[English] 日本語
 Yorodumi
Yorodumi- EMDB-28781: Structure of a 12mer DegP cage bound to the client protein hTRF1 -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of a 12mer DegP cage bound to the client protein hTRF1 | |||||||||
 Map data Map data | Cryo-EM map of a 12mer DegP cage with constituent trimers bound to the client hTRF1. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Protease / chaperone / hydrolase / cage / complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of shelterin complex assembly / negative regulation of establishment of protein localization to telomere / negative regulation of establishment of RNA localization to telomere / negative regulation of establishment of protein-containing complex localization to telomere / negative regulation of telomere maintenance via semi-conservative replication / negative regulation of exonuclease activity / negative regulation of telomeric D-loop disassembly / meiotic telomere clustering / peptidase Do / t-circle formation ...positive regulation of shelterin complex assembly / negative regulation of establishment of protein localization to telomere / negative regulation of establishment of RNA localization to telomere / negative regulation of establishment of protein-containing complex localization to telomere / negative regulation of telomere maintenance via semi-conservative replication / negative regulation of exonuclease activity / negative regulation of telomeric D-loop disassembly / meiotic telomere clustering / peptidase Do / t-circle formation / telomeric D-loop disassembly / shelterin complex / Telomere C-strand synthesis initiation / double-stranded telomeric DNA binding / Telomere C-strand (Lagging Strand) Synthesis / : / positive regulation of telomere maintenance / nuclear telomere cap complex / ankyrin repeat binding / Processive synthesis on the C-strand of the telomere / Polymerase switching on the C-strand of the telomere / Removal of the Flap Intermediate from the C-strand / G-rich strand telomeric DNA binding / telomere capping / DNA binding, bending / negative regulation of telomere maintenance via telomere lengthening / response to temperature stimulus / protein quality control for misfolded or incompletely synthesized proteins / telomeric DNA binding / negative regulation of DNA replication / negative regulation of telomere maintenance via telomerase / telomere maintenance via telomerase / Telomere Extension By Telomerase / Packaging Of Telomere Ends / chaperone-mediated protein folding / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / Inhibition of DNA recombination at telomere / Meiotic synapsis / serine-type peptidase activity / telomere maintenance / DNA Damage/Telomere Stress Induced Senescence / fibrillar center / spindle / protein folding / peptidase activity / outer membrane-bounded periplasmic space / response to heat / microtubule binding / response to oxidative stress / chromosome, telomeric region / periplasmic space / nuclear body / cell division / serine-type endopeptidase activity / nucleolus / protein homodimerization activity / proteolysis / DNA binding / nucleoplasm / identical protein binding / nucleus / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
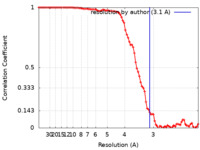
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Harkness RW / Ripstein ZA / Di Trani JM / Kay LE | |||||||||
| Funding support |  Canada, 1 items Canada, 1 items
| |||||||||
 Citation Citation |  Journal: J Am Chem Soc / Year: 2023 Journal: J Am Chem Soc / Year: 2023Title: Flexible Client-Dependent Cages in the Assembly Landscape of the Periplasmic Protease-Chaperone DegP. Authors: Robert W Harkness / Zev A Ripstein / Justin M Di Trani / Lewis E Kay /  Abstract: The periplasmic protein DegP, which is implicated in virulence factor transport leading to pathogenicity, is a bi-functional protease and chaperone that helps to maintain protein homeostasis in Gram- ...The periplasmic protein DegP, which is implicated in virulence factor transport leading to pathogenicity, is a bi-functional protease and chaperone that helps to maintain protein homeostasis in Gram-negative bacteria and is essential to bacterial survival under stress conditions. To perform these functions, DegP captures clients inside cage-like structures, which we have recently shown to form through the reorganization of high-order preformed apo oligomers, consisting of trimeric building blocks, that are structurally distinct from client-bound cages. Our previous studies suggested that these apo oligomers may allow DegP to encapsulate clients of various sizes under protein folding stresses by forming ensembles that can include extremely large cage particles, but how this occurs remains an open question. To explore the relation between cage and substrate sizes, we engineered a series of DegP clients of increasing hydrodynamic radii and analyzed their influence on DegP cage formation. We used dynamic light scattering and cryogenic electron microscopy to characterize the hydrodynamic properties and structures of the DegP cages that are adopted in response to each client. We present a series of density maps and structural models that include those for novel particles of approximately 30 and 60 monomers. Key interactions between DegP trimers and the bound clients that stabilize the cage assemblies and prime the clients for catalysis are revealed. We also provide evidence that DegP can form cages which approach subcellular organelles in terms of size. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28781.map.gz emd_28781.map.gz | 59.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28781-v30.xml emd-28781-v30.xml emd-28781.xml emd-28781.xml | 16.5 KB 16.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28781_fsc.xml emd_28781_fsc.xml | 8.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_28781.png emd_28781.png | 63.4 KB | ||
| Filedesc metadata |  emd-28781.cif.gz emd-28781.cif.gz | 5.5 KB | ||
| Others |  emd_28781_half_map_1.map.gz emd_28781_half_map_1.map.gz emd_28781_half_map_2.map.gz emd_28781_half_map_2.map.gz | 59.2 MB 59.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28781 http://ftp.pdbj.org/pub/emdb/structures/EMD-28781 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28781 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28781 | HTTPS FTP |
-Validation report
| Summary document |  emd_28781_validation.pdf.gz emd_28781_validation.pdf.gz | 815.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_28781_full_validation.pdf.gz emd_28781_full_validation.pdf.gz | 815.4 KB | Display | |
| Data in XML |  emd_28781_validation.xml.gz emd_28781_validation.xml.gz | 16.5 KB | Display | |
| Data in CIF |  emd_28781_validation.cif.gz emd_28781_validation.cif.gz | 21.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28781 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28781 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28781 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28781 | HTTPS FTP |
-Related structure data
| Related structure data |  8f0uMC  8f0aC  8f1tC  8f1uC  8f21C  8f26C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_28781.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28781.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM map of a 12mer DegP cage with constituent trimers bound to the client hTRF1. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_28781_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_28781_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Structure of a 12mer DegP cage bound to the client protein hTRF1
| Entire | Name: Structure of a 12mer DegP cage bound to the client protein hTRF1 |
|---|---|
| Components |
|
-Supramolecule #1: Structure of a 12mer DegP cage bound to the client protein hTRF1
| Supramolecule | Name: Structure of a 12mer DegP cage bound to the client protein hTRF1 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Periplasmic serine endoprotease DegP
| Macromolecule | Name: Periplasmic serine endoprotease DegP / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: peptidase Do |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 36.376176 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPSLAPMLEK VMPSVVSINV EGSTTVNTPR MPRNFQQFFG DDSPFCQEGS PFQSSPFCQG GQGGNGGGQQ QKFMALGSGV IIDADKGYV VTNNHVVDNA TVIKVQLSDG RKFDAKMVGK DPRSDIALIQ IQNPKNLTAI KMADSDALRV GDYTVAIGNP F GLGETVTS ...String: MPSLAPMLEK VMPSVVSINV EGSTTVNTPR MPRNFQQFFG DDSPFCQEGS PFQSSPFCQG GQGGNGGGQQ QKFMALGSGV IIDADKGYV VTNNHVVDNA TVIKVQLSDG RKFDAKMVGK DPRSDIALIQ IQNPKNLTAI KMADSDALRV GDYTVAIGNP F GLGETVTS GIVSALGRSG LNAENYENFI QTDAAINRGN AGGALVNLNG ELIGINTAIL APDGGNIGIG FAIPSNMVKN LT SQMVEYG QVKRGELGIM GTELNSELAK AMKVDAQRGA FVSQVLPNSS AAKAGIKAGD VITSLNGKPI SSFAALRAQV GTM PVGSKL TLGLLRDGKQ VNVNLELQQS SQ UniProtKB: Periplasmic serine endoprotease DegP |
-Macromolecule #2: Periplasmic serine endoprotease DegP
| Macromolecule | Name: Periplasmic serine endoprotease DegP / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: peptidase Do |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 7.970229 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AEMSNKGKDQ GVVVNNVKTG TPAAQIGLKK GDVIIGANQQ AVKNIAELRK VLDSKPSVLA LNIQRGDSTI YLLMQ UniProtKB: Periplasmic serine endoprotease DegP |
-Macromolecule #3: Telomeric repeat-binding factor 1
| Macromolecule | Name: Telomeric repeat-binding factor 1 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 3.417168 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SKILLHYKFN NRTSVMLKDR WRTMKKL UniProtKB: Telomeric repeat-binding factor 1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.9 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)