+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3894 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | S.aureus ClpC resting state, C2 symmetrised | |||||||||
 Map data Map data | Resting state of ClpC from S. aureus. C2 symmetry applied. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ClpC / AAA+ protease / oligomeric complex / CHAPERONE | |||||||||
| Function / homology |  Function and homology information Function and homology informationstress response to cadmium ion / stress response to copper ion / protein folding chaperone / peptidase activity / cellular response to heat / ATP hydrolysis activity / proteolysis / ATP binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
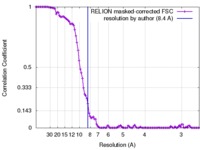
| Method | single particle reconstruction / cryo EM / Resolution: 8.4 Å | |||||||||
 Authors Authors | Carroni M / Mogk A | |||||||||
 Citation Citation |  Journal: Elife / Year: 2017 Journal: Elife / Year: 2017Title: Regulatory coiled-coil domains promote head-to-head assemblies of AAA+ chaperones essential for tunable activity control. Authors: Marta Carroni / Kamila B Franke / Michael Maurer / Jasmin Jäger / Ingo Hantke / Felix Gloge / Daniela Linder / Sebastian Gremer / Kürşad Turgay / Bernd Bukau / Axel Mogk /   Abstract: Ring-forming AAA+ chaperones exert ATP-fueled substrate unfolding by threading through a central pore. This activity is potentially harmful requiring mechanisms for tight repression and substrate- ...Ring-forming AAA+ chaperones exert ATP-fueled substrate unfolding by threading through a central pore. This activity is potentially harmful requiring mechanisms for tight repression and substrate-specific activation. The AAA+ chaperone ClpC with the peptidase ClpP forms a bacterial protease essential to virulence and stress resistance. The adaptor MecA activates ClpC by targeting substrates and stimulating ClpC ATPase activity. We show how ClpC is repressed in its ground state by determining ClpC cryo-EM structures with and without MecA. ClpC forms large two-helical assemblies that associate via head-to-head contacts between coiled-coil middle domains (MDs). MecA converts this resting state to an active planar ring structure by binding to MD interaction sites. Loss of ClpC repression in MD mutants causes constitutive activation and severe cellular toxicity. These findings unravel an unexpected regulatory concept executed by coiled-coil MDs to tightly control AAA+ chaperone activity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3894.map.gz emd_3894.map.gz | 8.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3894-v30.xml emd-3894-v30.xml emd-3894.xml emd-3894.xml | 16.3 KB 16.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3894_fsc.xml emd_3894_fsc.xml | 10.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_3894.png emd_3894.png | 126.2 KB | ||
| Masks |  emd_3894_msk_1.map emd_3894_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-3894.cif.gz emd-3894.cif.gz | 6.1 KB | ||
| Others |  emd_3894_half_map_1.map.gz emd_3894_half_map_1.map.gz emd_3894_half_map_2.map.gz emd_3894_half_map_2.map.gz | 79.4 MB 79.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3894 http://ftp.pdbj.org/pub/emdb/structures/EMD-3894 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3894 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3894 | HTTPS FTP |
-Related structure data
| Related structure data |  6em8MC  3895C  3897C  6em9C  6emwC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3894.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3894.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Resting state of ClpC from S. aureus. C2 symmetry applied. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_3894_msk_1.map emd_3894_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1. Resting state of ClpC from...
| File | emd_3894_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1. Resting state of ClpC from S. aureus. C2 symmetry applied. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2. Resting state of ClpC from...
| File | emd_3894_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2. Resting state of ClpC from S. aureus. C2 symmetry applied. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Resting-state oligomeric complex of S. aureus ClpC
| Entire | Name: Resting-state oligomeric complex of S. aureus ClpC |
|---|---|
| Components |
|
-Supramolecule #1: Resting-state oligomeric complex of S. aureus ClpC
| Supramolecule | Name: Resting-state oligomeric complex of S. aureus ClpC / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 900 KDa |
-Macromolecule #1: ATP-dependent Clp protease ATP-binding subunit ClpC
| Macromolecule | Name: ATP-dependent Clp protease ATP-binding subunit ClpC / type: protein_or_peptide / ID: 1 / Number of copies: 10 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 91.170352 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MLFGRLTERA QRVLAHAQEE AIRLNHSNIG TEHLLLGLMK EPEGIAAKVL ESFNITEDKV IEEVEKLIGH GQDHVGTLHY TPRAKKVIE LSMDEARKLH HNFVGTEHIL LGLIRENEGV AARVFANLDL NITKARAQVV KALGNPEMSN KNAQASKSNN T PTLDSLAR ...String: MLFGRLTERA QRVLAHAQEE AIRLNHSNIG TEHLLLGLMK EPEGIAAKVL ESFNITEDKV IEEVEKLIGH GQDHVGTLHY TPRAKKVIE LSMDEARKLH HNFVGTEHIL LGLIRENEGV AARVFANLDL NITKARAQVV KALGNPEMSN KNAQASKSNN T PTLDSLAR DLTVIAKDGT LDPVIGRDKE ITRVIEVLSR RTKNNPVLIG EPGVGKTAIA EGLAQAIVNN EVPETLKDKR VM SLDMGTV VAGTKYRGEF EERLKKVMEE IQQAGNVILF IDELHTLVGA GGAEGAIDAS NILKPALARG ELQCIGATTL DEY RKNIEK DAALERRFQP VQVDEPSVVD TVAILKGLRD RYEAHHRINI SDEAIEAAVK LSNRYVSDRF LPDKAIDLID EASS KVRLK SHTTPNNLKE IEQEIEKVKN EKDAAVHAQE FENAANLRDK QTKLEKQYEE AKNEWKNAQN GMSTSLSEED IAEVI AGWT GIPLTKINET ESEKLLSLED TLHERVIGQK DAVNSISKAV RRARAGLKDP KRPIGSFIFL GPTGVGKTEL ARALAE SMF GDDDAMIRVD MSEFMEKHAV SRLVGAPPGY VGHDDGGQLT EKVRRKPYSV ILFDEIEKAH PDVFNILLQV LDDGHLT DT KGRTVDFRNT IIIMTSNVGA QELQDQRFAG FGGSSDGQDY ETIRKTMLKE LKNSFRPEFL NRVDDIIVFH KLTKEELK E IVTMMVNKLT NRLSEQNINI IVTDKAKDKI AEEGYDPEYG ARPLIRAIQK TIEDNLSELI LDGNQIEGKK VTVDHDGKE FKYDIAEQTS ETKTPSQA UniProtKB: ATP-dependent Clp protease ATP-binding subunit ClpC |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: C-flat / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 40 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 297 K / Instrument: FEI VITROBOT MARK IV |
| Details | concentration used was 6uM |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Phyre2 base on the crystal structure PDB: 3pxi |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
| Output model |  PDB-6em8: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)