[English] 日本語
 Yorodumi
Yorodumi- EMDB-25882: Cryo-EM structure of respiratory super-complex CI+III2 from Tetra... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of respiratory super-complex CI+III2 from Tetrahymena thermophila | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationlipid-A-disaccharide synthase / lipid-A-disaccharide synthase activity / NADH:ubiquinone reductase (H+-translocating) / quinol-cytochrome-c reductase / medium-chain fatty acid-CoA ligase activity / P450-containing electron transport chain / protein processing involved in protein targeting to mitochondrion / ubiquinone-6 biosynthetic process / : / oxidoreductase activity, acting on NAD(P)H ...lipid-A-disaccharide synthase / lipid-A-disaccharide synthase activity / NADH:ubiquinone reductase (H+-translocating) / quinol-cytochrome-c reductase / medium-chain fatty acid-CoA ligase activity / P450-containing electron transport chain / protein processing involved in protein targeting to mitochondrion / ubiquinone-6 biosynthetic process / : / oxidoreductase activity, acting on NAD(P)H / ubiquinol-cytochrome-c reductase activity / lipid A biosynthetic process / mitochondrial electron transport, ubiquinol to cytochrome c / NADH:ubiquinone reductase (H+-translocating) / ubiquinone binding / NADH dehydrogenase activity / chloroplast thylakoid membrane / : / mitochondrial electron transport, NADH to ubiquinone / mitochondrial respiratory chain complex I assembly / electron transport coupled proton transport / acyl binding / acyl carrier activity / NADH dehydrogenase (ubiquinone) activity / quinone binding / : / ATP synthesis coupled electron transport / aerobic respiration / respiratory electron transport chain / chloroplast / mitochondrion organization / fatty acid metabolic process / mitochondrial membrane / electron transport chain / phospholipid binding / metalloendopeptidase activity / 2 iron, 2 sulfur cluster binding / NAD binding / FMN binding / 4 iron, 4 sulfur cluster binding / response to oxidative stress / membrane => GO:0016020 / mitochondrial inner membrane / electron transfer activity / oxidoreductase activity / ribosome / heme binding / protein-containing complex binding / negative regulation of apoptotic process / protein kinase binding / mitochondrion / membrane / metal ion binding / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
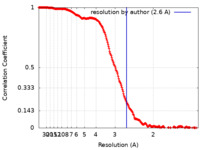
| Method | single particle reconstruction / cryo EM / Resolution: 2.6 Å | |||||||||
 Authors Authors | Zhou L / Maldonado M / Padavannil A / Guo F / Letts JA | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2022 Journal: Science / Year: 2022Title: Structures of 's respiratory chain reveal the diversity of eukaryotic core metabolism. Authors: Long Zhou / María Maldonado / Abhilash Padavannil / Fei Guo / James A Letts /   Abstract: Respiration is a core biological energy-converting process whose last steps are carried out by a chain of multisubunit complexes in the inner mitochondrial membrane. To probe the functional and ...Respiration is a core biological energy-converting process whose last steps are carried out by a chain of multisubunit complexes in the inner mitochondrial membrane. To probe the functional and structural diversity of eukaryotic respiration, we examined the respiratory chain of the ciliate (Tt). Using cryo-electron microscopy on a mixed sample, we solved structures of a supercomplex between Tt complex I (Tt-CI) and Tt-CIII (Tt-SC I+III) and a structure of Tt-CIV. Tt-SC I+III (~2.3 megadaltons) is a curved assembly with structural and functional symmetry breaking. Tt-CIV is a ~2.7-megadalton dimer with more than 50 subunits per protomer, including mitochondrial carriers and a TIM8-TIM13-like domain. Our structural and functional study of the respiratory chain reveals divergence in key components of eukaryotic respiration, thereby expanding our understanding of core metabolism. | |||||||||
| History |
|
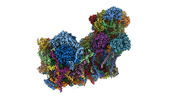
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25882.map.gz emd_25882.map.gz | 727.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25882-v30.xml emd-25882-v30.xml emd-25882.xml emd-25882.xml | 101.8 KB 101.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25882_fsc.xml emd_25882_fsc.xml | 20.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_25882.png emd_25882.png | 69.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25882 http://ftp.pdbj.org/pub/emdb/structures/EMD-25882 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25882 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25882 | HTTPS FTP |
-Validation report
| Summary document |  emd_25882_validation.pdf.gz emd_25882_validation.pdf.gz | 552.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_25882_full_validation.pdf.gz emd_25882_full_validation.pdf.gz | 552.2 KB | Display | |
| Data in XML |  emd_25882_validation.xml.gz emd_25882_validation.xml.gz | 18 KB | Display | |
| Data in CIF |  emd_25882_validation.cif.gz emd_25882_validation.cif.gz | 25.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25882 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25882 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25882 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25882 | HTTPS FTP |
-Related structure data
| Related structure data |  7tghMC  7w5zC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
| EM raw data |  EMPIAR-10919 (Title: Single particle cryogenic electron micrographs of tetrahymena mitochondrial electron transport chain complexes EMPIAR-10919 (Title: Single particle cryogenic electron micrographs of tetrahymena mitochondrial electron transport chain complexesData size: 7.8 TB Data #1: Unaligned multiframe micrographs of Tetrahymena respiratory chain complexes - Krios data [micrographs - multiframe] Data #2: Non-dose weighted motion corrected micrographs of Tetrahymena respiratory chain complexes - Krios data [micrographs - single frame] Data #3: Dose-weighted motion corrected micrographs of Tetrahymena respiratory chain complexes - Krios data [micrographs - single frame]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25882.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25882.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.835 Å | ||||||||||||||||||||||||||||||||||||
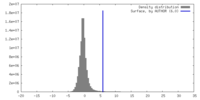
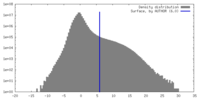
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
+Entire : Mitochondrial Super-complex I+III2 of Tetrahymena thermophila
+Supramolecule #1: Mitochondrial Super-complex I+III2 of Tetrahymena thermophila
+Macromolecule #1: NADH-ubiquinone oxidoreductase chain 1
+Macromolecule #2: NADH dehydrogenase subunit 1
+Macromolecule #3: Ymf65
+Macromolecule #4: NADH dehydrogenase subunit 2
+Macromolecule #5: NADH-ubiquinone oxidoreductase chain 3
+Macromolecule #6: M16 family peptidase, putative
+Macromolecule #7: Peptidase M16 inactive domain protein
+Macromolecule #8: Apocytochrome b
+Macromolecule #9: Cytochrome protein c1
+Macromolecule #10: Rieske iron-sulfur protein, ubiquinol-cytochrome C reductase iron...
+Macromolecule #11: Ubiquinol-cytochrome C reductase hinge protein
+Macromolecule #12: UQCRB
+Macromolecule #13: Transmembrane protein, putative
+Macromolecule #14: Transmembrane protein, putative
+Macromolecule #15: Transmembrane protein, putative
+Macromolecule #16: UNK1
+Macromolecule #17: UNK2
+Macromolecule #18: UNK3
+Macromolecule #19: NADH-ubiquinone oxidoreductase chain 4
+Macromolecule #20: Ymf58
+Macromolecule #21: NADH dehydrogenase subunit 5
+Macromolecule #22: Ymf57
+Macromolecule #23: Ymf62
+Macromolecule #24: Ribosomal protein L51/S25/CI-B8 domain protein
+Macromolecule #25: ETC complex I subunit motif protein
+Macromolecule #26: NADH dehydrogenase, putative
+Macromolecule #27: NDUA7
+Macromolecule #28: NAD-dependent epimerase/dehydratase family protein
+Macromolecule #29: Acyl carrier protein
+Macromolecule #30: Acyl carrier protein
+Macromolecule #31: NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 12
+Macromolecule #32: NDUA13
+Macromolecule #33: NDUB7
+Macromolecule #34: NDUB8
+Macromolecule #35: NDUB10
+Macromolecule #36: UNK4
+Macromolecule #37: Gamma-carbonic anhydrase
+Macromolecule #38: Gamma-carbonic anhydrase
+Macromolecule #39: Transcription factor apfi protein, putative
+Macromolecule #40: Transmembrane protein, putative
+Macromolecule #41: DnaJ domain protein
+Macromolecule #42: 2 iron, 2 sulfur cluster-binding protein
+Macromolecule #43: Acyl-CoA synthetase (AMP-forming)/AMP-acid ligase II
+Macromolecule #44: Lipid-A-disaccharide synthase
+Macromolecule #45: NDUA1
+Macromolecule #46: NADH-ubiquinone oxidoreductase complex I, 21 kDa subunit
+Macromolecule #47: NDUTT6
+Macromolecule #48: Transmembrane protein, putative
+Macromolecule #49: UNK5
+Macromolecule #50: NADH-ubiquinone oxidoreductase 75 kDa subunit
+Macromolecule #51: NADH dehydrogenase subunit 7
+Macromolecule #52: NADH dehydrogenase subunit 9
+Macromolecule #53: NADH dehydrogenase [ubiquinone] iron-sulfur protein 4, mitochondrial
+Macromolecule #54: Zinc-finger protein
+Macromolecule #55: NADH dehydrogenase subunit 10
+Macromolecule #56: NADH-ubiquinone oxidoreductase 1, chain, putative
+Macromolecule #57: NDUB9
+Macromolecule #58: Thioredoxin
+Macromolecule #59: NDUA8
+Macromolecule #60: Transmembrane protein, putative
+Macromolecule #61: NADH dehydrogenase [ubiquinone] flavoprotein 1, mitochondrial
+Macromolecule #62: NADH-ubiquinone oxidoreductase 24 kDa subunit
+Macromolecule #63: NDUB6
+Macromolecule #64: Transmembrane protein, putative
+Macromolecule #65: NDUC2
+Macromolecule #66: Transmembrane protein, putative
+Macromolecule #67: NDUB4
+Macromolecule #68: NDUTT4
+Macromolecule #69: NDUTT8
+Macromolecule #70: NDUB2
+Macromolecule #71: NDUTT3
+Macromolecule #72: Transmembrane protein, putative
+Macromolecule #73: Transmembrane protein, putative
+Macromolecule #74: NDUTT10
+Macromolecule #75: GRAM domain protein
+Macromolecule #76: NDUTT12
+Macromolecule #77: NDUPH2
+Macromolecule #78: Transmembrane protein, putative
+Macromolecule #79: NDUTT9
+Macromolecule #80: Transmembrane protein, putative
+Macromolecule #81: Transmembrane protein, putative
+Macromolecule #82: CARDIOLIPIN
+Macromolecule #83: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE
+Macromolecule #84: PROTOPORPHYRIN IX CONTAINING FE
+Macromolecule #85: UBIQUINONE-10
+Macromolecule #86: HEME C
+Macromolecule #87: FE2/S2 (INORGANIC) CLUSTER
+Macromolecule #88: 1,2-Distearoyl-sn-glycerophosphoethanolamine
+Macromolecule #89: ZINC ION
+Macromolecule #90: NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE
+Macromolecule #91: S-[2-({N-[(2S)-2-hydroxy-3,3-dimethyl-4-(phosphonooxy)butanoyl]-b...
+Macromolecule #92: ADENOSINE-5'-DIPHOSPHATE
+Macromolecule #93: IRON/SULFUR CLUSTER
+Macromolecule #94: FLAVIN MONONUCLEOTIDE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
Details: Solution were made fresh from concentrated stocks, filtered and de-gassed before equilibration onto Superose6 column | |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Details: 30 mA | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK III Details: incubation time before blotting: 60s, blot time: 9s, blot force: 25, offset: -2, incubation after blotting: 0s. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 7895 / Average exposure time: 5.9 sec. / Average electron dose: 66.69 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 58616 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-7tgh: |
 Movie
Movie Controller
Controller



































 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)