+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The structure of KATP H175K mutant in closed state | |||||||||
 Map data Map data | composite map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | KATP / Kir6.2 / SUR1 / NN414 / ADP / ATP / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationATP sensitive Potassium channels / response to resveratrol / ATP-activated inward rectifier potassium channel activity / glutamate secretion, neurotransmission / inward rectifying potassium channel / Regulation of insulin secretion / sulfonylurea receptor activity / cell body fiber / ventricular cardiac muscle tissue development / ABC-family proteins mediated transport ...ATP sensitive Potassium channels / response to resveratrol / ATP-activated inward rectifier potassium channel activity / glutamate secretion, neurotransmission / inward rectifying potassium channel / Regulation of insulin secretion / sulfonylurea receptor activity / cell body fiber / ventricular cardiac muscle tissue development / ABC-family proteins mediated transport / CAMKK-AMPK signaling cascade / voltage-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential / Ion homeostasis / ATPase-coupled monoatomic cation transmembrane transporter activity / inward rectifier potassium channel activity / regulation of monoatomic ion transmembrane transport / nervous system process / inorganic cation transmembrane transport / response to stress / ankyrin binding / neuromuscular process / response to ATP / potassium ion import across plasma membrane / response to testosterone / action potential / potassium ion binding / positive regulation of insulin secretion involved in cellular response to glucose stimulus / intercalated disc / potassium channel activity / axolemma / ABC-type transporter activity / negative regulation of insulin secretion / cellular response to nutrient levels / heat shock protein binding / T-tubule / acrosomal vesicle / response to ischemia / determination of adult lifespan / positive regulation of protein localization to plasma membrane / cellular response to glucose stimulus / ADP binding / potassium ion transport / sarcolemma / cellular response to nicotine / glucose metabolic process / cellular response to tumor necrosis factor / nuclear envelope / response to estradiol / presynapse / presynaptic membrane / transmembrane transporter binding / response to hypoxia / endosome / response to xenobiotic stimulus / neuronal cell body / glutamatergic synapse / apoptotic process / ATP hydrolysis activity / ATP binding / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Mesocricetus auratus (golden hamster) Mesocricetus auratus (golden hamster) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.19 Å | |||||||||
 Authors Authors | Chen L / Wang M | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structural insights into the mechanism of pancreatic K channel regulation by nucleotides. Authors: Mengmeng Wang / Jing-Xiang Wu / Dian Ding / Lei Chen /  Abstract: ATP-sensitive potassium channels (K) are metabolic sensors that convert the intracellular ATP/ADP ratio to the excitability of cells. They are involved in many physiological processes and implicated ...ATP-sensitive potassium channels (K) are metabolic sensors that convert the intracellular ATP/ADP ratio to the excitability of cells. They are involved in many physiological processes and implicated in several human diseases. Here we present the cryo-EM structures of the pancreatic K channel in both the closed state and the pre-open state, resolved in the same sample. We observe the binding of nucleotides at the inhibitory sites of the Kir6.2 channel in the closed but not in the pre-open state. Structural comparisons reveal the mechanism for ATP inhibition and Mg-ADP activation, two fundamental properties of K channels. Moreover, the structures also uncover the activation mechanism of diazoxide-type K openers. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32311.map.gz emd_32311.map.gz | 17.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32311-v30.xml emd-32311-v30.xml emd-32311.xml emd-32311.xml | 14.1 KB 14.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_32311.png emd_32311.png | 132.2 KB | ||
| Filedesc metadata |  emd-32311.cif.gz emd-32311.cif.gz | 6.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32311 http://ftp.pdbj.org/pub/emdb/structures/EMD-32311 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32311 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32311 | HTTPS FTP |
-Validation report
| Summary document |  emd_32311_validation.pdf.gz emd_32311_validation.pdf.gz | 604.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_32311_full_validation.pdf.gz emd_32311_full_validation.pdf.gz | 604.2 KB | Display | |
| Data in XML |  emd_32311_validation.xml.gz emd_32311_validation.xml.gz | 5.5 KB | Display | |
| Data in CIF |  emd_32311_validation.cif.gz emd_32311_validation.cif.gz | 6.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32311 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32311 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32311 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32311 | HTTPS FTP |
-Related structure data
| Related structure data |  7w4pMC  7w4oC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_32311.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32311.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | composite map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.324 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Kir6.2 H175K mutant in complex with SUR1
| Entire | Name: Kir6.2 H175K mutant in complex with SUR1 |
|---|---|
| Components |
|
-Supramolecule #1: Kir6.2 H175K mutant in complex with SUR1
| Supramolecule | Name: Kir6.2 H175K mutant in complex with SUR1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #2: Kir6.2 H175K mutant
| Supramolecule | Name: Kir6.2 H175K mutant / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Mesocricetus auratus (golden hamster) Mesocricetus auratus (golden hamster) |
-Supramolecule #3: SUR1
| Supramolecule | Name: SUR1 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|
-Macromolecule #1: ATP-sensitive inward rectifier potassium channel 11
| Macromolecule | Name: ATP-sensitive inward rectifier potassium channel 11 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 43.60677 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MLSRKGIIPE EYVLTRLAED PAEPRYRTRE RRARFVSKKG NCNVAHKNIR EQGRFLQDVF TTLVDLKWPH TLLIFTMSFL CSWLLFAMV WWLIAFAHGD LAPGEGTNVP CVTSIHSFSS AFLFSIEVQV TIGFGGRMVT EECPLAILIL IVQNIVGLMI N AIMLGCIF ...String: MLSRKGIIPE EYVLTRLAED PAEPRYRTRE RRARFVSKKG NCNVAHKNIR EQGRFLQDVF TTLVDLKWPH TLLIFTMSFL CSWLLFAMV WWLIAFAHGD LAPGEGTNVP CVTSIHSFSS AFLFSIEVQV TIGFGGRMVT EECPLAILIL IVQNIVGLMI N AIMLGCIF MKTAQAKRRA ETLIFSKHAV ITLRHGRLCF MLRVGDLRKS MIISATIHMQ VVRKTTSPEG EVVPLHQVDI PM ENGVGGN GIFLVAPLII YHVIDSNSPL YDLAPSDLHH HQDLEIIVIL EGVVETTGIT TQARTSYLAD EILWGQRFVP IVA EEDGRY SVDYSKFGNT IKVPTPLCTA RQLDEDRSLL DALTLASSRG PLRKRSVAVA KAKPKFSISP DSLS UniProtKB: ATP-sensitive inward rectifier potassium channel 11 |
-Macromolecule #2: ATP-binding cassette sub-family C member 8 isoform X2
| Macromolecule | Name: ATP-binding cassette sub-family C member 8 isoform X2 / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Mesocricetus auratus (golden hamster) Mesocricetus auratus (golden hamster) |
| Molecular weight | Theoretical: 177.295516 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MPLAFCGTEN HSAAYRVDQG VLNNGCFVDA LNVVPHVFLL FITFPILFIG WGSQSSKVHI HHSTWLHFPG HNLRWILTFI LLFVLVCEI AEGILSDGVT ESRHLHLYMP AGMAFMAAIT SVVYYHNIET SNFPKLLIAL LIYWTLAFIT KTIKFVKFYD H AIGFSQLR ...String: MPLAFCGTEN HSAAYRVDQG VLNNGCFVDA LNVVPHVFLL FITFPILFIG WGSQSSKVHI HHSTWLHFPG HNLRWILTFI LLFVLVCEI AEGILSDGVT ESRHLHLYMP AGMAFMAAIT SVVYYHNIET SNFPKLLIAL LIYWTLAFIT KTIKFVKFYD H AIGFSQLR FCLTGLLVIL YGMLLLVEVN VIRVRRYIFF KTPREVKPPE DLQDLGVRFL QPFVNLLSKG TYWWMNAFIK TA HKKPIDL RAIGKLPIAM RALTNYQRLC VAFDAQARKD TQSPQGARAI WRALCHAFGR RLILSSTFRI LADLLGFAGP LCI FGIVDH LGKENHVFQP KTQFLGVYFV SSQEFLGNAY VLAVLLFLAL LLQRTFLQAS YYVAIETGIN LRGAIQTKIY NKIM HLSTS NLSMGEMTAG QICNLVAIDT NQLMWFFFLC PNLWAMPVQI IVGVILLYYI LGVSALIGAA VIILLAPVQY FVATK LSQA QRSTLEHSNE RLKQTNEMLR GMKLLKLYAW ESIFCSRVEV TRRKEMTSLR AFAVYTSISI FMNTAIPIAA VLITFV GHV SFFKESDLSP SVAFASLSLF HILVTPLFLL SSVVRSTVKA LVSVQKLSEF LSSAEIREEQ CAPREPAPQG QAGKYQA VP LKVVNRKRPA REEVRDLLGP LQRLAPSMDG DADNFCVQII GGFFTWTPDG IPTLSNITIR IPRGQLTMIV GQVGCGKS S LLLATLGEMQ KVSGAVFWNS NLPDSEGEDP SSPERETAAG SDIRSRGPVA YASQKPWLLN ATVEENITFE SPFNKQRYK MVIEACSLQP DIDILPHGDQ TQIGERGINL SGGQRQRISV ARALYQQTNV VFLDDPFSAL DVHLSDHLMQ AGILELLRDD KRTVVLVTH KLQYLPHADW IIAMKDGTIQ REGTLKDFQR SECQLFEHWK TLMNRQDQEL EKETVMERKA SEPSQGLPRA M SSRDGLLL DEEEEEEEAA ESEEDDNLSS VLHQRAKIPW RACTKYLSSA GILLLSLLVF SQLLKHMVLV AIDYWLAKWT DS ALVLSPA ARNCSLSQEC DLDQSVYAMV FTLLCSLGIV LCLVTSVTVE WTGLKVAKRL HRSLLNRIIL APMRFFETTP LGS ILNRFS SDCNTIDQHI PSTLECLSRS TLLCVSALTV ISYVTPVFLV ALLPLAVVCY FIQKYFRVAS RDLQQLDDTT QLPL LSHFA ETVEGLTTIR AFRYEARFQQ KLLEYTDSNN IASLFLTAAN RWLEVRMEYI GACVVLIAAA TSISNSLHRE LSAGL VGLG LTYALMVSNY LNWMVRNLAD MEIQLGAVKR IHALLKTEAE SYEGLLAPSL IPKNWPDQGK IQIQNLSVRY DSSLKP VLK HVNALISPGQ KIGICGRTGS GKSSFSLAFF RMVDMFEGRI IIDGIDIAKL PLHTLRSRLS IILQDPVLFS GTIRFNL DP EKKCSDSTLW EALEIAQLKL VVKALPGGLD AIITEGGENF SQGQRQLFCL ARAFVRKTSI FIMDEATASI DMATENIL Q KVVMTAFADR TVVTIAHRVH TILSADLVMV LKRGAILEFD KPETLLSQKD SVFASFVRAD K UniProtKB: ATP-binding cassette sub-family C member 8 isoform X2 |
-Macromolecule #3: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 8 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 8 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #5: 6-chloranyl-~{N}-(1-methylcyclopropyl)-1,1-bis(oxidanylidene)-4~{...
| Macromolecule | Name: 6-chloranyl-~{N}-(1-methylcyclopropyl)-1,1-bis(oxidanylidene)-4~{H}-thieno[3,2-e][1,2,4]thiadiazin-3-amine type: ligand / ID: 5 / Number of copies: 4 / Formula: E2H |
|---|---|
| Molecular weight | Theoretical: 291.778 Da |
| Chemical component information | 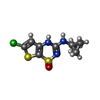 ChemComp-E2H: |
-Macromolecule #6: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 6 / Number of copies: 4 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.19 Å / Resolution method: FSC 0.143 CUT-OFF Details: This is a composite map. Resolutions for each individual map are "3.16/3.19/2.72" respectively Number images used: 216000 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: OTHER |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)