[English] 日本語
 Yorodumi
Yorodumi- EMDB-26492: Cryo-EM structure of BG24 inferred germline Fabs with mature CDR3... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
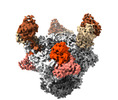
| Title | Cryo-EM structure of BG24 inferred germline Fabs with mature CDR3s and 10-1074 Fabs in complex with HIV-1 Env immunogen BG505-SOSIPv4.1-GT1 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / positive regulation of establishment of T cell polarity / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / virus-mediated perturbation of host defense response / fusion of virus membrane with host endosome membrane / viral envelope ...positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / positive regulation of establishment of T cell polarity / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / viral protein processing / fusion of virus membrane with host plasma membrane / virus-mediated perturbation of host defense response / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / apoptotic process / host cell plasma membrane / structural molecule activity / virion membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | Dam KA / Bjorkman PJ | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: HIV-1 CD4-binding site germline antibody-Env structures inform vaccine design. Authors: Kim-Marie A Dam / Christopher O Barnes / Harry B Gristick / Till Schoofs / Priyanthi N P Gnanapragasam / Michel C Nussenzweig / Pamela J Bjorkman /    Abstract: BG24, a VRC01-class broadly neutralizing antibody (bNAb) against HIV-1 Env with relatively few somatic hypermutations (SHMs), represents a promising target for vaccine strategies to elicit CD4- ...BG24, a VRC01-class broadly neutralizing antibody (bNAb) against HIV-1 Env with relatively few somatic hypermutations (SHMs), represents a promising target for vaccine strategies to elicit CD4-binding site (CD4bs) bNAbs. To understand how SHMs correlate with BG24 neutralization of HIV-1, we report 4.1 Å and 3.4 Å single-particle cryo-EM structures of two inferred germline (iGL) BG24 precursors complexed with engineered Env-based immunogens lacking CD4bs N-glycans. Structures reveal critical Env contacts by BG24 and identify antibody light chain structural features that impede Env recognition. In addition, biochemical data and cryo-EM structures of BG24 variants bound to Envs with CD4bs glycans present provide insights into N-glycan accommodation, including structural modes of light chain adaptations in the presence of the N276 glycan. Together, these findings reveal Env regions critical for germline antibody recognition and potential sites to alter in immunogen design. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_26492.map.gz emd_26492.map.gz | 6.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-26492-v30.xml emd-26492-v30.xml emd-26492.xml emd-26492.xml | 22.9 KB 22.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_26492.png emd_26492.png | 103.8 KB | ||
| Others |  emd_26492_additional_1.map.gz emd_26492_additional_1.map.gz emd_26492_half_map_1.map.gz emd_26492_half_map_1.map.gz emd_26492_half_map_2.map.gz emd_26492_half_map_2.map.gz | 59.8 MB 49.2 MB 49.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-26492 http://ftp.pdbj.org/pub/emdb/structures/EMD-26492 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26492 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-26492 | HTTPS FTP |
-Validation report
| Summary document |  emd_26492_validation.pdf.gz emd_26492_validation.pdf.gz | 569.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_26492_full_validation.pdf.gz emd_26492_full_validation.pdf.gz | 569 KB | Display | |
| Data in XML |  emd_26492_validation.xml.gz emd_26492_validation.xml.gz | 12.2 KB | Display | |
| Data in CIF |  emd_26492_validation.cif.gz emd_26492_validation.cif.gz | 14.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26492 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26492 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26492 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-26492 | HTTPS FTP |
-Related structure data
| Related structure data |  7ugoMC  7ugmC  7ugnC  7ugpC  7ugqC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_26492.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_26492.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.14 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Local refinement map used to build BG24-iGL CDR3mat Fab CDRL1
| File | emd_26492_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local refinement map used to build BG24-iGL CDR3mat Fab CDRL1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_26492_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_26492_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM map of BG24-iGL CDR3mat and 10-1074 Fabs in complex with ...
| Entire | Name: Cryo-EM map of BG24-iGL CDR3mat and 10-1074 Fabs in complex with BG505 SOSIP.664v4.1-GT1 Env trimer immunogen |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM map of BG24-iGL CDR3mat and 10-1074 Fabs in complex with ...
| Supramolecule | Name: Cryo-EM map of BG24-iGL CDR3mat and 10-1074 Fabs in complex with BG505 SOSIP.664v4.1-GT1 Env trimer immunogen type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
-Macromolecule #1: Envelope glycoprotein gp120
| Macromolecule | Name: Envelope glycoprotein gp120 / type: protein_or_peptide / ID: 1 / Details: Env mimic BG505-SOSIPv4.1-GT1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 52.322422 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: ENLWVTVYYG VPVWKDAETT LFCASDAKAY ETKKHNVWAT HACVPTDPNP QEIHLENVTE EFNMWKNNMV EQMHTDIISL WDQSLKPCV KLTPLCVTLQ CTNVTNNITD DMRGELKNCS FNMTTELRDK RQKVHALFYK LDIVPINENQ NTSYRLINCN T AAITQACP ...String: ENLWVTVYYG VPVWKDAETT LFCASDAKAY ETKKHNVWAT HACVPTDPNP QEIHLENVTE EFNMWKNNMV EQMHTDIISL WDQSLKPCV KLTPLCVTLQ CTNVTNNITD DMRGELKNCS FNMTTELRDK RQKVHALFYK LDIVPINENQ NTSYRLINCN T AAITQACP KVSFEPIPIH YCAPAGFAIL KCKDKKFNGT GPCPSVSTVQ CTHGIKPVVS TQLLLNGSLA EEEVMIRSED IR NNAKNIL VQFNTPVQIN CTRPNNNTRK SIRIGPGQWF YATGDIIGDI RQAHCNVSKA TWNETLGKVV KQLRKHFGNN TII RFANSS GGDLEVTTHS FNCGGEFFYC DTSGLFNSTW ISNTSVQGSN STGSNDSITL PCRIKQIINM WQRIGQAMYA PPIQ GVIRC VSNITGLILT RDGGSTDSTT ETFRPSGGDM RDNWRSELYK YKVVKIEPLG VAPTRCKRRV V |
-Macromolecule #2: Envelope glycoprotein gp41
| Macromolecule | Name: Envelope glycoprotein gp41 / type: protein_or_peptide / ID: 2 / Details: Env mimic BG505-SOSIPv4.1-GT1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 14.676723 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: VFLGFLGAAG STMGAASMTL TVQARNLLSL LKLTVWGIKQ LQARVLAVER YLRDQQLLGI WGCSGKLICC TNVPWNSSWS NRNLSEIWD NMTWLQWDKE ISNYTQIIYG LLEESQNQQE KNEQDLLALD |
-Macromolecule #3: 10-1074 Fab heavy chain
| Macromolecule | Name: 10-1074 Fab heavy chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 14.791433 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLQESGPG LVKPSETLSV TCSVSGDSMN NYYWTWIRQS PGKGLEWIGY ISDRESATYN PSLNSRVVIS RDTSKNQLSL KLNSVTPAD TAVYYCATAR RGQRIYGVVS FGEFFYYYSM DVWGKGTTVT VSSA |
-Macromolecule #4: 10-1074 Fab light chain
| Macromolecule | Name: 10-1074 Fab light chain / type: protein_or_peptide / ID: 4 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.701023 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: VRPLSVALGE TARISCGRQA LGSRAVQWYQ HRPGQAPILL IYNNQDRPSG IPERFSGTPD INFGTRATLT ISGVEAGDEA DYYCHMWDS RSGFSWSFGG ATRLTVLG |
-Macromolecule #5: BG24 inferred germline Fab with mature CDR3s heavy chain
| Macromolecule | Name: BG24 inferred germline Fab with mature CDR3s heavy chain type: protein_or_peptide / ID: 5 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 13.697314 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLVQSGAE VKKPGASVKV SCKASGYTFT GYYMHWVRQA PGQGLEWMGW INPNSGGTNY AQKFQGRVTM TRDTSISTAY MELSRLRSD DTAVYYCATQ VKLDSSAGYP FDIWGQGTMV TVSSAS |
-Macromolecule #6: BG24 inferred germline Fab with mature CDR3s light chain
| Macromolecule | Name: BG24 inferred germline Fab with mature CDR3s light chain type: protein_or_peptide / ID: 6 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 11.180226 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QSALTQPRSV SGSPGQSVTI SCTGTSSDVG GYNYVSWYQQ HPGKAPKLMI YDVSKRPSGV PDRFSGSKSG NTASLTISGL QAEDEADYY CSAFEYFGGG TKLTVLS |
-Macromolecule #14: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 14 / Number of copies: 21 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #15: alpha-D-mannopyranose
| Macromolecule | Name: alpha-D-mannopyranose / type: ligand / ID: 15 / Number of copies: 4 / Formula: MAN |
|---|---|
| Molecular weight | Theoretical: 180.156 Da |
| Chemical component information |  ChemComp-MAN: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.2 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: Details: Closed conformation Env trimer (gp120/gp41 subunits) low-pass filtered to 80 angstrom |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 225140 |
| Initial angle assignment | Type: RANDOM ASSIGNMENT |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller









 Z
Z Y
Y X
X