+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Tracing p2 phiPA3 PhuN Tetramer Interfaces | |||||||||
 Map data Map data | Core PhuN tetramer processed with larger mask and an interior mask to better resolve contacts made between tetramers. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | phiPA3 protein / shell protein / PhuN / phage nucleus / VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationdetection of maltose stimulus / maltose transport complex / carbohydrate transport / carbohydrate transmembrane transporter activity / maltose binding / maltose transport / maltodextrin transmembrane transport / ATP-binding cassette (ABC) transporter complex, substrate-binding subunit-containing / ATP-binding cassette (ABC) transporter complex / cell chemotaxis ...detection of maltose stimulus / maltose transport complex / carbohydrate transport / carbohydrate transmembrane transporter activity / maltose binding / maltose transport / maltodextrin transmembrane transport / ATP-binding cassette (ABC) transporter complex, substrate-binding subunit-containing / ATP-binding cassette (ABC) transporter complex / cell chemotaxis / outer membrane-bounded periplasmic space / host cell cytoplasm / periplasmic space / DNA damage response / membrane Similarity search - Function | |||||||||
| Biological species |  Pseudomonas phage PhiPA3 (virus) Pseudomonas phage PhiPA3 (virus) | |||||||||
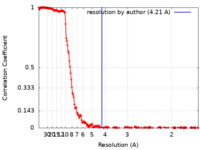
| Method | single particle reconstruction / cryo EM / Resolution: 4.21 Å | |||||||||
 Authors Authors | Nieweglowska ES / Brilot AF / Mendez-Moran M / Kokontis C / Baek M / Li J / Cheng Y / Baker D / Bondy-Denomy J / Agard DA | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: The ϕPA3 phage nucleus is enclosed by a self-assembling 2D crystalline lattice. Authors: Eliza S Nieweglowska / Axel F Brilot / Melissa Méndez-Moran / Claire Kokontis / Minkyung Baek / Junrui Li / Yifan Cheng / David Baker / Joseph Bondy-Denomy / David A Agard /  Abstract: To protect themselves from host attack, numerous jumbo bacteriophages establish a phage nucleus-a micron-scale, proteinaceous structure encompassing the replicating phage DNA. Bacteriophage and host ...To protect themselves from host attack, numerous jumbo bacteriophages establish a phage nucleus-a micron-scale, proteinaceous structure encompassing the replicating phage DNA. Bacteriophage and host proteins associated with replication and transcription are concentrated inside the phage nucleus while other phage and host proteins are excluded, including CRISPR-Cas and restriction endonuclease host defense systems. Here, we show that nucleus fragments isolated from ϕPA3 infected Pseudomonas aeruginosa form a 2-dimensional lattice, having p2 or p4 symmetry. We further demonstrate that recombinantly purified primary Phage Nuclear Enclosure (PhuN) protein spontaneously assembles into similar 2D sheets with p2 and p4 symmetry. We resolve the dominant p2 symmetric state to 3.9 Å by cryo-EM. Our structure reveals a two-domain core, organized into quasi-symmetric tetramers. Flexible loops and termini mediate adaptable inter-tetramer contacts that drive subunit assembly into a lattice and enable the adoption of different symmetric states. While the interfaces between subunits are mostly well packed, two are open, forming channels that likely have functional implications for the transport of proteins, mRNA, and small molecules. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29451.map.gz emd_29451.map.gz | 473.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29451-v30.xml emd-29451-v30.xml emd-29451.xml emd-29451.xml | 18 KB 18 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29451_fsc.xml emd_29451_fsc.xml | 23.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_29451.png emd_29451.png | 85.7 KB | ||
| Others |  emd_29451_half_map_1.map.gz emd_29451_half_map_1.map.gz emd_29451_half_map_2.map.gz emd_29451_half_map_2.map.gz | 162.4 MB 162.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29451 http://ftp.pdbj.org/pub/emdb/structures/EMD-29451 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29451 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29451 | HTTPS FTP |
-Related structure data
| Related structure data |  8fv5MC  8fneC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_29451.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29451.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Core PhuN tetramer processed with larger mask and an interior mask to better resolve contacts made between tetramers. | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.834 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_29451_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
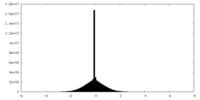
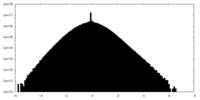
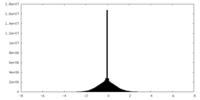
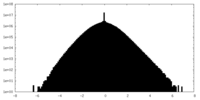
| Density Histograms |
-Half map: #1
| File | emd_29451_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Core tetramer assembly (p2) of the phiPA3 bacteriophage PhuN prot...
| Entire | Name: Core tetramer assembly (p2) of the phiPA3 bacteriophage PhuN protein with neighboring subunits |
|---|---|
| Components |
|
-Supramolecule #1: Core tetramer assembly (p2) of the phiPA3 bacteriophage PhuN prot...
| Supramolecule | Name: Core tetramer assembly (p2) of the phiPA3 bacteriophage PhuN protein with neighboring subunits type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Pseudomonas phage PhiPA3 (virus) Pseudomonas phage PhiPA3 (virus) |
-Macromolecule #1: Maltose/maltodextrin-binding periplasmic protein, phiPA3 PhuN
| Macromolecule | Name: Maltose/maltodextrin-binding periplasmic protein, phiPA3 PhuN type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pseudomonas phage PhiPA3 (virus) Pseudomonas phage PhiPA3 (virus) |
| Recombinant expression | Organism:  |
| Sequence | String: HHHHHHMKIE EGKLVIWING DKGYNGLAEV GKKFEKDTGI KVTVEHPDKL EEKFPQVAAT GDGPDIIFWA HDRFGGYAQS GLLAEITPDK AFQDKLYPFT WDAVRYNGKL IAYPIAVEAL SLIYNKDLLP NPPKTWEEIP ALDKELKAKG KSALMFNLQE PYFTWPLIAA ...String: HHHHHHMKIE EGKLVIWING DKGYNGLAEV GKKFEKDTGI KVTVEHPDKL EEKFPQVAAT GDGPDIIFWA HDRFGGYAQS GLLAEITPDK AFQDKLYPFT WDAVRYNGKL IAYPIAVEAL SLIYNKDLLP NPPKTWEEIP ALDKELKAKG KSALMFNLQE PYFTWPLIAA DGGYAFKYEN GKYDIKDVGV DNAGAKAGLT FLVDLIKNKH MNADTDYSIA EAAFNKGETA MTINGPWAWS NIDTSKVNYG VTVLPTFKGQ PSKPFVGVLS AGINAASPNK ELAKEFLENY LLTDEGLEAV NKDKPLGAVA LKSYEEELAK DPRIAATMEN AQKGEIMPNI PQMSAFWYAV RTAVINAASG RQTVDEALKD AQTGKPIPNP LLGLDSTENL YFQGMQQTQQ GPKVQTQTLQ GGAGNLNSIF QRSGRTDGGD ARASEALAVF NKLKEEAIAQ QDLHDDFLVF RFDRDQNRVG YSALLVVKRA AINGQQVIVT RPLVMPNDQI TLPTKKLTIQ NGMHQETIEA EADVQDVFTT QYWNRICDSI RQQTGKHDAM VINAGPTVIP ADFDLKDELV LKQLLIKSVN LCDDMLAKRS GEQPFSVAML KGTDETLAAR LNFTGKPMHD SLGYPIRSDI LVSLNRVKKP GQQENEFYEA EDKLNQVSCF VNLEYTPQPQ QAIYGAPQQT QQLPPLTPAI VITDVRQAEW LKANTMELYL FALSNAFRVT ANQSWARSLL PQLGKVKDMR DIGAIGYLSR LAARVETKTE TFTDQNFAEL LYNMVRPSPV FMSDLNRFGD NAAIENVFID ALGGVNQQRA VAAIIAGVNN LIGGGFEKFF DHNTMPIIQP YGTDIQLGYY LDGEGEKQDR RDLDVLGALN ASDGNIQEWM SWYGTQCNVA VHPELRARQS KNFDRQYLGN SVTYTTRAHR GIWNPKFIEA LDKAIASVGL TVAMDNVAQV FGAQRFSGNL AIADYAVTGT AQVSSGLVSN GGYNPQFGVG QGSGFY |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | 2D array |
- Sample preparation
Sample preparation
| Buffer | pH: 6.5 Component:
Details: 0.25 Complete Protease Inhibitor Tablet also included | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 67.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-8fv5: |
 Movie
Movie Controller
Controller







 Z
Z Y
Y X
X