[English] 日本語
 Yorodumi
Yorodumi- EMDB-32591: Cryo-EM structure of the nucleosome containing Komagataella pasto... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the nucleosome containing Komagataella pastoris histones | ||||||||||||||||||||||||
 Map data Map data | Cryo-EM map of the nucleosome containing Komagataella pastoris histones | ||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||
 Keywords Keywords | Chromatin / Nucleosome / Komagataella pastoris / Histone H2A / Histone H2B / Histone H3 / Histone H4 / DNA / Epigenetic / Gene regulation / cryo-EM / GENE REGULATION-DNA complex | ||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationchromosome, centromeric region / DNA-templated transcription initiation / nucleosome / protein heterodimerization activity / DNA binding / nucleus Similarity search - Function | ||||||||||||||||||||||||
| Biological species |  Komagataella pastoris (fungus) / synthetic construct (others) Komagataella pastoris (fungus) / synthetic construct (others) | ||||||||||||||||||||||||
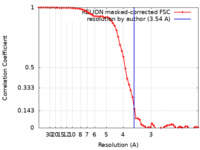
| Method | single particle reconstruction / cryo EM / Resolution: 3.54 Å | ||||||||||||||||||||||||
 Authors Authors | Fukushima Y / Hatazawa S / Hirai S / Kujirai T / Takizawa Y / Kurumizaka H | ||||||||||||||||||||||||
| Funding support |  Japan, 7 items Japan, 7 items
| ||||||||||||||||||||||||
 Citation Citation |  Journal: J Biochem / Year: 2022 Journal: J Biochem / Year: 2022Title: Structural and biochemical analyses of the nucleosome containing Komagataella pastoris histones. Authors: Yutaro Fukushima / Suguru Hatazawa / Seiya Hirai / Tomoya Kujirai / Haruhiko Ehara / Shun-Ichi Sekine / Yoshimasa Takizawa / Hitoshi Kurumizaka /  Abstract: Komagataella pastoris is a methylotrophic yeast that is commonly used as a host cell for protein production. In the present study, we reconstituted the nucleosome with K. pastoris histones and ...Komagataella pastoris is a methylotrophic yeast that is commonly used as a host cell for protein production. In the present study, we reconstituted the nucleosome with K. pastoris histones and determined the structure of the nucleosome core particle by cryogenic electron microscopy. In the K. pastoris nucleosome, the histones form an octamer and the DNA is left-handedly wrapped around it. Micrococcal nuclease assays revealed that the DNA ends of the K. pastoris nucleosome are somewhat more accessible, as compared with those of the human nucleosome. In vitro transcription assays demonstrated that the K. pastoris nucleosome is transcribed by the K. pastoris RNA polymerase II (RNAPII) more efficiently than the human nucleosome, while the RNAPII pausing positions of the K. pastoris nucleosome are the same as those of the human nucleosome. These results suggested that the DNA end flexibility may enhance the transcription efficiency in the nucleosome but minimally affect the nucleosomal pausing positions of RNAPII. | ||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32591.map.gz emd_32591.map.gz | 2.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32591-v30.xml emd-32591-v30.xml emd-32591.xml emd-32591.xml | 18.4 KB 18.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_32591_fsc.xml emd_32591_fsc.xml | 6.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_32591.png emd_32591.png | 124.2 KB | ||
| Filedesc metadata |  emd-32591.cif.gz emd-32591.cif.gz | 5.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32591 http://ftp.pdbj.org/pub/emdb/structures/EMD-32591 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32591 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32591 | HTTPS FTP |
-Validation report
| Summary document |  emd_32591_validation.pdf.gz emd_32591_validation.pdf.gz | 436.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_32591_full_validation.pdf.gz emd_32591_full_validation.pdf.gz | 435.7 KB | Display | |
| Data in XML |  emd_32591_validation.xml.gz emd_32591_validation.xml.gz | 9.6 KB | Display | |
| Data in CIF |  emd_32591_validation.cif.gz emd_32591_validation.cif.gz | 12.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32591 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32591 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32591 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32591 | HTTPS FTP |
-Related structure data
| Related structure data |  7wlrMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_32591.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32591.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM map of the nucleosome containing Komagataella pastoris histones | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : K. pastoris nucleosome
| Entire | Name: K. pastoris nucleosome |
|---|---|
| Components |
|
-Supramolecule #1: K. pastoris nucleosome
| Supramolecule | Name: K. pastoris nucleosome / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Komagataella pastoris (fungus) Komagataella pastoris (fungus) |
-Supramolecule #2: Histone
| Supramolecule | Name: Histone / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#4 |
|---|
-Supramolecule #3: DNA
| Supramolecule | Name: DNA / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #1-#2 |
|---|
-Macromolecule #1: Histone H3
| Macromolecule | Name: Histone H3 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Komagataella pastoris (fungus) Komagataella pastoris (fungus) |
| Molecular weight | Theoretical: 11.601565 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: KPHRYKPGTV ALREIRRFQK STELLIRKLP FQRLVREIAQ DFKTDLRFQS SAIGALQESV EAYLVSLFED TNLCAIHAKR VTIQKKDIL LARRLRGERS UniProtKB: Histone H3 |
-Macromolecule #2: Histone H4
| Macromolecule | Name: Histone H4 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Komagataella pastoris (fungus) Komagataella pastoris (fungus) |
| Molecular weight | Theoretical: 10.030811 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: KRHRKILRDN IQGITKPAIR RLARRGGVKR ISALIYEEVR AVLKTFLENV IRDAVTYTEH AKRKTVTSLD VVYALKRQGR TLYGFGG UniProtKB: Histone H4 |
-Macromolecule #3: Histone H2A
| Macromolecule | Name: Histone H2A / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Komagataella pastoris (fungus) Komagataella pastoris (fungus) |
| Molecular weight | Theoretical: 11.810682 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: KASTSRSSKA GLTFPVGRVH RLLRRGNYAQ RIGSGAPVYL TAVLEYLAAE ILELAGNAAR DNKKSRIIPR HLQLAIRNDE ELNKLLGHV TIAQGGVLPN IQSELLPKK UniProtKB: Histone H2A |
-Macromolecule #4: Histone H2B
| Macromolecule | Name: Histone H2B / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Komagataella pastoris (fungus) Komagataella pastoris (fungus) |
| Molecular weight | Theoretical: 11.018554 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: RSKVRKESYA SYIYKVLKQT HPDTGISQKA MSIMNSFVND IFERIASEAS KLASYNKKST ISAREIQTAV RLILPGELSK HAVSEGTRA VTKYTSSTQA UniProtKB: Histone H2B |
-Macromolecule #5: DNA (145-MER)
| Macromolecule | Name: DNA (145-MER) / type: dna / ID: 5 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 44.520383 KDa |
| Sequence | String: (DA)(DT)(DC)(DA)(DG)(DA)(DA)(DT)(DC)(DC) (DC)(DG)(DG)(DT)(DG)(DC)(DC)(DG)(DA)(DG) (DG)(DC)(DC)(DG)(DC)(DT)(DC)(DA)(DA) (DT)(DT)(DG)(DG)(DT)(DC)(DG)(DT)(DA)(DG) (DA) (DC)(DA)(DG)(DC)(DT)(DC) ...String: (DA)(DT)(DC)(DA)(DG)(DA)(DA)(DT)(DC)(DC) (DC)(DG)(DG)(DT)(DG)(DC)(DC)(DG)(DA)(DG) (DG)(DC)(DC)(DG)(DC)(DT)(DC)(DA)(DA) (DT)(DT)(DG)(DG)(DT)(DC)(DG)(DT)(DA)(DG) (DA) (DC)(DA)(DG)(DC)(DT)(DC)(DT)(DA) (DG)(DC)(DA)(DC)(DC)(DG)(DC)(DT)(DT)(DA) (DA)(DA) (DC)(DG)(DC)(DA)(DC)(DG)(DT) (DA)(DC)(DG)(DC)(DG)(DC)(DT)(DG)(DT)(DC) (DC)(DC)(DC) (DC)(DG)(DC)(DG)(DT)(DT) (DT)(DT)(DA)(DA)(DC)(DC)(DG)(DC)(DC)(DA) (DA)(DG)(DG)(DG) (DG)(DA)(DT)(DT)(DA) (DC)(DT)(DC)(DC)(DC)(DT)(DA)(DG)(DT)(DC) (DT)(DC)(DC)(DA)(DG) (DG)(DC)(DA)(DC) (DG)(DT)(DG)(DT)(DC)(DA)(DG)(DA)(DT)(DA) (DT)(DA)(DT)(DA)(DC)(DA) (DT)(DC)(DG) (DA)(DT) |
-Macromolecule #6: DNA (145-MER)
| Macromolecule | Name: DNA (145-MER) / type: dna / ID: 6 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 44.99166 KDa |
| Sequence | String: (DA)(DT)(DC)(DG)(DA)(DT)(DG)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DC)(DT)(DG)(DA)(DC)(DA) (DC)(DG)(DT)(DG)(DC)(DC)(DT)(DG)(DG) (DA)(DG)(DA)(DC)(DT)(DA)(DG)(DG)(DG)(DA) (DG) (DT)(DA)(DA)(DT)(DC)(DC) ...String: (DA)(DT)(DC)(DG)(DA)(DT)(DG)(DT)(DA)(DT) (DA)(DT)(DA)(DT)(DC)(DT)(DG)(DA)(DC)(DA) (DC)(DG)(DT)(DG)(DC)(DC)(DT)(DG)(DG) (DA)(DG)(DA)(DC)(DT)(DA)(DG)(DG)(DG)(DA) (DG) (DT)(DA)(DA)(DT)(DC)(DC)(DC)(DC) (DT)(DT)(DG)(DG)(DC)(DG)(DG)(DT)(DT)(DA) (DA)(DA) (DA)(DC)(DG)(DC)(DG)(DG)(DG) (DG)(DG)(DA)(DC)(DA)(DG)(DC)(DG)(DC)(DG) (DT)(DA)(DC) (DG)(DT)(DG)(DC)(DG)(DT) (DT)(DT)(DA)(DA)(DG)(DC)(DG)(DG)(DT)(DG) (DC)(DT)(DA)(DG) (DA)(DG)(DC)(DT)(DG) (DT)(DC)(DT)(DA)(DC)(DG)(DA)(DC)(DC)(DA) (DA)(DT)(DT)(DG)(DA) (DG)(DC)(DG)(DG) (DC)(DC)(DT)(DC)(DG)(DG)(DC)(DA)(DC)(DC) (DG)(DG)(DG)(DA)(DT)(DT) (DC)(DT)(DG) (DA)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.3 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 57.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.25 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller