[English] 日本語
 Yorodumi
Yorodumi- EMDB-27258: Human DNA polymerase-alpha/primase elongation complex II bound to... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human DNA polymerase-alpha/primase elongation complex II bound to primer/template | |||||||||
 Map data Map data | Unsharpened and flipped map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | DNA replication / human DNA polymerase alpha/primase / human primosome / elongation complex / REPLICATION | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of DNA primase activity / DNA primase AEP / ribonucleotide binding / DNA replication initiation / DNA/RNA hybrid binding / Telomere C-strand synthesis initiation / Inhibition of replication initiation of damaged DNA by RB1/E2F1 / alpha DNA polymerase:primase complex / Polymerase switching / regulation of type I interferon production ...positive regulation of DNA primase activity / DNA primase AEP / ribonucleotide binding / DNA replication initiation / DNA/RNA hybrid binding / Telomere C-strand synthesis initiation / Inhibition of replication initiation of damaged DNA by RB1/E2F1 / alpha DNA polymerase:primase complex / Polymerase switching / regulation of type I interferon production / Processive synthesis on the lagging strand / DNA primase activity / Removal of the Flap Intermediate / Polymerase switching on the C-strand of the telomere / lagging strand elongation / DNA replication, synthesis of primer / mitotic DNA replication initiation / DNA strand elongation involved in DNA replication / DNA synthesis involved in DNA repair / G1/S-Specific Transcription / leading strand elongation / DNA replication origin binding / Activation of the pre-replicative complex / DNA replication initiation / Defective pyroptosis / nuclear matrix / double-strand break repair via nonhomologous end joining / protein import into nucleus / nuclear envelope / single-stranded DNA binding / 4 iron, 4 sulfur cluster binding / DNA replication / DNA-directed DNA polymerase / DNA-directed DNA polymerase activity / DNA repair / nucleotide binding / chromatin binding / chromatin / nucleolus / protein kinase binding / magnesium ion binding / DNA binding / zinc ion binding / nucleoplasm / membrane / nucleus / metal ion binding / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
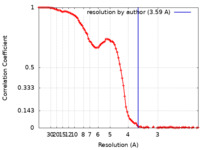
| Method | single particle reconstruction / cryo EM / Resolution: 3.59 Å | |||||||||
 Authors Authors | He Q / Baranovskiy A / Lim C / Tahirov T | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2023 Journal: Nat Struct Mol Biol / Year: 2023Title: Structures of human primosome elongation complexes. Authors: Qixiang He / Andrey G Baranovskiy / Lucia M Morstadt / Alisa E Lisova / Nigar D Babayeva / Benjamin L Lusk / Ci Ji Lim / Tahir H Tahirov /  Abstract: The synthesis of RNA-DNA primer by primosome requires coordination between primase and DNA polymerase α subunits, which is accompanied by unknown architectural rearrangements of multiple domains. ...The synthesis of RNA-DNA primer by primosome requires coordination between primase and DNA polymerase α subunits, which is accompanied by unknown architectural rearrangements of multiple domains. Using cryogenic electron microscopy, we solved a 3.6 Å human primosome structure caught at an early stage of RNA primer elongation with deoxynucleotides. The structure confirms a long-standing role of primase large subunit and reveals new insights into how primosome is limited to synthesizing short RNA-DNA primers. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27258.map.gz emd_27258.map.gz | 32.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27258-v30.xml emd-27258-v30.xml emd-27258.xml emd-27258.xml | 27.5 KB 27.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27258_fsc.xml emd_27258_fsc.xml | 11.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_27258.png emd_27258.png | 120.2 KB | ||
| Filedesc metadata |  emd-27258.cif.gz emd-27258.cif.gz | 8.6 KB | ||
| Others |  emd_27258_half_map_1.map.gz emd_27258_half_map_1.map.gz emd_27258_half_map_2.map.gz emd_27258_half_map_2.map.gz | 59.4 MB 59.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27258 http://ftp.pdbj.org/pub/emdb/structures/EMD-27258 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27258 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27258 | HTTPS FTP |
-Validation report
| Summary document |  emd_27258_validation.pdf.gz emd_27258_validation.pdf.gz | 816.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_27258_full_validation.pdf.gz emd_27258_full_validation.pdf.gz | 815.7 KB | Display | |
| Data in XML |  emd_27258_validation.xml.gz emd_27258_validation.xml.gz | 15.7 KB | Display | |
| Data in CIF |  emd_27258_validation.cif.gz emd_27258_validation.cif.gz | 20.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27258 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27258 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27258 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-27258 | HTTPS FTP |
-Related structure data
| Related structure data |  8d9dMC  8d96C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_27258.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27258.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened and flipped map | ||||||||||||||||||||
| Voxel size | X=Y=Z: 1.12 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map B
| File | emd_27258_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map A
| File | emd_27258_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Elongation complex II of human DNA polymerase alpha/primase bound...
+Supramolecule #1: Elongation complex II of human DNA polymerase alpha/primase bound...
+Macromolecule #1: DNA primase small subunit
+Macromolecule #2: DNA primase large subunit
+Macromolecule #3: DNA polymerase alpha catalytic subunit
+Macromolecule #4: DNA polymerase alpha subunit B
+Macromolecule #5: DNA/RNA (5'-D(*(GTP))-R(P*GP*CP*GP*GP*CP*AP*CP*G)-D(P*AP*CP*C)-3')
+Macromolecule #6: DNA (5'-D(*AP*TP*GP*GP*TP*CP*GP*TP*GP*CP*CP*GP*CP*CP*AP*AP*TP*AP*...
+Macromolecule #7: ZINC ION
+Macromolecule #8: IRON/SULFUR CLUSTER
+Macromolecule #9: MAGNESIUM ION
+Macromolecule #10: 2'-DEOXYADENOSINE 5'-TRIPHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 6 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: CHAPSO is made fresh at 80 mM before added to the sample at a final concentration of 4-8 mM immediately before vitrification. | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV | ||||||||||||||||||
| Details | This sample was monodisperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 2 / Number real images: 13243 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Protocol: RIGID BODY FIT | ||||||||||
| Output model |  PDB-8d9d: |
 Movie
Movie Controller
Controller













 Z
Z Y
Y X
X