[English] 日本語
 Yorodumi
Yorodumi- EMDB-4873: Cryo-EM structure of respiratory complex I from Yarrowia lipolyti... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4873 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of respiratory complex I from Yarrowia lipolytica at 3.2 A resolution | |||||||||
 Map data Map data | complex I from Yarrowia lipolytica | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex I / NADH dehydrogenase / Mitochondrion Proton pumping / Ubiquinone / OXIDOREDUCTASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationNADH dehydrogenase / TIM23 mitochondrial import inner membrane translocase complex / membrane protein complex / NADH dehydrogenase complex / mitochondrial [2Fe-2S] assembly complex / ubiquinone biosynthetic process / protein import into mitochondrial matrix / NADH:ubiquinone reductase (H+-translocating) / ubiquinone binding / mitochondrial electron transport, NADH to ubiquinone ...NADH dehydrogenase / TIM23 mitochondrial import inner membrane translocase complex / membrane protein complex / NADH dehydrogenase complex / mitochondrial [2Fe-2S] assembly complex / ubiquinone biosynthetic process / protein import into mitochondrial matrix / NADH:ubiquinone reductase (H+-translocating) / ubiquinone binding / mitochondrial electron transport, NADH to ubiquinone / electron transport coupled proton transport / acyl binding / mitochondrial respiratory chain complex I assembly / NADH dehydrogenase activity / oxidoreductase activity, acting on NAD(P)H / respiratory chain complex I / NADH dehydrogenase (ubiquinone) activity / acyl carrier activity / catalytic complex / quinone binding / ATP synthesis coupled electron transport / aerobic respiration / respiratory electron transport chain / electron transport chain / mitochondrial intermembrane space / 2 iron, 2 sulfur cluster binding / mitochondrial membrane / NAD binding / FMN binding / 4 iron, 4 sulfur cluster binding / response to oxidative stress / oxidoreductase activity / mitochondrial inner membrane / protein-containing complex binding / mitochondrion / membrane / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Yarrowia lipolytica (yeast) Yarrowia lipolytica (yeast) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | |||||||||
 Authors Authors | Parey K / Vonck J | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2019 Journal: Sci Adv / Year: 2019Title: High-resolution cryo-EM structures of respiratory complex I: Mechanism, assembly, and disease. Authors: Kristian Parey / Outi Haapanen / Vivek Sharma / Harald Köfeler / Thomas Züllig / Simone Prinz / Karin Siegmund / Ilka Wittig / Deryck J Mills / Janet Vonck / Werner Kühlbrandt / Volker Zickermann /    Abstract: Respiratory complex I is a redox-driven proton pump, accounting for a large part of the electrochemical gradient that powers mitochondrial adenosine triphosphate synthesis. Complex I dysfunction is ...Respiratory complex I is a redox-driven proton pump, accounting for a large part of the electrochemical gradient that powers mitochondrial adenosine triphosphate synthesis. Complex I dysfunction is associated with severe human diseases. Assembly of the one-megadalton complex I in the inner mitochondrial membrane requires assembly factors and chaperones. We have determined the structure of complex I from the aerobic yeast by electron cryo-microscopy at 3.2-Å resolution. A ubiquinone molecule was identified in the access path to the active site. The electron cryo-microscopy structure indicated an unusual lipid-protein arrangement at the junction of membrane and matrix arms that was confirmed by molecular simulations. The structure of a complex I mutant and an assembly intermediate provide detailed molecular insights into the cause of a hereditary complex I-linked disease and complex I assembly in the inner mitochondrial membrane. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4873.map.gz emd_4873.map.gz | 338.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4873-v30.xml emd-4873-v30.xml emd-4873.xml emd-4873.xml | 64.1 KB 64.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4873_fsc.xml emd_4873_fsc.xml | 16.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_4873.png emd_4873.png | 146.1 KB | ||
| Filedesc metadata |  emd-4873.cif.gz emd-4873.cif.gz | 14.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4873 http://ftp.pdbj.org/pub/emdb/structures/EMD-4873 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4873 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4873 | HTTPS FTP |
-Related structure data
| Related structure data |  6rfrMC  4872C  4874C  6rfqC  6rfsC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4873.map.gz / Format: CCP4 / Size: 361.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4873.map.gz / Format: CCP4 / Size: 361.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | complex I from Yarrowia lipolytica | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.077 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
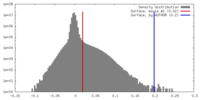
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Mitochondrial NADH:ubiquinone oxidoreductase
+Supramolecule #1: Mitochondrial NADH:ubiquinone oxidoreductase
+Macromolecule #1: Subunit NUAM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #2: Subunit NUBM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #3: Subunit NUCM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #4: Subunit NIMM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #5: Subunit NUEM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #6: Subunit NUFM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #7: Subunit NUGM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #8: Subunit NUHM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #9: Subunit NUIM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #10: Subunit NUJM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #11: Subunit NUKM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #12: Subunit NULM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #13: Subunit NUMM of protein NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #14: Acyl carrier protein ACPM1 of NADH:Ubiquinone Oxidoreductase (Com...
+Macromolecule #15: Subunit NB4M of protein NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #16: Acyl carrier protein ACPM2 of NADH:Ubiquinone Oxidoreductase (Com...
+Macromolecule #17: Subunit NI2M of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #18: Subunit NESM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #19: Subunit NUPM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #20: Subunit NB6M of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #21: Subunit NUXM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #22: Subunit NUYM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #23: Subunit NUZM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #24: Subunit NIAM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #25: Subunit NEBM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #26: Subunit NB2M of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #27: Subunit NIDM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #28: Subunit NUVM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #29: Subunit NI8M of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #30: Subunit NI9M of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #31: Subunit N7BM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #32: Subunit NUUM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #33: Subunit NB5M of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #34: Subunit NUNM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #35: Subunit NU1M of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #36: Subunit NU2M of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #37: Subunit NU3M of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #38: Subunit NU4M of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #39: Subunit NU5M of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #40: Subunit NU6M of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #41: Subunit NB8M of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #42: Subunit NIPM of NADH:Ubiquinone Oxidoreductase (Complex I)
+Macromolecule #43: IRON/SULFUR CLUSTER
+Macromolecule #44: FE2/S2 (INORGANIC) CLUSTER
+Macromolecule #45: FLAVIN MONONUCLEOTIDE
+Macromolecule #46: 1,2-Distearoyl-sn-glycerophosphoethanolamine
+Macromolecule #47: NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE
+Macromolecule #48: CARDIOLIPIN
+Macromolecule #49: Lauryl Maltose Neopentyl Glycol
+Macromolecule #50: DIUNDECYL PHOSPHATIDYL CHOLINE
+Macromolecule #51: ZINC ION
+Macromolecule #52: S-[2-({N-[(2S)-2-hydroxy-3,3-dimethyl-4-(phosphonooxy)butanoyl]-b...
+Macromolecule #53: Ubiquinone-9
+Macromolecule #54: Phosphatidylinositol
+Macromolecule #55: 1-PALMITOYL-2-LINOLEOYL-SN-GLYCERO-3-PHOSPHOCHOLINE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.2 Component:
| |||||||||||||||
| Grid | Model: C-flat-1/1 / Material: COPPER / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 80 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number real images: 4043 / Average exposure time: 8.0 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 46425 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: -2.5 µm / Nominal defocus min: -1.5 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)