+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | G4 RNA-mediated PRC2 dimer | |||||||||
 Map data Map data | consensus map of G4 RNA-mediated PRC2 dimer | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | PRC2 / G-quadruplex RNA / RNP complex / chromatin modifier / GENE REGULATION | |||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of kidney development / hepatocyte homeostasis / cellular response to trichostatin A / regulation of gliogenesis / negative regulation of striated muscle cell differentiation / [histone H3]-lysine27 N-trimethyltransferase / sex chromatin / negative regulation of keratinocyte differentiation / random inactivation of X chromosome / CAF-1 complex ...regulation of kidney development / hepatocyte homeostasis / cellular response to trichostatin A / regulation of gliogenesis / negative regulation of striated muscle cell differentiation / [histone H3]-lysine27 N-trimethyltransferase / sex chromatin / negative regulation of keratinocyte differentiation / random inactivation of X chromosome / CAF-1 complex / histone H3K27 trimethyltransferase activity / negative regulation of retinoic acid receptor signaling pathway / regulatory ncRNA-mediated heterochromatin formation / skeletal muscle satellite cell maintenance involved in skeletal muscle regeneration / response to tetrachloromethane / cerebellar cortex development / primary miRNA binding / histone H3K27 methyltransferase activity / facultative heterochromatin formation / positive regulation of cell cycle G1/S phase transition / NURF complex / NuRD complex / regulation of cell fate specification / negative regulation of stem cell population maintenance / DNA replication-dependent chromatin assembly / ESC/E(Z) complex / chromatin silencing complex / Transcription of E2F targets under negative control by p107 (RBL1) and p130 (RBL2) in complex with HDAC1 / regulation of stem cell differentiation / RSC-type complex / protein-lysine N-methyltransferase activity / cardiac muscle hypertrophy in response to stress / negative regulation of stem cell differentiation / Transcription of E2F targets under negative control by DREAM complex / pronucleus / Polo-like kinase mediated events / synaptic transmission, GABAergic / positive regulation of dendrite development / histone H3 methyltransferase activity / lncRNA binding / negative regulation of G1/S transition of mitotic cell cycle / negative regulation of gene expression, epigenetic / G1 to G0 transition / spinal cord development / positive regulation of stem cell population maintenance / ATPase complex / G1/S-Specific Transcription / histone methyltransferase activity / Sin3-type complex / oligodendrocyte differentiation / negative regulation of transcription elongation by RNA polymerase II / Transcriptional Regulation by E2F6 / RNA Polymerase I Transcription Initiation / negative regulation of cell differentiation / histone deacetylase complex / G0 and Early G1 / subtelomeric heterochromatin formation / negative regulation of cytokine production involved in inflammatory response / RNA polymerase II core promoter sequence-specific DNA binding / pericentric heterochromatin / ribonucleoprotein complex binding / Cyclin E associated events during G1/S transition / Transcriptional regulation of brown and beige adipocyte differentiation by EBF2 / positive regulation of epithelial to mesenchymal transition / Cyclin A:Cdk2-associated events at S phase entry / keratinocyte differentiation / Deposition of new CENPA-containing nucleosomes at the centromere / enzyme activator activity / Regulation of TP53 Activity through Acetylation / protein localization to chromatin / methylated histone binding / B cell differentiation / SUMOylation of chromatin organization proteins / transcription corepressor binding / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / positive regulation of GTPase activity / negative regulation of cell migration / PRC2 methylates histones and DNA / Regulation of PTEN gene transcription / stem cell differentiation / Defective pyroptosis / HDACs deacetylate histones / liver regeneration / promoter-specific chromatin binding / hippocampus development / transcription coregulator activity / negative regulation of transforming growth factor beta receptor signaling pathway / positive regulation of MAP kinase activity / protein modification process / positive regulation of protein serine/threonine kinase activity / brain development / regulation of circadian rhythm / heterochromatin formation / chromatin DNA binding / PKMTs methylate histone lysines / histone deacetylase binding / Activation of anterior HOX genes in hindbrain development during early embryogenesis / HCMV Early Events / cellular response to hydrogen peroxide / G1/S transition of mitotic cell cycle Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
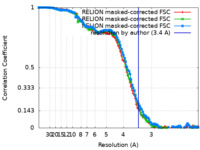
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Song J / Kasinath V | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2023 Journal: Science / Year: 2023Title: Structural basis for inactivation of PRC2 by G-quadruplex RNA. Authors: Jiarui Song / Anne R Gooding / Wayne O Hemphill / Brittney D Love / Anne Robertson / Liqi Yao / Leonard I Zon / Trista E North / Vignesh Kasinath / Thomas R Cech /  Abstract: Polycomb repressive complex 2 (PRC2) silences genes through trimethylation of histone H3K27. PRC2 associates with numerous precursor messenger RNAs (pre-mRNAs) and long noncoding RNAs (lncRNAs) with ...Polycomb repressive complex 2 (PRC2) silences genes through trimethylation of histone H3K27. PRC2 associates with numerous precursor messenger RNAs (pre-mRNAs) and long noncoding RNAs (lncRNAs) with a binding preference for G-quadruplex RNA. In this work, we present a 3.3-Å-resolution cryo-electron microscopy structure of PRC2 bound to a G-quadruplex RNA. Notably, RNA mediates the dimerization of PRC2 by binding both protomers and inducing a protein interface composed of two copies of the catalytic subunit EZH2, thereby blocking nucleosome DNA interaction and histone H3 tail accessibility. Furthermore, an RNA-binding loop of EZH2 facilitates the handoff between RNA and DNA, another activity implicated in PRC2 regulation by RNA. We identified a gain-of-function mutation in this loop that activates PRC2 in zebrafish. Our results reveal mechanisms for RNA-mediated regulation of a chromatin-modifying enzyme. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29578.map.gz emd_29578.map.gz | 80.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29578-v30.xml emd-29578-v30.xml emd-29578.xml emd-29578.xml | 31.6 KB 31.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29578_fsc.xml emd_29578_fsc.xml emd_29578_fsc_2.xml emd_29578_fsc_2.xml emd_29578_fsc_3.xml emd_29578_fsc_3.xml | 10.7 KB 10.7 KB 10.6 KB | Display Display Display |  FSC data file FSC data file |
| Images |  emd_29578.png emd_29578.png | 117.5 KB | ||
| Masks |  emd_29578_msk_1.map emd_29578_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-29578.cif.gz emd-29578.cif.gz | 9.7 KB | ||
| Others |  emd_29578_additional_1.map.gz emd_29578_additional_1.map.gz emd_29578_half_map_1.map.gz emd_29578_half_map_1.map.gz emd_29578_half_map_2.map.gz emd_29578_half_map_2.map.gz | 8.3 MB 80.9 MB 80.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29578 http://ftp.pdbj.org/pub/emdb/structures/EMD-29578 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29578 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29578 | HTTPS FTP |
-Validation report
| Summary document |  emd_29578_validation.pdf.gz emd_29578_validation.pdf.gz | 877.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_29578_full_validation.pdf.gz emd_29578_full_validation.pdf.gz | 877.1 KB | Display | |
| Data in XML |  emd_29578_validation.xml.gz emd_29578_validation.xml.gz | 17.9 KB | Display | |
| Data in CIF |  emd_29578_validation.cif.gz emd_29578_validation.cif.gz | 23.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29578 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29578 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29578 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-29578 | HTTPS FTP |
-Related structure data
| Related structure data |  8fyhMC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29578.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29578.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | consensus map of G4 RNA-mediated PRC2 dimer | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06344 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_29578_msk_1.map emd_29578_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: consensus map of G4 RNA-mediated PRC2 dimer after...
| File | emd_29578_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | consensus map of G4 RNA-mediated PRC2 dimer after postprocessing with B-factor of -40 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2 of consensus map
| File | emd_29578_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 of consensus map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1 of consensus map
| File | emd_29578_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 of consensus map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : G4 RNA-mediated dimer of polycomb repressive complex 2
| Entire | Name: G4 RNA-mediated dimer of polycomb repressive complex 2 |
|---|---|
| Components |
|
-Supramolecule #1: G4 RNA-mediated dimer of polycomb repressive complex 2
| Supramolecule | Name: G4 RNA-mediated dimer of polycomb repressive complex 2 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#7 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 800 KDa |
-Macromolecule #1: Polycomb protein SUZ12
| Macromolecule | Name: Polycomb protein SUZ12 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 83.181922 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MAPQKHGGGG GGGSGPSAGS GGGGFGGSAA VAAATASGGK SGGGSCGGGG SYSASSSSSA AAAAGAAVLP VKKPKMEHVQ ADHELFLQA FEKPTQIYRF LRTRNLIAPI FLHRTLTYMS HRNSRTNIKR KTFKVDDMLS KVEKMKGEQE SHSLSAHLQL T FTGFFHKN ...String: MAPQKHGGGG GGGSGPSAGS GGGGFGGSAA VAAATASGGK SGGGSCGGGG SYSASSSSSA AAAAGAAVLP VKKPKMEHVQ ADHELFLQA FEKPTQIYRF LRTRNLIAPI FLHRTLTYMS HRNSRTNIKR KTFKVDDMLS KVEKMKGEQE SHSLSAHLQL T FTGFFHKN DKPSPNSENE QNSVTLEVLL VKVCHKKRKD VSCPIRQVPT GKKQVPLNPD LNQTKPGNFP SLAVSSNEFE PS NSHMVKS YSLLFRVTRP GRREFNGMIN GETNENIDVN EELPARRKRN REDGEKTFVA QMTVFDKNRR LQLLDGEYEV AMQ EMEECP ISKKRATWET ILDGKRLPPF ETFSQGPTLQ FTLRWTGETN DKSTAPIAKP LATRNSESLH QENKPGSVKP TQTI AVKES LTTDLQTRKE KDTPNENRQK LRIFYQFLYN NNTRQQTEAR DDLHCPWCTL NCRKLYSLLK HLKLCHSRFI FNYVY HPKG ARIDVSINEC YDGSYAGNPQ DIHRQPGFAF SRNGPVKRTP ITHILVCRPK RTKASMSEFL ESEDGEVEQQ RTYSSG HNR LYFHSDTCLP LRPQEMEVDS EDEKDPEWLR EKTITQIEEF SDVNEGEKEV MKLWNLHVMK HGFIADNQMN HACMLFV EN YGQKIIKKNL CRNFMLHLVS MHDFNLISIM SIDKAVTKLR EMQQKLEKGE SASPANEEIT EEQNGTANGF SEINSKEK A LETDSVSGVS KQSKKQKL UniProtKB: Polycomb protein SUZ12 |
-Macromolecule #2: Polycomb protein EED
| Macromolecule | Name: Polycomb protein EED / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 50.267691 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MSEREVSTAP AGTDMPAAKK QKLSSDENSN PDLSGDENDD AVSIESGTNT ERPDTPTNTP NAPGRKSWGK GKWKSKKCKY SFKCVNSLK EDHNQPLFGV QFNWHSKEGD PLVFATVGSN RVTLYECHSQ GEIRLLQSYV DADADENFYT CAWTYDSNTS H PLLAVAGS ...String: MSEREVSTAP AGTDMPAAKK QKLSSDENSN PDLSGDENDD AVSIESGTNT ERPDTPTNTP NAPGRKSWGK GKWKSKKCKY SFKCVNSLK EDHNQPLFGV QFNWHSKEGD PLVFATVGSN RVTLYECHSQ GEIRLLQSYV DADADENFYT CAWTYDSNTS H PLLAVAGS RGIIRIINPI TMQCIKHYVG HGNAINELKF HPRDPNLLLS VSKDHALRLW NIQTDTLVAI FGGVEGHRDE VL SADYDLL GEKIMSCGMD HSLKLWRINS KRMMNAIKES YDYNPNKTNR PFISQKIHFP DFSTRDIHRN YVDCVRWLGD LIL SKSCEN AIVCWKPGKM EDDIDKIKPS ESNVTILGRF DYSQCDIWYM RFSMDFWQKM LALGNQVGKL YVWDLEVEDP HKAK CTTLT HHKCGAAIRQ TSFSRDSSIL IAVCDDASIW RWDRLR UniProtKB: Polycomb protein EED |
-Macromolecule #3: Histone-binding protein RBBP4
| Macromolecule | Name: Histone-binding protein RBBP4 / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 47.709527 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MADKEAAFDD AVEERVINEE YKIWKKNTPF LYDLVMTHAL EWPSLTAQWL PDVTRPEGKD FSIHRLVLGT HTSDEQNHLV IASVQLPND DAQFDASHYD SEKGEFGGFG SVSGKIEIEI KINHEGEVNR ARYMPQNPCI IATKTPSSDV LVFDYTKHPS K PDPSGECN ...String: MADKEAAFDD AVEERVINEE YKIWKKNTPF LYDLVMTHAL EWPSLTAQWL PDVTRPEGKD FSIHRLVLGT HTSDEQNHLV IASVQLPND DAQFDASHYD SEKGEFGGFG SVSGKIEIEI KINHEGEVNR ARYMPQNPCI IATKTPSSDV LVFDYTKHPS K PDPSGECN PDLRLRGHQK EGYGLSWNPN LSGHLLSASD DHTICLWDIS AVPKEGKVVD AKTIFTGHTA VVEDVSWHLL HE SLFGSVA DDQKLMIWDT RSNNTSKPSH SVDAHTAEVN CLSFNPYSEF ILATGSADKT VALWDLRNLK LKLHSFESHK DEI FQVQWS PHNETILASS GTDRRLNVWD LSKIGEEQSP EDAEDGPPEL LFIHGGHTAK ISDFSWNPNE PWVICSVSED NIMQ VWQMA ENIYNDEDPE GSVDPEGQGS UniProtKB: Histone-binding protein RBBP4 |
-Macromolecule #4: Histone-lysine N-methyltransferase EZH2
| Macromolecule | Name: Histone-lysine N-methyltransferase EZH2 / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO / EC number: [histone H3]-lysine27 N-trimethyltransferase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 86.149055 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MGQTGKKSEK GPVCWRKRVK SEYMRLRQLK RFRRADEVKS MFSSNRQKIL ERTEILNQEW KQRRIQPVHI LTSVSSLRGT RECSVTSDL DFPTQVIPLK TLNAVASVPI MYSWSPLQQN FMVEDETVLH NIPYMGDEVL DQDGTFIEEL IKNYDGKVHG D RECGFIND ...String: MGQTGKKSEK GPVCWRKRVK SEYMRLRQLK RFRRADEVKS MFSSNRQKIL ERTEILNQEW KQRRIQPVHI LTSVSSLRGT RECSVTSDL DFPTQVIPLK TLNAVASVPI MYSWSPLQQN FMVEDETVLH NIPYMGDEVL DQDGTFIEEL IKNYDGKVHG D RECGFIND EIFVELVNAL GQYNDDDDDD DGDDPEEREE KQKDLEDHRD DKESRPPRKF PSDKIFEAIS SMFPDKGTAE EL KEKYKEL TEQQLPGALP PECTPNIDGP NAKSVQREQS LHSFHTLFCR RCFKYDCFLH RKCNYSFHAT PNTYKRKNTE TAL DNKPCG PQCYQHLEGA KEFAAALTAE RIKTPPKRPG GRRRGRLPNN SSRPSTPTIN VLESKDTDSD REAGTETGGE NNDK EEEEK KDETSSSSEA NSRCQTPIKM KPNIEPPENV EWSGAEASMF RVLIGTYYDN FCAIARLIGT KTCRQVYEFR VKESS IIAP APAEDVDTPP RKKKRKHRLW AAHCRKIQLK KDGSSNHVYN YQPCDHPRQP CDSSCPCVIA QNFCEKFCQC SSECQN RFP GCRCKAQCNT KQCPCYLAVR ECDPDLCLTC GAADHWDSKN VSCKNCSIQR GSKKHLLLAP SDVAGWGIFI KDPVQKN EF ISEYCGEIIS QDEADRRGKV YDKYMCSFLF NLNNDFVVDA TRKGNKIRFA NHSVNPNCYA KVMMVNGDHR IGIFAKRA I QTGEELFFDY RYSQADALKY VGIEREMEIP UniProtKB: Histone-lysine N-methyltransferase EZH2 |
-Macromolecule #5: protein Jumonji isoform X3
| Macromolecule | Name: protein Jumonji isoform X3 / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 138.406219 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MSKERPKRNI IQKKYDDSDG IPWSEERVVR KVLYLSLKEF KNAQKRQHGE GIAGSLKSVN GLLGNDQSKA LGPASEQSEN EKDDASQVS STSNDVSSSD FEEGPSRKRP RLQAQRKFAQ SQPNSPSTTP VKTVEPLLPP PATQISDLSK RKPKTEDFLT F LCLRGSPA ...String: MSKERPKRNI IQKKYDDSDG IPWSEERVVR KVLYLSLKEF KNAQKRQHGE GIAGSLKSVN GLLGNDQSKA LGPASEQSEN EKDDASQVS STSNDVSSSD FEEGPSRKRP RLQAQRKFAQ SQPNSPSTTP VKTVEPLLPP PATQISDLSK RKPKTEDFLT F LCLRGSPA LPSSMVYFGS SQDEEDVEEE DDETEDVKTA NNNASSSCQS TPRKGKTHKH VHNGHVFNGS NRSTREKEPA QK HKSKETT PAKEKHIDHR ADSRREPASV AQPTATPSAG SLAKGLPANH QPPPPHRSAQ DLRKQVTLHV SKVNGVTRMS SLG AGTTSA KKIREVRPSP SKTVKYTATV TKGTVTYTKA KRELVKETKP THHKPSSAVN HTISGKTESS NAKTRKQVLS LGGA STSTG PAASGLKASS RLNPKSCTKE VGGRQLREGL RNSKRRLEEA QQVDKPQSPP KKMKGAAGIA EAPGKKASAA SAEKS LLNG HVKKEVPERS LERNRPKRAT AGKNMPGKQA HGKAEGTPCE NRSTSQPESS HKPHDPQGKP EKGIGKSGWT AMDEIP VLR PSAKEFHDPL IYIESVRAQV EKYGMCRVIP PPDWRPECKL NDEMRFVTQI QHIHKLGRRW GPNVQRLACI KKHLRSQ GI TMDELPLIGG CELDLACFFR LINEMGGMQQ VTDLKKWNKL ADMLRIPKTA QDRLAKLQEA YCQYLLSYDS LSPEEHRR L EKEVLMEKEI LEKRKGPLEG HTENDHHKFH SLPRFEPKNG LIHGVTPRNG FRSKLKEVGQ APLKTGRRRL FAQEKEVVK EEEEDKGVLN DFHKCIYKGR SVSLTTFYRT ARNIMNMCFS KEPAPAEIEQ EYWRLVEEKD CHVAVHCGKV DTNTHGSGFP VGKSEPFSR HGWNLTVLPN NTGSILRHLG AVPGVTIPWL NIGMVFSTSC WSRDQNHLPY IDYLHTGADC IWYCIPAEEE N KLEDVVHT LLQANGTPGL QMLESNVMIS PEVLCKEGIK VHRTVQQSGQ FVVCFPGSFV SKVCCGYSVS ETVHFATTQW TS MGFETAK EMKRRHIAKP FSMEKLLYQI AQAEAKKENG PTLSTISALL DELRDTELRQ RRQLFEAGLH SSARYGSHDG NST VADGKK KPRKWLQLET SERRCQICQH LCYLSMVVQE NENVVFCLEC ALRHVEKQKS CRGLKLMYRY DEEQIISLVN QICG KVSGK HGGIENCLNK PTPKRGPRKR ATVDVPPSRL PSS UniProtKB: Protein Jumonji isoform X3 |
-Macromolecule #6: Zinc finger protein AEBP2
| Macromolecule | Name: Zinc finger protein AEBP2 / type: protein_or_peptide / ID: 6 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 54.535496 KDa |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Sequence | String: MAAAITDMAD LEELSRLSPL PPGSPGSAAR GRAEPPEEEE EEEEEEEEAE AEAVAALLLN GGSGGGGGGG GGGVGGGEAE TMSEPSPES ASQAGEDEDE EEDDEEEEDE SSSSGGGEEE SSAESLVGSS GGSSSDETRS LSPGAASSSS GDGDGKEGLE E PKGPRGSQ ...String: MAAAITDMAD LEELSRLSPL PPGSPGSAAR GRAEPPEEEE EEEEEEEEAE AEAVAALLLN GGSGGGGGGG GGGVGGGEAE TMSEPSPES ASQAGEDEDE EEDDEEEEDE SSSSGGGEEE SSAESLVGSS GGSSSDETRS LSPGAASSSS GDGDGKEGLE E PKGPRGSQ GGGGGGSSSS SVVSSGGDEG YGTGGGGSSA TSGGRRGSLE MSSDGEPLSR MDSEDSISST IMDVDSTISS GR STPAMMN GQGSTTSSSK NIAYNCCWDQ CQACFNSSPD LADHIRSIHV DGQRGGVFVC LWKGCKVYNT PSTSQSWLQR HML THSGDK PFKCVVGGCN ASFASQGGLA RHVPTHFSQQ NSSKVSSQPK AKEESPSKAG MNKRRKLKNK RRRSLPRPHD FFDA QTLDA IRHRAICFNL SAHIESLGKG HSVVFHSTVI AKRKEDSGKI KLLLHWMPED ILPDVWVNES ERHQLKTKVV HLSKL PKDT ALLLDPNIYR TMPQKRLKRT LIRKVFNLYL SKQ UniProtKB: Zinc finger protein AEBP2 |
-Macromolecule #7: G4 RNA
| Macromolecule | Name: G4 RNA / type: rna / ID: 7 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 7.955823 KDa |
| Sequence | String: GGGUAAGGGU AAGGGUAAGG GUAA |
-Macromolecule #8: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 8 / Number of copies: 14 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 Component:
Details: RNP complex buffer (25 mM HEPES pH 7.9, 50 mM KCl, 2 mM MgCl2, 10% glycerol, and 1mM TCEP) EM preparation buffer I (25 mM HEPES pH 7.9, 50 mM KCl, 2.5% glycerol, and 1mM TCEP) EM preparation ...Details: RNP complex buffer (25 mM HEPES pH 7.9, 50 mM KCl, 2 mM MgCl2, 10% glycerol, and 1mM TCEP) EM preparation buffer I (25 mM HEPES pH 7.9, 50 mM KCl, 2.5% glycerol, and 1mM TCEP) EM preparation buffer II (25 mM HEPES pH 7.9, 50 mM KCl, 2.5% glycerol, 0.01%NP-40, and 1mM TCEP). | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R2/1 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 1.5 | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 281 K / Instrument: LEICA PLUNGER / Details: 3s of single side blotting. | ||||||||||||||||||
| Details | We used streptavidin-affinity grid preparation method with biotin-labeled RNA at 100 nM concentration. PRC2 was applied in excess at 600 nM. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 18632 / Average electron dose: 60.0 e/Å2 / Details: 60 frames per movie |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 0.01 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)