+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | BANAL-20-236 S1 in complex with R. Affinis ACE2 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | BANAL-20-236 / Bat / Spike / S1 / R. Affinis / ACE2 / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationHydrolases; Acting on peptide bonds (peptidases) / peptidyl-dipeptidase activity / carboxypeptidase activity / endocytosis involved in viral entry into host cell / cilium / metallopeptidase activity / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / receptor-mediated virion attachment to host cell / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane ...Hydrolases; Acting on peptide bonds (peptidases) / peptidyl-dipeptidase activity / carboxypeptidase activity / endocytosis involved in viral entry into host cell / cilium / metallopeptidase activity / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / receptor-mediated virion attachment to host cell / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / host cell plasma membrane / virion membrane / proteolysis / extracellular region / membrane / metal ion binding / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  unclassified Coronavirinae (virus) / unclassified Coronavirinae (virus) /  Rhinolophus affinis (intermediate horseshoe bat) Rhinolophus affinis (intermediate horseshoe bat) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Wang X / Xu G | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: The selective effect of fecal-oral transmission on the S proteins of bat SARS-CoV-2 related coronaviruses in favor of stability over infectivity Authors: Wang X / Xu G | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35079.map.gz emd_35079.map.gz | 54.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35079-v30.xml emd-35079-v30.xml emd-35079.xml emd-35079.xml | 15.3 KB 15.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_35079.png emd_35079.png | 43.6 KB | ||
| Filedesc metadata |  emd-35079.cif.gz emd-35079.cif.gz | 6.5 KB | ||
| Others |  emd_35079_half_map_1.map.gz emd_35079_half_map_1.map.gz emd_35079_half_map_2.map.gz emd_35079_half_map_2.map.gz | 45.1 MB 45.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35079 http://ftp.pdbj.org/pub/emdb/structures/EMD-35079 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35079 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35079 | HTTPS FTP |
-Validation report
| Summary document |  emd_35079_validation.pdf.gz emd_35079_validation.pdf.gz | 897.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_35079_full_validation.pdf.gz emd_35079_full_validation.pdf.gz | 897 KB | Display | |
| Data in XML |  emd_35079_validation.xml.gz emd_35079_validation.xml.gz | 11.8 KB | Display | |
| Data in CIF |  emd_35079_validation.cif.gz emd_35079_validation.cif.gz | 13.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35079 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35079 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35079 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-35079 | HTTPS FTP |
-Related structure data
| Related structure data |  8hxkMC  8hxjC  8i3wC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35079.map.gz / Format: CCP4 / Size: 58.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35079.map.gz / Format: CCP4 / Size: 58.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
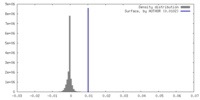
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_35079_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
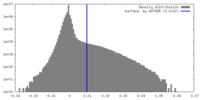
| Density Histograms |
-Half map: #2
| File | emd_35079_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : BANAL-20-236 S1 in complex with R. Affinis ACE2
| Entire | Name: BANAL-20-236 S1 in complex with R. Affinis ACE2 |
|---|---|
| Components |
|
-Supramolecule #1: BANAL-20-236 S1 in complex with R. Affinis ACE2
| Supramolecule | Name: BANAL-20-236 S1 in complex with R. Affinis ACE2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  unclassified Coronavirinae (virus) unclassified Coronavirinae (virus) |
-Macromolecule #1: Angiotensin-converting enzyme
| Macromolecule | Name: Angiotensin-converting enzyme / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Rhinolophus affinis (intermediate horseshoe bat) Rhinolophus affinis (intermediate horseshoe bat) |
| Molecular weight | Theoretical: 76.340852 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: METDTLLLWV LLLWVPGSTG KLSTTEDRAK IFLDNFNHEA EDLSYQSSLA SWEYNTNISD ENVQKMDEAG AKWSAFYEEQ SKLAKNYPL EEIQTVPVKL QLQILQQSGS PVLSEDKSKR LNSILNAMST IYSTGKVCKP NNPQECFLLE PGLDNIMGTS K DYNERLWA ...String: METDTLLLWV LLLWVPGSTG KLSTTEDRAK IFLDNFNHEA EDLSYQSSLA SWEYNTNISD ENVQKMDEAG AKWSAFYEEQ SKLAKNYPL EEIQTVPVKL QLQILQQSGS PVLSEDKSKR LNSILNAMST IYSTGKVCKP NNPQECFLLE PGLDNIMGTS K DYNERLWA WEGWRAEVGK QLRPLYEEYV ALKNEMARGY HYEDYGDYWR RDYETEESSG SGYSRDQLMK DVDRIFTEIK PL YEHLHAY VRTKLMDTYP FHISPTGCLP AHLLGDMWGR FWTNLYPLTV PFGQKPNIDV TDAMVNQGWD ANRIFKEAEK FFV SVGLPN MTEGFWNNSM LTEPGDGRKV VCHPTAWDLG KGDFRIKMCT KVTMEDFLTA HHEMGHIQYD MAYATQPYLL RNGA NEGFH EAVGEVMSLS VATPKHLKTM GLLSPDFLED NETEINFLLK QALNIVGTLP FTYMLEKWRW MVFRGEIPKE EWMKK WWEM KRDLVGVVEP VPHDETYCDP ASLFHVANDY SFIRYYTRTI FEFQFHEALC RIAQHDGPLH KCDISNSTDA GKKLHQ MLS VGKSQPWTVT LKDIVDSRNM DVGPLLRYFE PLYTWLQEQN RKSYVGWNTD WSPYSDSRGS GLEVLFQGPG SWSHPQF EK GGGSGGGSGG SAWSHPQFEK UniProtKB: Angiotensin-converting enzyme |
-Macromolecule #2: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  unclassified Coronavirinae (virus) unclassified Coronavirinae (virus) |
| Molecular weight | Theoretical: 141.914672 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MLFFFFLCFA SVNSQCVNLT GRATIQPSFT NSSHRGVYYP DTIFRSNSLV LSQGYFLPFY SNISWYYALT KTNGAEKRVD NPILDFKDG IYFAATEKSN IVRGWIFGTT LDNTSQSLLI VNNATNVIIK VCNFQFCYDP YLSGYFHNNK TWSTREFAVY S SYANCTFE ...String: MLFFFFLCFA SVNSQCVNLT GRATIQPSFT NSSHRGVYYP DTIFRSNSLV LSQGYFLPFY SNISWYYALT KTNGAEKRVD NPILDFKDG IYFAATEKSN IVRGWIFGTT LDNTSQSLLI VNNATNVIIK VCNFQFCYDP YLSGYFHNNK TWSTREFAVY S SYANCTFE YVSKPFMLDI SGKSGLFDTL REFVFRNVDG YFKIYSKYSP VNVNSNLPSG FSALEPLVEL PAGINITRFR TL LTIHRGD PMPNNGWTVF SAAYYVGYLA PRTFMLKYNE NGTITDAVDC SLDPLSEAKC TLKSFTVEKG IYQTSNFRVQ PTD SIVRFP NITNLCPFGE VFNATTFASV YAWNRKRISN CVADYSVLYN STSFSTFKCY GVSPTKLNDL CFTNVYADSF VVRG DEVRQ IAPGQTGKIA DYNYKLPDDF TGCVIAWNSN NLDSKVGGNY NYLYRLFRKS NLKPFERDIS TEIYQAGSTP CNGVE GFNC YFPLKSYGFH PTNGVGYQPY RVVVLSFELL NAPATVCGPK KSTNLIKNKC VNFNFNGLTG TGVLTESNKK FLPFQQ FGR DIADTTDAVR DPQTLEILDI TPCSFGGVSV ITPGTNASNQ VAVLYQDVNC TEVPVAIHAD QLTPTWRVYS TGSNVFQ TR AGCLIGAEHV NNSYECDIPI GAGICASYQT QTNSRSVASQ SIIAYTMSLG AENSVAYSNN SIAIPTNFTI SVTTEILP V SMTKTSVDCT MYICGDSTEC SNLLLQYGSF CTQLNRALTG IAVEQDKNTQ EVFAQVKQIY KTPQIKDFGG FNFSQILPD PSKPSKRSFI EDLLFNKVTL ADAGFIKQYG DCLGDIAARD LICAQKFNGL TVLPPLLTDE MIAQYTSALL AGTITSGWTF GAGAALQIP FAMQMAYRFN GIGVTQNVLY ENQKLIANQF NSAIGKIQDS LSSTASALGK LQDVVNQNAQ ALNTLVKQLS S NFGAISSV LNDILSRLDP PEAEVQIDRL ITGRLQSLQT YVTQQLIRAA EIRASANLAA TKMSECVLGQ SKRVDFCGKG YH LMSFPQS APHGVVFLHV TYVPAQEKNF TTAPAICHDG KAHFPREGVF VSNGTHWFVT QRNFYEPQII TTDNTFVSGN CDV VIGIVN NTVYDPLQPE LDSFKEELDK YFKNHTSPDV DLGDISGINA SVVNIQKEID RLNEVAKNLN ESLIDLQQLG KYEQ LEGSG YIPEAPRDGQ AYVRKDGEWV LLSTFLGRSL EVLFQGPGHH HHHHHHSAWS HPQFEKGGGS GGGGSGGSAW SHPQF EK UniProtKB: Spike glycoprotein |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 5 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 5.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: DARK FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 1.4000000000000001 µm |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.8 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 156892 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)