[English] 日本語
 Yorodumi
Yorodumi- PDB-3ux0: Crystal structure of human 14-3-3 sigma in complex with TASK-3 pe... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3ux0 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of human 14-3-3 sigma in complex with TASK-3 peptide and stabilizer Fusicoccin H | ||||||
 Components Components |
| ||||||
 Keywords Keywords | PEPTIDE BINDING PROTEIN / Helical protein / Phosphoprotein / Adapter protein / Nucleus | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of aldosterone secretion / TWIK-releated acid-sensitive K+ channel (TASK) / regulation of action potential firing rate / Phase 4 - resting membrane potential / potassium ion leak channel activity / regulation of resting membrane potential / outward rectifier potassium channel activity / regulation of epidermal cell division / protein kinase C inhibitor activity / positive regulation of epidermal cell differentiation ...negative regulation of aldosterone secretion / TWIK-releated acid-sensitive K+ channel (TASK) / regulation of action potential firing rate / Phase 4 - resting membrane potential / potassium ion leak channel activity / regulation of resting membrane potential / outward rectifier potassium channel activity / regulation of epidermal cell division / protein kinase C inhibitor activity / positive regulation of epidermal cell differentiation / keratinocyte development / keratinization / sodium channel activity / regulation of cell-cell adhesion / potassium ion import across plasma membrane / establishment of skin barrier / Regulation of localization of FOXO transcription factors / keratinocyte proliferation / Activation of BAD and translocation to mitochondria / phosphoserine residue binding / negative regulation of keratinocyte proliferation / cAMP/PKA signal transduction / negative regulation of protein localization to plasma membrane / potassium channel activity / SARS-CoV-2 targets host intracellular signalling and regulatory pathways / negative regulation of protein kinase activity / negative regulation of stem cell proliferation / SARS-CoV-1 targets host intracellular signalling and regulatory pathways / RHO GTPases activate PKNs / Chk1/Chk2(Cds1) mediated inactivation of Cyclin B:Cdk1 complex / cellular response to acidic pH / positive regulation of protein localization / visual perception / protein export from nucleus / TP53 Regulates Transcription of Genes Involved in G2 Cell Cycle Arrest / positive regulation of cell adhesion / negative regulation of innate immune response / release of cytochrome c from mitochondria / positive regulation of protein export from nucleus / stem cell proliferation / TP53 Regulates Metabolic Genes / Translocation of SLC2A4 (GLUT4) to the plasma membrane / potassium ion transport / protein sequestering activity / intrinsic apoptotic signaling pathway in response to DNA damage / synaptic vesicle / intracellular protein localization / regulation of protein localization / positive regulation of cell growth / sperm midpiece / regulation of cell cycle / mitochondrial inner membrane / cadherin binding / protein heterodimerization activity / dendrite / protein kinase binding / negative regulation of transcription by RNA polymerase II / signal transduction / : / extracellular exosome / metal ion binding / identical protein binding / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / MOLECULAR REPLACEMENT /  molecular replacement / Resolution: 1.75 Å molecular replacement / Resolution: 1.75 Å | ||||||
 Authors Authors | Thiel, P. / Bartel, M. / Anders, C. / Higuchi, Y. / Schumacher, B. / Kato, N. / Ottmann, C. | ||||||
 Citation Citation |  Journal: Chem.Biol. / Year: 2013 Journal: Chem.Biol. / Year: 2013Title: A semisynthetic fusicoccane stabilizes a protein-protein interaction and enhances the expression of k(+) channels at the cell surface. Authors: Anders, C. / Higuchi, Y. / Koschinsky, K. / Bartel, M. / Schumacher, B. / Thiel, P. / Nitta, H. / Preisig-Muller, R. / Schlichthorl, G. / Renigunta, V. / Ohkanda, J. / Daut, J. / Kato, N. / Ottmann, C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3ux0.cif.gz 3ux0.cif.gz | 77.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3ux0.ent.gz pdb3ux0.ent.gz | 56.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3ux0.json.gz 3ux0.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ux/3ux0 https://data.pdbj.org/pub/pdb/validation_reports/ux/3ux0 ftp://data.pdbj.org/pub/pdb/validation_reports/ux/3ux0 ftp://data.pdbj.org/pub/pdb/validation_reports/ux/3ux0 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  3p1nC 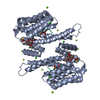 3p1oC  3p1pC  3p1qC  3p1rC  3p1sC  3smkC 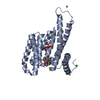 3smlC 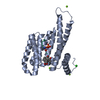 3smmC  3smnC  3smoC  3sp5C  3sprC  4fr3C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| |||||||||
| Unit cell |
| |||||||||
| Components on special symmetry positions |
|
- Components
Components
-Protein / Protein/peptide , 2 types, 2 molecules AP
| #1: Protein | Mass: 26501.865 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: HME1, SFN / Plasmid: PPROEX HTB / Production host: Homo sapiens (human) / Gene: HME1, SFN / Plasmid: PPROEX HTB / Production host:  |
|---|---|
| #2: Protein/peptide | Mass: 856.950 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: This sequence occurs naturally in humans. / Source: (synth.)  Homo sapiens (human) / References: UniProt: Q9NPC2 Homo sapiens (human) / References: UniProt: Q9NPC2 |
-Non-polymers , 5 types, 340 molecules 


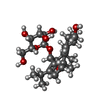





| #3: Chemical | ChemComp-MG / #4: Chemical | ChemComp-CL / | #5: Chemical | ChemComp-GOL / | #6: Chemical | ChemComp-0DV / ( | #7: Water | ChemComp-HOH / | |
|---|
-Details
| Has protein modification | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.59 Å3/Da / Density % sol: 52.52 % |
|---|---|
| Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 7.5 Details: 0.095M HEPES Na, 0.19M calcium chloride, 5% glycerol, 26.6% PEG400, pH 7.5, VAPOR DIFFUSION, HANGING DROP, temperature 277K |
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: BRUKER AXS MICROSTAR / Wavelength: 1.5418 Å ROTATING ANODE / Type: BRUKER AXS MICROSTAR / Wavelength: 1.5418 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: MAR scanner 345 mm plate / Detector: IMAGE PLATE / Date: Nov 11, 2011 / Details: mirrors | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Monochromator: CURVED MULTILAYER MIRRORS / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 1.75→19.464 Å / Num. all: 28378 / Num. obs: 28378 / % possible obs: 97.7 % / Observed criterion σ(F): -3 / Observed criterion σ(I): -3 / Redundancy: 5.86 % / Biso Wilson estimate: 24.991 Å2 / Rmerge(I) obs: 0.049 / Net I/σ(I): 23.75 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1
|
-Phasing
| Phasing | Method:  molecular replacement molecular replacement | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Phasing MR | Rfactor: 34.38 / Model details: Phaser MODE: MR_AUTO
|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 1.75→19.46 Å / Cor.coef. Fo:Fc: 0.967 / Cor.coef. Fo:Fc free: 0.944 / WRfactor Rfree: 0.185 / WRfactor Rwork: 0.1467 / Occupancy max: 1 / Occupancy min: 0.08 / FOM work R set: 0.9016 / SU B: 1.813 / SU ML: 0.059 / SU R Cruickshank DPI: 0.1027 / SU Rfree: 0.1033 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.103 / ESU R Free: 0.103 / Stereochemistry target values: MAXIMUM LIKELIHOOD MOLECULAR REPLACEMENT / Resolution: 1.75→19.46 Å / Cor.coef. Fo:Fc: 0.967 / Cor.coef. Fo:Fc free: 0.944 / WRfactor Rfree: 0.185 / WRfactor Rwork: 0.1467 / Occupancy max: 1 / Occupancy min: 0.08 / FOM work R set: 0.9016 / SU B: 1.813 / SU ML: 0.059 / SU R Cruickshank DPI: 0.1027 / SU Rfree: 0.1033 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.103 / ESU R Free: 0.103 / Stereochemistry target values: MAXIMUM LIKELIHOODDetails: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS U VALUES : REFINED INDIVIDUALLY
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.4 Å / Solvent model: MASK | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 60.36 Å2 / Biso mean: 19.6787 Å2 / Biso min: 2.32 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.75→19.46 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.75→1.795 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller












 PDBj
PDBj
















