[English] 日本語
 Yorodumi
Yorodumi- EMDB-27091: CryoEM structure of amplified alpha-synuclein fibril class A type... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of amplified alpha-synuclein fibril class A type I with extended core from DLB case VII | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  alpha-synuclein / Cerebrospinal Fluid (CSF) / PROTEIN FIBRIL alpha-synuclein / Cerebrospinal Fluid (CSF) / PROTEIN FIBRIL | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of mitochondrial electron transport, NADH to ubiquinone / neutral lipid metabolic process /  regulation of phospholipase activity / negative regulation of monooxygenase activity / regulation of acyl-CoA biosynthetic process / negative regulation of dopamine uptake involved in synaptic transmission / negative regulation of norepinephrine uptake / positive regulation of glutathione peroxidase activity / positive regulation of SNARE complex assembly / positive regulation of hydrogen peroxide catabolic process ...negative regulation of mitochondrial electron transport, NADH to ubiquinone / neutral lipid metabolic process / regulation of phospholipase activity / negative regulation of monooxygenase activity / regulation of acyl-CoA biosynthetic process / negative regulation of dopamine uptake involved in synaptic transmission / negative regulation of norepinephrine uptake / positive regulation of glutathione peroxidase activity / positive regulation of SNARE complex assembly / positive regulation of hydrogen peroxide catabolic process ...negative regulation of mitochondrial electron transport, NADH to ubiquinone / neutral lipid metabolic process /  regulation of phospholipase activity / negative regulation of monooxygenase activity / regulation of acyl-CoA biosynthetic process / negative regulation of dopamine uptake involved in synaptic transmission / negative regulation of norepinephrine uptake / positive regulation of glutathione peroxidase activity / positive regulation of SNARE complex assembly / positive regulation of hydrogen peroxide catabolic process / supramolecular fiber / negative regulation of transporter activity / mitochondrial membrane organization / negative regulation of chaperone-mediated autophagy / regulation of reactive oxygen species biosynthetic process / positive regulation of protein localization to cell periphery / regulation of phospholipase activity / negative regulation of monooxygenase activity / regulation of acyl-CoA biosynthetic process / negative regulation of dopamine uptake involved in synaptic transmission / negative regulation of norepinephrine uptake / positive regulation of glutathione peroxidase activity / positive regulation of SNARE complex assembly / positive regulation of hydrogen peroxide catabolic process / supramolecular fiber / negative regulation of transporter activity / mitochondrial membrane organization / negative regulation of chaperone-mediated autophagy / regulation of reactive oxygen species biosynthetic process / positive regulation of protein localization to cell periphery /  regulation of synaptic vesicle recycling / negative regulation of platelet-derived growth factor receptor signaling pathway / negative regulation of exocytosis / regulation of glutamate secretion / regulation of norepinephrine uptake / response to iron(II) ion / dopamine biosynthetic process / regulation of synaptic vesicle recycling / negative regulation of platelet-derived growth factor receptor signaling pathway / negative regulation of exocytosis / regulation of glutamate secretion / regulation of norepinephrine uptake / response to iron(II) ion / dopamine biosynthetic process /  SNARE complex assembly / positive regulation of neurotransmitter secretion / regulation of locomotion / positive regulation of inositol phosphate biosynthetic process / synaptic vesicle priming / SNARE complex assembly / positive regulation of neurotransmitter secretion / regulation of locomotion / positive regulation of inositol phosphate biosynthetic process / synaptic vesicle priming /  regulation of macrophage activation / dopamine uptake involved in synaptic transmission / negative regulation of microtubule polymerization / synaptic vesicle transport / dynein complex binding / positive regulation of receptor recycling / regulation of dopamine secretion / regulation of macrophage activation / dopamine uptake involved in synaptic transmission / negative regulation of microtubule polymerization / synaptic vesicle transport / dynein complex binding / positive regulation of receptor recycling / regulation of dopamine secretion /  protein kinase inhibitor activity / negative regulation of thrombin-activated receptor signaling pathway / response to type II interferon / cuprous ion binding / positive regulation of exocytosis / synaptic vesicle exocytosis / protein kinase inhibitor activity / negative regulation of thrombin-activated receptor signaling pathway / response to type II interferon / cuprous ion binding / positive regulation of exocytosis / synaptic vesicle exocytosis /  kinesin binding / positive regulation of endocytosis / mitochondrial ATP synthesis coupled electron transport / cysteine-type endopeptidase inhibitor activity involved in apoptotic process / response to magnesium ion / synaptic vesicle endocytosis / regulation of presynapse assembly / alpha-tubulin binding / negative regulation of serotonin uptake / localization / phospholipid metabolic process / supramolecular fiber organization / axon terminus / kinesin binding / positive regulation of endocytosis / mitochondrial ATP synthesis coupled electron transport / cysteine-type endopeptidase inhibitor activity involved in apoptotic process / response to magnesium ion / synaptic vesicle endocytosis / regulation of presynapse assembly / alpha-tubulin binding / negative regulation of serotonin uptake / localization / phospholipid metabolic process / supramolecular fiber organization / axon terminus /  inclusion body / cellular response to copper ion / inclusion body / cellular response to copper ion /  Hsp70 protein binding / cellular response to epinephrine stimulus / Hsp70 protein binding / cellular response to epinephrine stimulus /  excitatory postsynaptic potential / response to interleukin-1 / adult locomotory behavior / excitatory postsynaptic potential / response to interleukin-1 / adult locomotory behavior /  SNARE binding / positive regulation of release of sequestered calcium ion into cytosol / fatty acid metabolic process / long-term synaptic potentiation / regulation of transmembrane transporter activity / SNARE binding / positive regulation of release of sequestered calcium ion into cytosol / fatty acid metabolic process / long-term synaptic potentiation / regulation of transmembrane transporter activity /  phosphoprotein binding / protein tetramerization / synapse organization / microglial cell activation / regulation of long-term neuronal synaptic plasticity / negative regulation of protein kinase activity / phosphoprotein binding / protein tetramerization / synapse organization / microglial cell activation / regulation of long-term neuronal synaptic plasticity / negative regulation of protein kinase activity /  ferrous iron binding / protein destabilization / PKR-mediated signaling / negative regulation of cysteine-type endopeptidase activity involved in apoptotic process / tau protein binding / positive regulation of protein serine/threonine kinase activity / ferrous iron binding / protein destabilization / PKR-mediated signaling / negative regulation of cysteine-type endopeptidase activity involved in apoptotic process / tau protein binding / positive regulation of protein serine/threonine kinase activity /  receptor internalization / receptor internalization /  phospholipid binding / synaptic vesicle membrane / positive regulation of inflammatory response / activation of cysteine-type endopeptidase activity involved in apoptotic process / phospholipid binding / synaptic vesicle membrane / positive regulation of inflammatory response / activation of cysteine-type endopeptidase activity involved in apoptotic process /  actin cytoskeleton / positive regulation of peptidyl-serine phosphorylation / actin cytoskeleton / positive regulation of peptidyl-serine phosphorylation /  actin binding / actin binding /  cell cortex / cellular response to oxidative stress / cell cortex / cellular response to oxidative stress /  histone binding / histone binding /  growth cone / postsynapse / chemical synaptic transmission / neuron apoptotic process / negative regulation of neuron apoptotic process / amyloid fibril formation / response to lipopolysaccharide / growth cone / postsynapse / chemical synaptic transmission / neuron apoptotic process / negative regulation of neuron apoptotic process / amyloid fibril formation / response to lipopolysaccharide /  lysosome / molecular adaptor activity / lysosome / molecular adaptor activity /  oxidoreductase activity / transcription cis-regulatory region binding oxidoreductase activity / transcription cis-regulatory region bindingSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
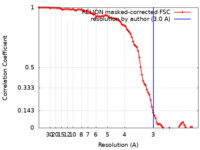
| Method | helical reconstruction /  cryo EM / Resolution: 3.0 Å cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Zhou Y / Sokratian A / Xu E / Viverette E / Dillard L / Yuan Y / Li JY / Matarangas A / Bouvette J / Borgnia M ...Zhou Y / Sokratian A / Xu E / Viverette E / Dillard L / Yuan Y / Li JY / Matarangas A / Bouvette J / Borgnia M / Bartesaghi A / West A | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structural and functional landscape of alpha-synuclein fibril assemblies amplified from cerebrospinal fluid Authors: Zhou Y / Sokratian A / Xu E / Viverette E / Dillard L / Yuan Y / Li JY / Matarangas A / Bouvette J / Borgnia M / Bartesaghi A / West A | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_27091.map.gz emd_27091.map.gz | 1.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-27091-v30.xml emd-27091-v30.xml emd-27091.xml emd-27091.xml | 17.7 KB 17.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_27091_fsc.xml emd_27091_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_27091.png emd_27091.png | 97.4 KB | ||
| Masks |  emd_27091_msk_1.map emd_27091_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-27091.cif.gz emd-27091.cif.gz | 5.6 KB | ||
| Others |  emd_27091_half_map_1.map.gz emd_27091_half_map_1.map.gz emd_27091_half_map_2.map.gz emd_27091_half_map_2.map.gz | 49.6 MB 49.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-27091 http://ftp.pdbj.org/pub/emdb/structures/EMD-27091 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27091 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-27091 | HTTPS FTP |
-Related structure data
| Related structure data |  8cz2MC  8cysC  8cytC  8cyvC  8cywC  8cyxC  8cyyC  8cz0C  8cz1C  8cz3C  8cz6C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_27091.map.gz / Format: CCP4 / Size: 6.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_27091.map.gz / Format: CCP4 / Size: 6.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_27091_msk_1.map emd_27091_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
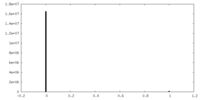
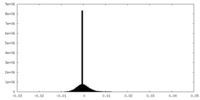
| Density Histograms |
-Half map: #2
| File | emd_27091_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
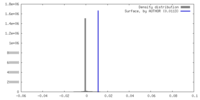
| Density Histograms |
-Half map: #1
| File | emd_27091_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
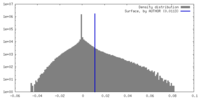
| Density Histograms |
- Sample components
Sample components
-Entire : Amplified alpha-synuclein fibrils from Lewy body dementia CSF
| Entire | Name: Amplified alpha-synuclein fibrils from Lewy body dementia CSF |
|---|---|
| Components |
|
-Supramolecule #1: Amplified alpha-synuclein fibrils from Lewy body dementia CSF
| Supramolecule | Name: Amplified alpha-synuclein fibrils from Lewy body dementia CSF type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all Details: alpha-synuclein fibril produced in presence of postmortem cerebrospinal fluid from specimens affected by synucleinopathies |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) / Tissue: cerebrospinal fluid Homo sapiens (human) / Tissue: cerebrospinal fluid |
| Molecular weight | Theoretical: 60 kDa/nm |
-Macromolecule #1: Alpha-synuclein
| Macromolecule | Name: Alpha-synuclein / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 14.476108 KDa |
| Recombinant expression | Organism:   Escherichia coli 'BL21-Gold(DE3)pLysS AG' (bacteria) Escherichia coli 'BL21-Gold(DE3)pLysS AG' (bacteria) |
| Sequence | String: MDVFMKGLSK AKEGVVAAAE KTKQGVAEAA GKTKEGVLYV GSKTKEGVVH GVATVAEKTK EQVTNVGGAV VTGVTAVAQK TVEGAGSIA AATGFVKKDQ LGKNEEGAPQ EGILEDMPVD PDNEAYEMPS EEGYQDYEPE A UniProtKB:  Alpha-synuclein Alpha-synuclein |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: Gibco Cat # 10010072 | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-8cz2: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)