+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Capsid structure of Staphylococcus phage Andhra | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | phage capsid / HK97-like fold / VIRUS | |||||||||
| Function / homology | Major capsid protein / DUF4025 domain-containing protein Function and homology information Function and homology information | |||||||||
| Biological species |  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Hawkins NC / Kizziah JL / Dokland T | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2022 Journal: Sci Adv / Year: 2022Title: Structure and host specificity of bacteriophage Andhra. Authors: N'Toia C Hawkins / James L Kizziah / Asma Hatoum-Aslan / Terje Dokland /  Abstract: is an opportunistic pathogen of the human skin, often associated with infections of implanted medical devices. Staphylococcal picoviruses are a group of strictly lytic, short-tailed bacteriophages ... is an opportunistic pathogen of the human skin, often associated with infections of implanted medical devices. Staphylococcal picoviruses are a group of strictly lytic, short-tailed bacteriophages with compact genomes that are attractive candidates for therapeutic use. Here, we report the structure of the complete virion of -infecting phage Andhra, determined using high-resolution cryo-electron microscopy, allowing atomic modeling of 11 capsid and tail proteins. The capsid is a = 4 icosahedron containing a unique stabilizing capsid lining protein. The tail includes 12 trimers of a unique receptor binding protein (RBP), a lytic protein that also serves to anchor the RBPs to the tail stem, and a hexameric tail knob that acts as a gatekeeper for DNA ejection. Using structure prediction with AlphaFold, we identified the two proteins that comprise the tail tip heterooctamer. Our findings elucidate critical features for virion assembly, host recognition, and penetration. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28130.map.gz emd_28130.map.gz | 106.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28130-v30.xml emd-28130-v30.xml emd-28130.xml emd-28130.xml | 15 KB 15 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28130_fsc.xml emd_28130_fsc.xml | 18 KB | Display |  FSC data file FSC data file |
| Images |  emd_28130.png emd_28130.png | 219.9 KB | ||
| Filedesc metadata |  emd-28130.cif.gz emd-28130.cif.gz | 5.3 KB | ||
| Others |  emd_28130_half_map_1.map.gz emd_28130_half_map_1.map.gz emd_28130_half_map_2.map.gz emd_28130_half_map_2.map.gz | 405.3 MB 405.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28130 http://ftp.pdbj.org/pub/emdb/structures/EMD-28130 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28130 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28130 | HTTPS FTP |
-Validation report
| Summary document |  emd_28130_validation.pdf.gz emd_28130_validation.pdf.gz | 845.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_28130_full_validation.pdf.gz emd_28130_full_validation.pdf.gz | 845.1 KB | Display | |
| Data in XML |  emd_28130_validation.xml.gz emd_28130_validation.xml.gz | 25.2 KB | Display | |
| Data in CIF |  emd_28130_validation.cif.gz emd_28130_validation.cif.gz | 34.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28130 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28130 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28130 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-28130 | HTTPS FTP |
-Related structure data
| Related structure data |  8egtMC  8egrC  8egsC  8ej5C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_28130.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28130.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.328 Å | ||||||||||||||||||||||||||||||||||||
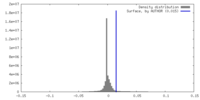
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_28130_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
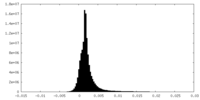
| Density Histograms |
-Half map: #2
| File | emd_28130_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
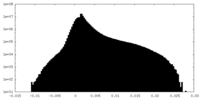
| Density Histograms |
- Sample components
Sample components
-Entire : Staphylococcus phage Andhra
| Entire | Name:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Staphylococcus phage Andhra
| Supramolecule | Name: Staphylococcus phage Andhra / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 1958907 / Sci species name: Staphylococcus phage Andhra / Virus type: VIRION / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  |
-Macromolecule #1: gp19, capsid lining protein
| Macromolecule | Name: gp19, capsid lining protein / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Molecular weight | Theoretical: 7.141647 KDa |
| Recombinant expression | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Sequence | String: MANFDGNEMR GMTHANYEDS RLNKSRELNA NMSIGTSKSE DEYGRQVHSL TKQSYSDDSV QEA UniProtKB: DUF4025 domain-containing protein |
-Macromolecule #2: Major capsid protein
| Macromolecule | Name: Major capsid protein / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Molecular weight | Theoretical: 46.35309 KDa |
| Recombinant expression | Organism:  Staphylococcus phage Andhra (virus) Staphylococcus phage Andhra (virus) |
| Sequence | String: MADKKTDIPT LIADSTKASL QDFNHDYGKQ WTFGENWSNV NTMFETYVNK YLFPKINETL LIDIALGNRF NWLAKEQDFI GQYSEEYVI MDTIPIEMNL SKSEELMLKR NYPQMATRLY GSGIVKKQKF TLNNNDVRFN FQTLGDATNY ALGVLRKKIS D INVQEEKE ...String: MADKKTDIPT LIADSTKASL QDFNHDYGKQ WTFGENWSNV NTMFETYVNK YLFPKINETL LIDIALGNRF NWLAKEQDFI GQYSEEYVI MDTIPIEMNL SKSEELMLKR NYPQMATRLY GSGIVKKQKF TLNNNDVRFN FQTLGDATNY ALGVLRKKIS D INVQEEKE IRAMMVDYAI NQLQDSNRRT ASSKEDLTER VFEAILNMQN NSAKYNEVHK ASGGSVGQYT TVSKLSDIAI LT TDSLKSY LLDTKIANTF QMAGIDFTDH IISFDDLGGV YKTTKDVTLA NEDTINYLRA FGDYQAMIGD VIPTGSVFTF NVS DLKEFK GNIEEIKPQG ELFAFIFDIN ALKYKRNTKG MLKEPFYNGE FDEVTHWIHY YSFKAMSPFF NKILITEAPK EQPD AGATE UniProtKB: Major capsid protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 39.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)