+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4162 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
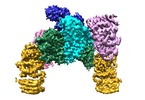
| Title | Eukaryotic initiation factor EIF2B in complex with ISRIB | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | GEF / Complex / ISRIB / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationeukaryotic translation initiation factor 2B complex / Recycling of eIF2:GDP / astrocyte development / astrocyte differentiation / oligodendrocyte development / regulation of translational initiation / cytoplasmic translational initiation / positive regulation of translational initiation / response to glucose / ovarian follicle development ...eukaryotic translation initiation factor 2B complex / Recycling of eIF2:GDP / astrocyte development / astrocyte differentiation / oligodendrocyte development / regulation of translational initiation / cytoplasmic translational initiation / positive regulation of translational initiation / response to glucose / ovarian follicle development / myelination / translation initiation factor binding / translation initiation factor activity / guanyl-nucleotide exchange factor activity / response to endoplasmic reticulum stress / central nervous system development / hippocampus development / translational initiation / response to peptide hormone / regulation of translation / T cell receptor signaling pathway / response to heat / positive regulation of apoptotic process / GTP binding / ATP binding / membrane / identical protein binding / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  Homo Sapiens (human) Homo Sapiens (human) | |||||||||
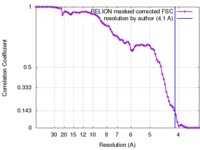
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | Faille A / Weis F | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2018 Journal: Science / Year: 2018Title: Binding of ISRIB reveals a regulatory site in the nucleotide exchange factor eIF2B. Authors: Alisa F Zyryanova / Félix Weis / Alexandre Faille / Akeel Abo Alard / Ana Crespillo-Casado / Yusuke Sekine / Heather P Harding / Felicity Allen / Leopold Parts / Christophe Fromont / Peter ...Authors: Alisa F Zyryanova / Félix Weis / Alexandre Faille / Akeel Abo Alard / Ana Crespillo-Casado / Yusuke Sekine / Heather P Harding / Felicity Allen / Leopold Parts / Christophe Fromont / Peter M Fischer / Alan J Warren / David Ron /  Abstract: The integrated stress response (ISR) is a conserved translational and transcriptional program affecting metabolism, memory, and immunity. The ISR is mediated by stress-induced phosphorylation of ...The integrated stress response (ISR) is a conserved translational and transcriptional program affecting metabolism, memory, and immunity. The ISR is mediated by stress-induced phosphorylation of eukaryotic translation initiation factor 2α (eIF2α) that attenuates the guanine nucleotide exchange factor eIF2B. A chemical inhibitor of the ISR, ISRIB, reverses the attenuation of eIF2B by phosphorylated eIF2α, protecting mice from neurodegeneration and traumatic brain injury. We describe a 4.1-angstrom-resolution cryo-electron microscopy structure of human eIF2B with an ISRIB molecule bound at the interface between the β and δ regulatory subunits. Mutagenesis of residues lining this pocket altered the hierarchical cellular response to ISRIB analogs in vivo and ISRIB binding in vitro. Our findings point to a site in eIF2B that can be exploited by ISRIB to regulate translation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4162.map.gz emd_4162.map.gz | 4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4162-v30.xml emd-4162-v30.xml emd-4162.xml emd-4162.xml | 23.8 KB 23.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_4162_fsc.xml emd_4162_fsc.xml | 8.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_4162.png emd_4162.png | 214.2 KB | ||
| Masks |  emd_4162_msk_1.map emd_4162_msk_1.map | 52.7 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-4162.cif.gz emd-4162.cif.gz | 7.9 KB | ||
| Others |  emd_4162_half_map_1.map.gz emd_4162_half_map_1.map.gz emd_4162_half_map_2.map.gz emd_4162_half_map_2.map.gz | 39.7 MB 39.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4162 http://ftp.pdbj.org/pub/emdb/structures/EMD-4162 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4162 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4162 | HTTPS FTP |
-Related structure data
| Related structure data |  6ezoMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4162.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4162.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.75 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
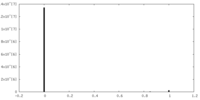
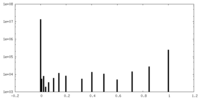
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_4162_msk_1.map emd_4162_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
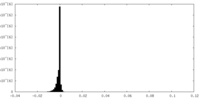
| Density Histograms |
-Half map: #1
| File | emd_4162_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
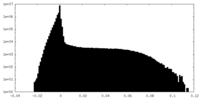
| Density Histograms |
-Half map: #2
| File | emd_4162_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
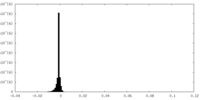
| Density Histograms |
- Sample components
Sample components
-Entire : Eukaryotic initiation factor EIF2B in complex with ISRIB
| Entire | Name: Eukaryotic initiation factor EIF2B in complex with ISRIB |
|---|---|
| Components |
|
-Supramolecule #1: Eukaryotic initiation factor EIF2B in complex with ISRIB
| Supramolecule | Name: Eukaryotic initiation factor EIF2B in complex with ISRIB type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Molecular weight | Theoretical: 521 KDa |
-Supramolecule #3: ISRIB
| Supramolecule | Name: ISRIB / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #2: Eukaryotic translation initiation factor EIF2B
| Supramolecule | Name: Eukaryotic translation initiation factor EIF2B / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1, #3-#5 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Translation initiation factor eIF-2B subunit alpha
| Macromolecule | Name: Translation initiation factor eIF-2B subunit alpha / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Tissue: cervix Homo sapiens (human) / Tissue: cervix |
| Molecular weight | Theoretical: 33.754148 KDa |
| Sequence | String: MDDKELIEYF KSQMKEDPDM ASAVAAIRTL LEFLKRDKGE TIQGLRANLT SAIETLCGVD SSVAVSSGGE LFLRFISLAS LEYSDYSKC KKIMIERGEL FLRRISLSRN KIADLCHTFI KDGATILTHA YSRVVLRVLE AAVAAKKRFS VYVTESQPDL S GKKMAKAL ...String: MDDKELIEYF KSQMKEDPDM ASAVAAIRTL LEFLKRDKGE TIQGLRANLT SAIETLCGVD SSVAVSSGGE LFLRFISLAS LEYSDYSKC KKIMIERGEL FLRRISLSRN KIADLCHTFI KDGATILTHA YSRVVLRVLE AAVAAKKRFS VYVTESQPDL S GKKMAKAL CHLNVPVTVV LDAAVGYIME KADLVIVGAE GVVENGGIIN KIGTNQMAVC AKAQNKPFYV VAESFKFVRL FP LNQQDVP DKFKYKADTL KVAQTGQDLK EEHPWVDYTA PSLITLLFTD LGVLTPSAVS DELIKLYL UniProtKB: Translation initiation factor eIF2B subunit alpha |
-Macromolecule #2: Translation initiation factor eIF-2B subunit beta
| Macromolecule | Name: Translation initiation factor eIF-2B subunit beta / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Tissue: cervix Homo sapiens (human) / Tissue: cervix |
| Molecular weight | Theoretical: 41.758242 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MPGSDYKDHD GDYKDHDIDY KDDDDKAAKG SELSERIESF VETLKRGGGP RSSEEMARET LGLLRQIITD HRWSNAGELM ELIRREGRR MTAAQPSETT VGNMVRRVLK IIREEYGRLH GRSDESDQQE SLHKLLTSGG LNEDFSFHYA QLQSNIIEAI N ELLVELEG ...String: MPGSDYKDHD GDYKDHDIDY KDDDDKAAKG SELSERIESF VETLKRGGGP RSSEEMARET LGLLRQIITD HRWSNAGELM ELIRREGRR MTAAQPSETT VGNMVRRVLK IIREEYGRLH GRSDESDQQE SLHKLLTSGG LNEDFSFHYA QLQSNIIEAI N ELLVELEG TMENIAAQAL EHIHSNEVIM TIGFSRTVEA FLKEAARKRK FHVIVAECAP FCQGHEMAVN LSKAGIETTV MT DAAIFAV MSRVNKVIIG TKTILANGAL RAVTGTHTLA LAAKHHSTPL IVCAPMFKLS PQFPNEEDSF HKFVAPEEVL PFT EGDILE KVSVHCPVFD YVPPELITLF ISNIGGNAPS YIYRLMSELY HPDDHVL UniProtKB: Translation initiation factor eIF2B subunit beta |
-Macromolecule #3: Translation initiation factor eIF-2B subunit gamma
| Macromolecule | Name: Translation initiation factor eIF-2B subunit gamma / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Tissue: cervix Homo sapiens (human) / Tissue: cervix |
| Molecular weight | Theoretical: 50.30423 KDa |
| Sequence | String: MEFQAVVMAV GGGSRMTDLT SSIPKPLLPV GNKPLIWYPL NLLERVGFEE VIVVTTRDVQ KALCAEFKMK MKPDIVCIPD DADMGTADS LRYIYPKLKT DVLVLSCDLI TDVALHEVVD LFRAYDASLA MLMRKGQDSI EPVPGQKGKK KAVEQRDFIG V DSTGKRLL ...String: MEFQAVVMAV GGGSRMTDLT SSIPKPLLPV GNKPLIWYPL NLLERVGFEE VIVVTTRDVQ KALCAEFKMK MKPDIVCIPD DADMGTADS LRYIYPKLKT DVLVLSCDLI TDVALHEVVD LFRAYDASLA MLMRKGQDSI EPVPGQKGKK KAVEQRDFIG V DSTGKRLL FMANEADLDE ELVIKGSILQ KHPRIRFHTG LVDAHLYCLK KYIVDFLMEN GSITSIRSEL IPYLVRKQFS SA SSQQGQE EKEEDLKKKE LKSLDIYSFI KEANTLNLAP YDACWNACRG DRWEDLSRSQ VRCYVHIMKE GLCSRVSTLG LYM EANRQV PKLLSALCPE EPPVHSSAQI VSKHLVGVDS LIGPETQIGE KSSIKRSVIG SSCLIKDRVT ITNCLLMNSV TVEE GSNIQ GSVICNNAVI EKGADIKDCL IGSGQRIEAK AKRVNEVIVG NDQLMEI UniProtKB: Translation initiation factor eIF2B subunit gamma |
-Macromolecule #4: Translation initiation factor eIF-2B subunit delta
| Macromolecule | Name: Translation initiation factor eIF-2B subunit delta / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Tissue: cervix Homo sapiens (human) / Tissue: cervix |
| Molecular weight | Theoretical: 57.640168 KDa |
| Sequence | String: MAAVAVAVRE DSGSGMKAEL PPGPGAVGRE MTKEEKLQLR KEKKQQKKKR KEEKGAEPET GSAVSAAQCQ VGPTRELPES GIQLGTPRE KVPAGRSKAE LRAERRAKQE AERALKQARK GEQGGPPPKA SPSTAGETPS GVKRLPEYPQ VDDLLLRRLV K KPERQQVP ...String: MAAVAVAVRE DSGSGMKAEL PPGPGAVGRE MTKEEKLQLR KEKKQQKKKR KEEKGAEPET GSAVSAAQCQ VGPTRELPES GIQLGTPRE KVPAGRSKAE LRAERRAKQE AERALKQARK GEQGGPPPKA SPSTAGETPS GVKRLPEYPQ VDDLLLRRLV K KPERQQVP TRKDYGSKVS LFSHLPQYSR QNSLTQFMSI PSSVIHPAMV RLGLQYSQGL VSGSNARCIA LLRALQQVIQ DY TTPPNEE LSRDLVNKLK PYMSFLTQCR PLSASMHNAI KFLNKEITSV GSSKREEEAK SELRAAIDRY VQEKIVLAAQ AIS RFAYQK ISNGDVILVY GCSSLVSRIL QEAWTEGRRF RVVVVDSRPW LEGRHTLRSL VHAGVPASYL LIPAASYVLP EVSK VLLGA HALLANGSVM SRVGTAQLAL VARAHNVPVL VCCETYKFCE RVQTDAFVSN ELDDPDDLQC KRGEHVALAN WQNHA SLRL LNLVYDVTPP ELVDLVITEL GMIPCSSVPV VLRVKSSDQ UniProtKB: Translation initiation factor eIF2B subunit delta |
-Macromolecule #5: Human eukaryotic initiation factor EIF2B epsilon subunits
| Macromolecule | Name: Human eukaryotic initiation factor EIF2B epsilon subunits type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo Sapiens (human) / Tissue: cervix Homo Sapiens (human) / Tissue: cervix |
| Molecular weight | Theoretical: 80.466609 KDa |
| Sequence | String: MAAPVVAPPG VVVSRANKRS GAGPGGSGGG GARGAEEEPP PPLQAVLVAD SFDRRFFPIS KDQPRVLLPL ANVALIDYTL EFLTATGVQ ETFVFCCWKA AQIKEHLLKS KWCRPTSLNV VRIITSELYR SLGDVLRDVD AKALVRSDFL LVYGDVISNI N ITRALEEH ...String: MAAPVVAPPG VVVSRANKRS GAGPGGSGGG GARGAEEEPP PPLQAVLVAD SFDRRFFPIS KDQPRVLLPL ANVALIDYTL EFLTATGVQ ETFVFCCWKA AQIKEHLLKS KWCRPTSLNV VRIITSELYR SLGDVLRDVD AKALVRSDFL LVYGDVISNI N ITRALEEH RLRRKLEKNV SVMTMIFKES SPSHPTRCHE DNVVVAVDST TNRVLHFQKT QGLRRFAFPL SLFQGSSDGV EV RYDLLDC HISICSPQVA QLFTDNFDYQ TRDDFVRGLL VNEEILGNQI HMHVTAKEYG ARVSNLHMYS AVCADVIRRW VYP LTPEAN FTDSTTQSCT HSRHNIYRGP EVSLGHGSIL EENVLLGSGT VIGSNCFITN SVIGPGCHIG DNVVLDQTYL WQGV RVAAG AQIHQSLLCD NAEVKERVTL KPRSVLTSQV VVGPNITLPE GSVISLHPPD AEEDEDDGEF SDDSGADQEK DKVKM KGYN PAEVGAAGKG YLWKAAGMNM EEEEELQQNL WGLKINMEEE SESESEQSMD SEEPDSRGGS PQMDDIKVFQ NEVLGT LQR GKEENISCDN LVLEINSLKY AYNISLKEVM QVLSHVVLEF PLQQMDSPLD SSRYCALLLP LLKAWSPVFR NYIKRAA DH LEALAAIEDF FLEHEALGIS MAKVLMAFYQ LEILAEETIL SWFSQRDTTD KGQQLRKNQQ LQRFIQWLKE AEEESSED D UniProtKB: Translation initiation factor eIF2B subunit epsilon |
-Macromolecule #6: 2-(4-chloranylphenoxy)-~{N}-[4-[2-(4-chloranylphenoxy)ethanoylami...
| Macromolecule | Name: 2-(4-chloranylphenoxy)-~{N}-[4-[2-(4-chloranylphenoxy)ethanoylamino]cyclohexyl]ethanamide type: ligand / ID: 6 / Number of copies: 1 / Formula: C7B |
|---|---|
| Molecular weight | Theoretical: 451.343 Da |
| Chemical component information | 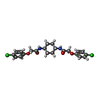 ChemComp-C7B: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.65 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Number real images: 765 / Average exposure time: 1.5 sec. / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 80000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 2.5 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6ezo: |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)