+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of AP2 bound to MSP2N2 nanodisc | |||||||||
 Map data Map data | Full map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | clathrin-dependent endocytosis / peripheral membrane protein / ENDOCYTOSIS | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
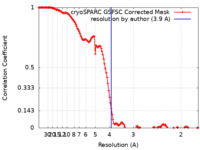
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | |||||||||
 Authors Authors | Sarsam RD / Cannon KS / Baker RW | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: J Struct Biol / Year: 2023 Journal: J Struct Biol / Year: 2023Title: Lipid nanodiscs as a template for high-resolution cryo-EM structures of peripheral membrane proteins. Authors: Kevin S Cannon / Reta D Sarsam / Tanita Tedamrongwanish / Kevin Zhang / Richard W Baker /  Abstract: Peripheral membrane proteins are ubiquitous throughout cell biology and are required for a variety of cellular processes such as signal transduction, membrane trafficking, and autophagy. Transient ...Peripheral membrane proteins are ubiquitous throughout cell biology and are required for a variety of cellular processes such as signal transduction, membrane trafficking, and autophagy. Transient binding to the membrane has a profound impact on protein function, serving to induce conformational changes and alter biochemical and biophysical parameters by increasing the local concentration of factors and restricting diffusion to two dimensions. Despite the centrality of the membrane in serving as a template for cell biology, there are few reported high-resolution structures of peripheral membrane proteins bound to the membrane. We analyzed the utility of lipid nanodiscs to serve as a template for cryo-EM analysis of peripheral membrane proteins. We tested a variety of nanodiscs and we report a 3.3 Å structure of the AP2 clathrin adaptor complex bound to a 17-nm nanodisc, with sufficient resolution to visualize a bound lipid head group. Our data demonstrate that lipid nanodiscs are amenable to high-resolution structure determination of peripheral membrane proteins and provide a framework for extending this analysis to other systems. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40035.map.gz emd_40035.map.gz | 107.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40035-v30.xml emd-40035-v30.xml emd-40035.xml emd-40035.xml | 28.4 KB 28.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_40035_fsc.xml emd_40035_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_40035.png emd_40035.png | 33 KB | ||
| Masks |  emd_40035_msk_1.map emd_40035_msk_1.map | 216 MB |  Mask map Mask map | |
| Others |  emd_40035_additional_1.map.gz emd_40035_additional_1.map.gz emd_40035_additional_2.map.gz emd_40035_additional_2.map.gz emd_40035_additional_3.map.gz emd_40035_additional_3.map.gz emd_40035_additional_4.map.gz emd_40035_additional_4.map.gz emd_40035_half_map_1.map.gz emd_40035_half_map_1.map.gz emd_40035_half_map_2.map.gz emd_40035_half_map_2.map.gz | 189.8 MB 4.4 MB 4.2 MB 203.8 MB 200.2 MB 200.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40035 http://ftp.pdbj.org/pub/emdb/structures/EMD-40035 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40035 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40035 | HTTPS FTP |
-Validation report
| Summary document |  emd_40035_validation.pdf.gz emd_40035_validation.pdf.gz | 850.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_40035_full_validation.pdf.gz emd_40035_full_validation.pdf.gz | 849.8 KB | Display | |
| Data in XML |  emd_40035_validation.xml.gz emd_40035_validation.xml.gz | 21.3 KB | Display | |
| Data in CIF |  emd_40035_validation.cif.gz emd_40035_validation.cif.gz | 27.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40035 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40035 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40035 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40035 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_40035.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40035.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Full map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.88 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_40035_msk_1.map emd_40035_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
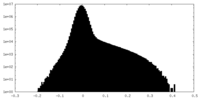
| Density Histograms |
-Additional map: deepEMhancer sharpened map
| File | emd_40035_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | deepEMhancer sharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
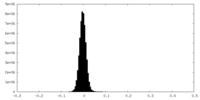
| Density Histograms |
-Additional map: locally filtered map
| File | emd_40035_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | locally filtered map | ||||||||||||
| Projections & Slices |
| ||||||||||||
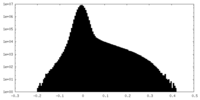
| Density Histograms |
-Additional map: Map for coloring by local resolution
| File | emd_40035_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map for coloring by local resolution | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: cryoSPARC sharpened map
| File | emd_40035_additional_4.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryoSPARC sharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1
| File | emd_40035_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2
| File | emd_40035_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : AP2 bound to MSP2N2 nanodisc
| Entire | Name: AP2 bound to MSP2N2 nanodisc |
|---|---|
| Components |
|
-Supramolecule #1: AP2 bound to MSP2N2 nanodisc
| Supramolecule | Name: AP2 bound to MSP2N2 nanodisc / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Nanodiscs were assembled with a lipid mixture containing 75 mol% DOPC, 15 mol% DOPS, 10 mol% PIP2. Complex was formed by co-elution via gel filtration chromatography. |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 203.98 KDa |
-Macromolecule #1: AP-2 complex subunit alpha-2
| Macromolecule | Name: AP-2 complex subunit alpha-2 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MPAVSKGDGM RGLAVFISDI RNCKSKEAEI KRINKELANI RSKFKGDKAL DGYSKKKYVC KLLFIFLLGH DIDFGHMEAV NLLSSNRYTE KQIGYLFISV LVNSNSELIR LINNAIKNDL ASRNPTFMGL ALHCIANVGS REMAEAFAGE IPKILVAGDT MDSVKQSAAL ...String: MPAVSKGDGM RGLAVFISDI RNCKSKEAEI KRINKELANI RSKFKGDKAL DGYSKKKYVC KLLFIFLLGH DIDFGHMEAV NLLSSNRYTE KQIGYLFISV LVNSNSELIR LINNAIKNDL ASRNPTFMGL ALHCIANVGS REMAEAFAGE IPKILVAGDT MDSVKQSAAL CLLRLYRTSP DLVPMGDWTS RVVHLLNDQH LGVVTAATSL ITTLAQKNPE EFKTSVSLAV SRLSRIVTSA STDLQDYTYY FVPAPWLSVK LLRLLQCYPP PEDPAVRGRL TECLETILNK AQEPPKSKKV QHSNAKNAVL FEAISLIIHH DSEPNLLVRA CNQLGQFLQH RETNLRYLAL ESMCTLASSE FSHEAVKTHI ETVINALKTE RDVSVRQRAV DLLYAMCDRS NAQQIVAEML SYLETADYSI REEIVLKVAI LAEKYAVDYT WYVDTILNLI RIAGDYVSEE VWYRVIQIVI NRDDVQGYAA KTVFEALQAP ACHENLVKVG GYILGEFGNL IAGDPRSSPL IQFNLLHSKF HLCSVPTRAL LLSTYIKFVN LFPEVKATIQ DVLRSDSQLK NADVELQQRA VEYLRLSTVA STDILATVLE EMPPFPERES SILAKLKKKK GGSGLEVLFQ |
-Macromolecule #2: AP-2 complex subunit beta
| Macromolecule | Name: AP-2 complex subunit beta / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MTDSKYFTTN KKGEIFELKA ELNNEKKEKR KEAVKKVIAA MTVGKDVSSL FPDVVNCMQT DNLELKKLVY LYLMNYAKSQ PDMAIMAVNS FVKDCEDPNP LIRALAVRTM GCIRVDKITE YLCEPLRKCL KDEDPYVRKT AAVCVAKLHD INAQMVEDQG FLDSLRDLIA ...String: MTDSKYFTTN KKGEIFELKA ELNNEKKEKR KEAVKKVIAA MTVGKDVSSL FPDVVNCMQT DNLELKKLVY LYLMNYAKSQ PDMAIMAVNS FVKDCEDPNP LIRALAVRTM GCIRVDKITE YLCEPLRKCL KDEDPYVRKT AAVCVAKLHD INAQMVEDQG FLDSLRDLIA DSNPMVVANA VAALSEISES HPNSNLLDLN PQNINKLLTA LNECTEWGQI FILDCLSNYN PKDDREAQSI CERVTPRLSH ANSAVVLSAV KVLMKFLELL PKDSDYYNML LKKLAPPLVT LLSGEPEVQY VALRNINLIV QKRPEILKQE IKVFFVKYND PIYVKLEKLD IMIRLASQAN IAQVLAELKE YATEVDVDFV RKAVRAIGRC AIKVEQSAER CVSTLLDLIQ TKVNYVVQEA IVVIRDIFRK YPNKYESIIA TLCENLDSLD EPDARAAMIW IVGEYAERID NADELLESFL EGFHDESTQV QLTLLTAIVK LFLKKPSETQ ELVQQVLSLA TQDSDNPDLR DRGYIYWRLL STDPVTAKEV VLSEKPLISE ETDLIEPTLL DELICHIGSL ASVYHKPPNA FVEGSHGIHR K |
-Macromolecule #3: AP-2 complex subunit mu
| Macromolecule | Name: AP-2 complex subunit mu / type: protein_or_peptide / ID: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MIGGLFIYNH KGEVLISRVY RDDIGRNAVD AFRVNVIHAR QQVRSPVTNI ARTSFFHVKR SNIWLAAVTK QNVNAAMVFE FLYKMCDVMA AYFGKISEEN IKNNFVLIYE LLDEILDFGY PQNSETGALK TFITQQGIKS QHQTKEEQSQ ITSQVTGQIG WRREGIKYRR ...String: MIGGLFIYNH KGEVLISRVY RDDIGRNAVD AFRVNVIHAR QQVRSPVTNI ARTSFFHVKR SNIWLAAVTK QNVNAAMVFE FLYKMCDVMA AYFGKISEEN IKNNFVLIYE LLDEILDFGY PQNSETGALK TFITQQGIKS QHQTKEEQSQ ITSQVTGQIG WRREGIKYRR NELFLDVLES VNLLMSPQGQ VLSAHVSGRV VMKSYLSGMP ECKFGMNDKI VIEKQGKGTA DETSKSGKQS IAIDDCTFHQ CVRLSKFDSE RSISFIPPDG EFELMRYRTT KDIILPFRVI PLVREVGRTK LEVKVVIKSN FKPSLLAQKI EVRIPTPLNT SGVQVICMKG KAKYKASENA IVWKIKRMAG MKESQISAEI ELLPTNDKKK WARPPISMNF EVPFAPSGLK VRYLKVFEPK LNYSDHDVIK WVRYIGRSGI YETRC |
-Macromolecule #4: AP-2 complex subunit sigma
| Macromolecule | Name: AP-2 complex subunit sigma / type: protein_or_peptide / ID: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MIRFILIQNR AGKTRLAKWY MQFDDDEKQK LIEEVHAVVT VRDAKHTNFV EFRNFKIIYR RYAGLYFCIC VDVNDNNLAY LEAIHNFVEV LNEYFHNVCE LDLVFNFYKV YTVVDEMFLA GEIRETSQTK VLKQLLMLQS LE |
-Macromolecule #5: MSP1E3D1
| Macromolecule | Name: MSP1E3D1 / type: protein_or_peptide / ID: 5 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  |
| Sequence | String: MGHHHHHHHD YDIPTTENLY FQGSTFSKLR EQLGPVTQEF WDNLEKETEG LRQEMSKDLE EVKAKVQPYL DDFQKKWQEE MELYRQKVEP LRAELQEGAR QKLHELQEKL SPLGEEMRDR ARAHVDALRT HLAPYLDDFQ KKWQEEMELY RQKVEPLRAE LQEGARQKLH ...String: MGHHHHHHHD YDIPTTENLY FQGSTFSKLR EQLGPVTQEF WDNLEKETEG LRQEMSKDLE EVKAKVQPYL DDFQKKWQEE MELYRQKVEP LRAELQEGAR QKLHELQEKL SPLGEEMRDR ARAHVDALRT HLAPYLDDFQ KKWQEEMELY RQKVEPLRAE LQEGARQKLH ELQEKLSPLG EEMRDRARAH VDALRTHLAP YSDELRQRLA ARLEALKENG GARLAEYHAK ATEHLSTLSE KAKPALEDLR QGLLPVLESF KVSFLSALEE YTKKLNTQ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 20 mM HEPES pH 7.4, 100 mM NaCl |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: OTHER / Details: Tergeo EM plasma cleaner |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: Two samples applications.. |
| Details | Nanodiscs were assembled with a lipid mixture containing 75 mol% DOPC, 15 mol% DOPS, 10 mol% PIP2. Complex was formed by co-elution via gel filtration chromatography. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 55.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated magnification: 45000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 45000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)