.062 Å / Actual pixel size: 1.062 Å / Magnification calibration: LAYER LINE CORRELATION Details: SUBMISSION BASED ON EXPERIMENTAL DATA FROM EMDB EMD-2842. (DEPOSITION ID: 12988). Symmetry type: HELICAL | ||||||||||||
| Atomic model building | B value: 90 / Protocol: OTHER / Space: REAL / Target criteria: REAL SPACE CORRELATION Details: METHOD--LOCAL AND GLOBAL CORRELATION REFINEMENT PROTOCOL--FIBRE DIFFRACTION | ||||||||||||
| Refinement | Highest resolution: 3.35 Å | ||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 3.35 Å
|
 Movie
Movie Controller
Controller









 PDBj
PDBj
