+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4902 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of St1Cas9-sgRNA-tDNA59-ntPAM complex. | |||||||||
 Map data Map data | Postprocessed 3D reconstruction of St1Cas9%u2022sgRNA%u2022tDNA59-ntPAM complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CRISPR-Cas9 / anti-CRISPR protein / bacteriophages / Streptococcus thermophilus Cas9 / St1Cas9 / hydrolase | |||||||||
| Function / homology |  Function and homology information Function and homology informationmaintenance of CRISPR repeat elements / endonuclease activity / defense response to virus / Hydrolases; Acting on ester bonds / hydrolase activity / DNA binding / RNA binding / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Streptococcus thermophilus DGCC 7710 (bacteria) / Streptococcus thermophilus DGCC 7710 (bacteria) /  Streptococcus thermophilus (bacteria) / Streptococcus thermophilus (bacteria) /  Streptococcus phage D1811 (virus) / Streptococcus phage D1811 (virus) /  Streptococcus phage 2972 (virus) / Streptococcus phage 2972 (virus) /  Streptococcus Phage 2972 (virus) Streptococcus Phage 2972 (virus) | |||||||||
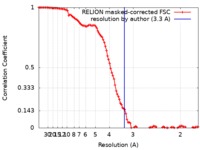
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Swuec P | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2019 Journal: Mol Cell / Year: 2019Title: Cas9 Allosteric Inhibition by the Anti-CRISPR Protein AcrIIA6. Authors: Olivier Fuchsbauer / Paolo Swuec / Claire Zimberger / Béatrice Amigues / Sébastien Levesque / Daniel Agudelo / Alexis Duringer / Antonio Chaves-Sanjuan / Silvia Spinelli / Geneviève M ...Authors: Olivier Fuchsbauer / Paolo Swuec / Claire Zimberger / Béatrice Amigues / Sébastien Levesque / Daniel Agudelo / Alexis Duringer / Antonio Chaves-Sanjuan / Silvia Spinelli / Geneviève M Rousseau / Minja Velimirovic / Martino Bolognesi / Alain Roussel / Christian Cambillau / Sylvain Moineau / Yannick Doyon / Adeline Goulet /    Abstract: In the arms race against bacteria, bacteriophages have evolved diverse anti-CRISPR proteins (Acrs) that block CRISPR-Cas immunity. Acrs play key roles in the molecular coevolution of bacteria with ...In the arms race against bacteria, bacteriophages have evolved diverse anti-CRISPR proteins (Acrs) that block CRISPR-Cas immunity. Acrs play key roles in the molecular coevolution of bacteria with their predators, use a variety of mechanisms of action, and provide tools to regulate Cas-based genome manipulation. Here, we present structural and functional analyses of AcrIIA6, an Acr from virulent phages, exploring its unique anti-CRISPR action. Our cryo-EM structures and functional data of AcrIIA6 binding to Streptococcus thermophilus Cas9 (St1Cas9) show that AcrIIA6 acts as an allosteric inhibitor and induces St1Cas9 dimerization. AcrIIA6 reduces St1Cas9 binding affinity for DNA and prevents DNA binding within cells. The PAM and AcrIIA6 recognition sites are structurally close and allosterically linked. Mechanistically, AcrIIA6 affects the St1Cas9 conformational dynamics associated with PAM binding. Finally, we identify a natural St1Cas9 variant resistant to AcrIIA6 illustrating Acr-driven mutational escape and molecular diversification of Cas9 proteins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4902.map.gz emd_4902.map.gz | 92.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4902-v30.xml emd-4902-v30.xml emd-4902.xml emd-4902.xml | 19.2 KB 19.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4902_fsc.xml emd_4902_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_4902.png emd_4902.png | 116.8 KB | ||
| Masks |  emd_4902_msk_1.map emd_4902_msk_1.map | 98.9 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-4902.cif.gz emd-4902.cif.gz | 7.1 KB | ||
| Others |  emd_4902_additional.map.gz emd_4902_additional.map.gz | 77.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4902 http://ftp.pdbj.org/pub/emdb/structures/EMD-4902 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4902 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4902 | HTTPS FTP |
-Related structure data
| Related structure data |  6rjdMC  4900C  4901C  4904C  6rj9C  6rjaC  6rjgC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4902.map.gz / Format: CCP4 / Size: 98.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4902.map.gz / Format: CCP4 / Size: 98.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Postprocessed 3D reconstruction of St1Cas9%u2022sgRNA%u2022tDNA59-ntPAM complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.889 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_4902_msk_1.map emd_4902_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Unsharpened 3D reconstruction of St1Cas9%u2022sgRNA%u2022tDNA59-ntPAM compelx...
| File | emd_4902_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened 3D reconstruction of St1Cas9%u2022sgRNA%u2022tDNA59-ntPAM compelx | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of St1Cas9-sgRNA-tDNA59-ntPAM complex.
| Entire | Name: Cryo-EM structure of St1Cas9-sgRNA-tDNA59-ntPAM complex. |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of St1Cas9-sgRNA-tDNA59-ntPAM complex.
| Supramolecule | Name: Cryo-EM structure of St1Cas9-sgRNA-tDNA59-ntPAM complex. type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: Streptococcus Thermophilus 1 Cas9
| Supramolecule | Name: Streptococcus Thermophilus 1 Cas9 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Streptococcus thermophilus DGCC 7710 (bacteria) Streptococcus thermophilus DGCC 7710 (bacteria) |
-Supramolecule #3: sgRNA (78-MER)
| Supramolecule | Name: sgRNA (78-MER) / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Streptococcus thermophilus (bacteria) Streptococcus thermophilus (bacteria) |
-Supramolecule #4: tDNA59
| Supramolecule | Name: tDNA59 / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #3 |
|---|---|
| Source (natural) | Organism:  Streptococcus phage D1811 (virus) Streptococcus phage D1811 (virus) |
-Supramolecule #5: ntPAM
| Supramolecule | Name: ntPAM / type: complex / ID: 5 / Parent: 1 / Macromolecule list: #4 |
|---|---|
| Source (natural) | Organism:  Streptococcus phage 2972 (virus) Streptococcus phage 2972 (virus) |
-Macromolecule #1: Streptococcus Thermophilus 1 Cas9
| Macromolecule | Name: Streptococcus Thermophilus 1 Cas9 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: Hydrolases; Acting on ester bonds |
|---|---|
| Source (natural) | Organism:  Streptococcus thermophilus DGCC 7710 (bacteria) Streptococcus thermophilus DGCC 7710 (bacteria) |
| Molecular weight | Theoretical: 129.700961 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSDLVLGLDI GIGSVGVGIL NKVTGEIIHK NSRIFPAAQA ENNLVRRTNR QGRRLTRRKK HRRVRLNRLF EESGLITDFT KISINLNPY QLRVKGLTDE LSNEELFIAL KNMVKHRGIS YLDDASDDGN SSIGDYAQIV KENSKQLETK TPGQIQLERY Q TYGQLRGD ...String: MSDLVLGLDI GIGSVGVGIL NKVTGEIIHK NSRIFPAAQA ENNLVRRTNR QGRRLTRRKK HRRVRLNRLF EESGLITDFT KISINLNPY QLRVKGLTDE LSNEELFIAL KNMVKHRGIS YLDDASDDGN SSIGDYAQIV KENSKQLETK TPGQIQLERY Q TYGQLRGD FTVEKDGKKH RLINVFPTSA YRSEALRILQ TQQEFNPQIT DEFINRYLEI LTGKRKYYHG PGNEKSRTDY GR YRTSGET LDNIFGILIG KCTFYPDEFR AAKASYTAQE FNLLNDLNNL TVPTETKKLS KEQKNQIINY VKNEKAMGPA KLF KYIAKL LSCDVADIKG YRIDKSGKAE IHTFEAYRKM KTLETLDIEQ MDRETLDKLA YVLTLNTERE GIQEALEHEF ADGS FSQKQ VDELVQFRKA NSSIFGKGWH NFSVKLMMEL IPELYETSEE QMTILTRLGK QKTTSSSNKT KYIDEKLLTE EIYNP VVAK SVRQAIKIVN AAIKEYGDFD NIVIEMARET NEDDEKKAIQ KIQKANKDEK DAAMLKAANQ YNGKAELPHS VFHGHK QLA TKIRLWHQQG ERCLYTGKTI SIHDLINNSN QFEVDHILPL SITFDDSLAN KVLVYATANQ EKGQRTPYQA LDSMDDA WS FRELKAFVRE SKTLSNKKKE YLLTEEDISK FDVRKKFIER NLVDTRYASR VVLNALQEHF RAHKIDTKVS VVRGQFTS Q LRRHWGIEKT RDTYHHHAVD ALIIAASSQL NLWKKQKNTL VSYSEDQLLD IETGELISDD EYKESVFKAP YQHFVDTLK SKEFEDSILF SYQVDSKFNR KISDATIYAT RQAKVGKDKA DETYVLGKIK DIYTQDGYDA FMKIYKKDKS KFLMYRHDPQ TFEKVIEPI LENYPNKQIN EKGKEVPCNP FLKYKEEHGY IRKYSKKGNG PEIKSLKYYD SKLGNHIDIT PKDSNNKVVL Q SVSPWRAD VYFNKTTGKY EILGLKYADL QFEKGTGTYK ISQEKYNDIK KKEGVDSDSE FKFTLYKNDL LLVKDTETKE QQ LFRFLSR TMPKQKHYVE LKPYDKQKFE GGEALIKVLG NVANSGQCKK GLGKSNISIY KVRTDVLGNQ HIIKNEGDKP KLD F |
-Macromolecule #2: sgRNA (78-MER)
| Macromolecule | Name: sgRNA (78-MER) / type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Streptococcus thermophilus DGCC 7710 (bacteria) Streptococcus thermophilus DGCC 7710 (bacteria) |
| Molecular weight | Theoretical: 37.438039 KDa |
| Sequence | String: GUUGCGUUGA UAAAAGUAUU GUUUUUGUAC UCUCAAGAUU CAAUAAUCUU GCAGAAGCUA CAAAGAUAAG GCUUCAUGCC GAAAUCAAC ACCCUGUCAU UUUAUGGCAG GGUGUUUU |
-Macromolecule #3: tDNA59
| Macromolecule | Name: tDNA59 / type: dna / ID: 3 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Streptococcus phage D1811 (virus) Streptococcus phage D1811 (virus) |
| Molecular weight | Theoretical: 18.160625 KDa |
| Sequence | String: (DA)(DT)(DG)(DG)(DT)(DT)(DC)(DT)(DG)(DG) (DT)(DT)(DT)(DC)(DA)(DT)(DT)(DT)(DT)(DC) (DT)(DG)(DC)(DA)(DA)(DT)(DA)(DC)(DT) (DT)(DT)(DT)(DA)(DT)(DC)(DA)(DA)(DC)(DG) (DC) (DA)(DA)(DG)(DA)(DG)(DG) ...String: (DA)(DT)(DG)(DG)(DT)(DT)(DC)(DT)(DG)(DG) (DT)(DT)(DT)(DC)(DA)(DT)(DT)(DT)(DT)(DC) (DT)(DG)(DC)(DA)(DA)(DT)(DA)(DC)(DT) (DT)(DT)(DT)(DA)(DT)(DC)(DA)(DA)(DC)(DG) (DC) (DA)(DA)(DG)(DA)(DG)(DG)(DT)(DG) (DC)(DT)(DT)(DC)(DT)(DG)(DT)(DT)(DA)(DT) (DG) |
-Macromolecule #4: ntPAM
| Macromolecule | Name: ntPAM / type: dna / ID: 4 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Streptococcus Phage 2972 (virus) Streptococcus Phage 2972 (virus) |
| Molecular weight | Theoretical: 7.084642 KDa |
| Sequence | String: (DG)(DC)(DA)(DG)(DA)(DA)(DA)(DA)(DT)(DG) (DA)(DA)(DA)(DC)(DC)(DA)(DG)(DA)(DA)(DC) (DC)(DA)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.75 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER |
|---|---|
| Output model |  PDB-6rjd: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)