+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30372 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of P.aeruginosa MlaFEBD | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Mla complex / Lipid transporter / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationresponse to nitrite / phospholipid transporter activity / phospholipid transport / ATP-binding cassette (ABC) transporter complex / phospholipid binding / ATP hydrolysis activity / ATP binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Pseudomonas aeruginosa PAO1 (bacteria) / Pseudomonas aeruginosa PAO1 (bacteria) /  Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) | |||||||||
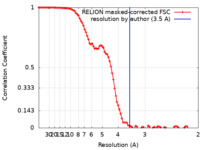
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Zhou C / Shi H | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2021 Journal: J Mol Biol / Year: 2021Title: Structural Insight into Phospholipid Transport by the MlaFEBD Complex from P. aeruginosa. Authors: Changping Zhou / Huigang Shi / Manfeng Zhang / Lijun Zhou / Le Xiao / Shasha Feng / Wonpil Im / Min Zhou / Xinzheng Zhang / Yihua Huang /   Abstract: The outer membrane (OM) of Gram-negative bacteria, which consists of lipopolysaccharides (LPS) in the outer leaflet and phospholipids (PLs) in the inner leaflet, plays a key role in antibiotic ...The outer membrane (OM) of Gram-negative bacteria, which consists of lipopolysaccharides (LPS) in the outer leaflet and phospholipids (PLs) in the inner leaflet, plays a key role in antibiotic resistance and pathogen virulence. The maintenance of lipid asymmetry (Mla) pathway is known to be involved in PL transport and contributes to the lipid homeostasis of the OM, yet the underlying molecular mechanism and the directionality of PL transport in this pathway remain elusive. Here, we reported the cryo-EM structures of the ATP-binding cassette (ABC) transporter MlaFEBD from P. areuginosa, the core complex in the Mla pathway, in nucleotide-free (apo)-, ADP (ATP + vanadate)- and ATP (AMPPNP)-bound states as well as the structures of MlaFEB from E. coli in apo- and AMPPNP-bound states at a resolution range of 3.4-3.9 Å. The structures show that the MlaFEBD complex contains a total of twelve protein molecules with a stoichiometry of MlaFEBD, and binds a plethora of PLs at different locations. In contrast to canonical ABC transporters, nucleotide binding fails to trigger significant conformational changes of both MlaFEBD and MlaFEB in the nucleotide-binding and transmembrane domains of the ABC transporter, correlated with their low ATPase activities exhibited in both detergent micelles and lipid nanodiscs. Intriguingly, PLs or detergents appeared to relocate to the membrane-proximal end from the distal end of the hydrophobic tunnel formed by the MlaD hexamer in MlaFEBD upon addition of ATP, indicating that retrograde PL transport might occur in the tunnel in an ATP-dependent manner. Site-specific photocrosslinking experiment confirms that the substrate-binding pocket in the dimeric MlaE and the MlaD hexamer are able to bind PLs in vitro, in line with the notion that MlaFEBD complex functions as a PL transporter. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30372.map.gz emd_30372.map.gz | 59.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30372-v30.xml emd-30372-v30.xml emd-30372.xml emd-30372.xml | 15.4 KB 15.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_30372_fsc.xml emd_30372_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_30372.png emd_30372.png | 57.7 KB | ||
| Filedesc metadata |  emd-30372.cif.gz emd-30372.cif.gz | 5.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30372 http://ftp.pdbj.org/pub/emdb/structures/EMD-30372 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30372 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30372 | HTTPS FTP |
-Validation report
| Summary document |  emd_30372_validation.pdf.gz emd_30372_validation.pdf.gz | 566.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_30372_full_validation.pdf.gz emd_30372_full_validation.pdf.gz | 565.6 KB | Display | |
| Data in XML |  emd_30372_validation.xml.gz emd_30372_validation.xml.gz | 10.3 KB | Display | |
| Data in CIF |  emd_30372_validation.cif.gz emd_30372_validation.cif.gz | 13.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30372 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30372 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30372 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30372 | HTTPS FTP |
-Related structure data
| Related structure data |  7ch9MC  7ch6C  7ch7C  7ch8C  7chaC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30372.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30372.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Cryo-EM structure of P.aeruginosa MlaFEBD
| Entire | Name: Cryo-EM structure of P.aeruginosa MlaFEBD |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of P.aeruginosa MlaFEBD
| Supramolecule | Name: Cryo-EM structure of P.aeruginosa MlaFEBD / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa PAO1 (bacteria) Pseudomonas aeruginosa PAO1 (bacteria) |
-Macromolecule #1: MlaD domain-containing protein
| Macromolecule | Name: MlaD domain-containing protein / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria)Strain: ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1 |
| Molecular weight | Theoretical: 16.571945 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MQTRTLEIGV GLFLLAGLLA LLLLALRVSG LSVGNAGDTY KVYAYFDNIA GVTVRGKVTL AGVTIGKVTA VDLDRDSYTG RVTMEINQN VNNLPVDSTA SILTAGLLGE KYIGISVGGD EDVLKDGSTI HDTQSALVLE DLIGKFLLNS VNKDEAKK UniProtKB: Mce/MlaD domain-containing protein |
-Macromolecule #2: Probable permease of ABC transporter
| Macromolecule | Name: Probable permease of ABC transporter / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria)Strain: ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1 |
| Molecular weight | Theoretical: 28.400326 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MRRVSPLERI RLFGRAGLDV VAALGRSTLF LGHALLGRRT PGTGLHLLVK QLYSVGVLSL AIIVVSGLFI GMVLALQGYN ILISYGSEQ AVGQMVALTL LRELGPVVTG LLFAGRAGSA LTAEIGNMKA TEQLSSLEMI GVDPLKYIVA PRLWAGFISM P LLAAIFSV ...String: MRRVSPLERI RLFGRAGLDV VAALGRSTLF LGHALLGRRT PGTGLHLLVK QLYSVGVLSL AIIVVSGLFI GMVLALQGYN ILISYGSEQ AVGQMVALTL LRELGPVVTG LLFAGRAGSA LTAEIGNMKA TEQLSSLEMI GVDPLKYIVA PRLWAGFISM P LLAAIFSV VGIWGGAMVA VDWLGVYEGS FWANMQNSVQ FTEDVLNGVI KSIVFAFVVT WIAVYQGYDC EPTSEGISRA TT RTVVYAS LAVLGLDFIL TALMFGDF UniProtKB: Intermembrane phospholipid transport system permease protein MlaE |
-Macromolecule #3: Probable ATP-binding component of ABC transporter
| Macromolecule | Name: Probable ATP-binding component of ABC transporter / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria)Strain: ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1 |
| Molecular weight | Theoretical: 29.657367 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSTDSAYAVE LKGLTFKRGS RAIFDNIDVR IPRGKVTGIM GPSGCGKTTL LRLIASQLRP SKGEVWVNGQ NLPQLSRGDL FDMRKQFGV LFQSGALFTD LDVFENVAFP LRVHTQLPEE MIRDIVLMKL QAVGLRGAVE LMPDELSGGM KRRVALARAI A LDPQILLY ...String: MSTDSAYAVE LKGLTFKRGS RAIFDNIDVR IPRGKVTGIM GPSGCGKTTL LRLIASQLRP SKGEVWVNGQ NLPQLSRGDL FDMRKQFGV LFQSGALFTD LDVFENVAFP LRVHTQLPEE MIRDIVLMKL QAVGLRGAVE LMPDELSGGM KRRVALARAI A LDPQILLY DEPFVGQDPI AMGVLVRLIR LLNDALGITS IVVSHDLAET ASIADYIYIV GDGRVLGHGT PDVLKETDDP RI RQFVKGI PDGPVPFHYP ARDYRADLLG ER UniProtKB: Probable ATP-binding component of ABC transporter |
-Macromolecule #4: STAS domain-containing protein
| Macromolecule | Name: STAS domain-containing protein / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria) Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1) (bacteria)Strain: ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1 |
| Molecular weight | Theoretical: 10.789388 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSQASLREGA AGELQLAGVL DYSSGPALRE QGGRLIRASQ AAELVVDCSA VERSSSVGIS LLLAFIRDAR KAGKVLSVRA LPDDMREIA KVSSLLEILP LQE UniProtKB: STAS domain-containing protein |
-Macromolecule #5: 2-(HEXADECANOYLOXY)-1-[(PHOSPHONOOXY)METHYL]ETHYL HEXADECANOATE
| Macromolecule | Name: 2-(HEXADECANOYLOXY)-1-[(PHOSPHONOOXY)METHYL]ETHYL HEXADECANOATE type: ligand / ID: 5 / Number of copies: 10 / Formula: LPP |
|---|---|
| Molecular weight | Theoretical: 648.891 Da |
| Chemical component information |  ChemComp-LPP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source: TUNGSTEN HAIRPIN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: SPOT SCAN / Imaging mode: DIFFRACTION / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller



 UCSF Chimera
UCSF Chimera
















 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)