+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-25127 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Xenopus laevis CRL2Lrr1 (State 1) | ||||||||||||
 Map data Map data | It's a composite map made with State1-map1, State1-map2 and State1-map3. | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Cullin RING E3 ubiquitin ligase / DNA replication termination / LIGASE | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationcullin-RING ubiquitin ligase complex / elongin complex / VCB complex / transcription elongation by RNA polymerase II / protein-macromolecule adaptor activity / ubiquitin-dependent protein catabolic process / protein ubiquitination / ubiquitin protein ligase binding / zinc ion binding Similarity search - Function | ||||||||||||
| Biological species | |||||||||||||
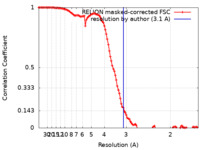
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | ||||||||||||
 Authors Authors | Zhou H / Brown A | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2021 Journal: Nucleic Acids Res / Year: 2021Title: Structure of CRL2Lrr1, the E3 ubiquitin ligase that promotes DNA replication termination in vertebrates. Authors: Haixia Zhou / Manal S Zaher / Johannes C Walter / Alan Brown /  Abstract: When vertebrate replisomes from neighboring origins converge, the Mcm7 subunit of the replicative helicase, CMG, is ubiquitylated by the E3 ubiquitin ligase, CRL2Lrr1. Polyubiquitylated CMG is then ...When vertebrate replisomes from neighboring origins converge, the Mcm7 subunit of the replicative helicase, CMG, is ubiquitylated by the E3 ubiquitin ligase, CRL2Lrr1. Polyubiquitylated CMG is then disassembled by the p97 ATPase, leading to replication termination. To avoid premature replisome disassembly, CRL2Lrr1 is only recruited to CMGs after they converge, but the underlying mechanism is unclear. Here, we use cryogenic electron microscopy to determine structures of recombinant Xenopus laevis CRL2Lrr1 with and without neddylation. The structures reveal that CRL2Lrr1 adopts an unusually open architecture, in which the putative substrate-recognition subunit, Lrr1, is located far from the catalytic module that catalyzes ubiquitin transfer. We further demonstrate that a predicted, flexible pleckstrin homology domain at the N-terminus of Lrr1 is essential to target CRL2Lrr1 to terminated CMGs. We propose a hypothetical model that explains how CRL2Lrr1's catalytic module is positioned next to the ubiquitylation site on Mcm7, and why CRL2Lrr1 binds CMG only after replisomes converge. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25127.map.gz emd_25127.map.gz | 113.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25127-v30.xml emd-25127-v30.xml emd-25127.xml emd-25127.xml | 23.8 KB 23.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25127_fsc.xml emd_25127_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_25127.png emd_25127.png | 75.7 KB | ||
| Masks |  emd_25127_msk_1.map emd_25127_msk_1.map emd_25127_msk_2.map emd_25127_msk_2.map emd_25127_msk_3.map emd_25127_msk_3.map | 125 MB 125 MB 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-25127.cif.gz emd-25127.cif.gz | 7 KB | ||
| Others |  emd_25127_additional_1.map.gz emd_25127_additional_1.map.gz emd_25127_additional_2.map.gz emd_25127_additional_2.map.gz emd_25127_additional_3.map.gz emd_25127_additional_3.map.gz | 4.4 MB 113.9 MB 3.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25127 http://ftp.pdbj.org/pub/emdb/structures/EMD-25127 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25127 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25127 | HTTPS FTP |
-Validation report
| Summary document |  emd_25127_validation.pdf.gz emd_25127_validation.pdf.gz | 537.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_25127_full_validation.pdf.gz emd_25127_full_validation.pdf.gz | 536.9 KB | Display | |
| Data in XML |  emd_25127_validation.xml.gz emd_25127_validation.xml.gz | 12.4 KB | Display | |
| Data in CIF |  emd_25127_validation.cif.gz emd_25127_validation.cif.gz | 16.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25127 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25127 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25127 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-25127 | HTTPS FTP |
-Related structure data
| Related structure data |  7shkMC  7shlC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25127.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25127.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | It's a composite map made with State1-map1, State1-map2 and State1-map3. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.825 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_25127_msk_1.map emd_25127_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Mask #2
| File |  emd_25127_msk_2.map emd_25127_msk_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Mask #3
| File |  emd_25127_msk_3.map emd_25127_msk_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #3
| File | emd_25127_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #2
| File | emd_25127_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_25127_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CRL2Lrr1
| Entire | Name: CRL2Lrr1 |
|---|---|
| Components |
|
-Supramolecule #1: CRL2Lrr1
| Supramolecule | Name: CRL2Lrr1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism: |
-Macromolecule #1: CULLIN_2 domain-containing protein
| Macromolecule | Name: CULLIN_2 domain-containing protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 87.244117 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSLKPRVVDF DETWNKLLTT IKAVVMLDYV ERATWNDRFS DIYALCVAYP EPLGERLYTE TKIFLENHVQ QLHTRVLDSA EQVLVMYFR YWEEYSRGAD YMDCLYRYLN TQYIKKNKLT EADLQYGYGG VDMNEPLMEI GELALDLWRK LMIEPLQDTL L IMLLREIK ...String: MSLKPRVVDF DETWNKLLTT IKAVVMLDYV ERATWNDRFS DIYALCVAYP EPLGERLYTE TKIFLENHVQ QLHTRVLDSA EQVLVMYFR YWEEYSRGAD YMDCLYRYLN TQYIKKNKLT EADLQYGYGG VDMNEPLMEI GELALDLWRK LMIEPLQDTL L IMLLREIK RDRCGEDPNQ KVIHGVINSF VHVEQYKKKF PLKFYQEIFE SPFLAETGEY YKQEASNLLQ ESNCSQYMEK IL GRLKDEE IRCRKYLHPS SYNKVIHECQ QRMVADHLQF LHAECHNIIR QERRNDMANM YTLLRAVSSG LPHMIQELQN HIH DEGLRA ISNLSQENMP TQFVESVLEV HSKFVQLVNC VLNGDQHFMS ALDKALTCVV NYREPKSVCK APELLAKYCD NMLK KSAKG MTENEVEDKL TSFITVFKYI DDKDVFQKFY ARMLAKRLIH GLSMSMDSEE TMINKLKQAC GYEFTSKLHR MYTDM SVSA DLNNKFNNFI KSQDTVIDLG ISFQIYVLQA GAWPLTQAPS STFAIPQELE KSVQMFELFY NQHFSGRKLT WLHYLC TGE VKMNYLCKPY VAMVTTYQMA VLLAFNNSEI ITYKELQDST QMNEKELTKT IKSLLDVKMI NHDSDKEDIE GESTFSL NM NFSSKRTKFK ITTPMQKDTP QEVEQTRSAV DEDRKMYLQA AIVRIMKARK VLRHNALIQE VISQSRARFN PSISMIKK C IEVLIDKQYI ERSQASADEY SYVA UniProtKB: CULLIN_2 domain-containing protein |
-Macromolecule #2: Tceb2-prov protein
| Macromolecule | Name: Tceb2-prov protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 13.325927 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDVFLMIRHH KTTIFTDAKE NTTVYELKRI VEGILKRPPE DQKLYKDDQL LDDNKTLGDC GFTSQTARPQ APATVGLAFR SSGDSFEPL RVEPFSSPPE LPDVMKPQET SGSANEQAVQ UniProtKB: Elongin B L homeolog |
-Macromolecule #3: Elongin-C
| Macromolecule | Name: Elongin-C / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 12.485135 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDGEEKTYGG CEGPDAMYVK LISSDGHEFI VKREHALTSG TIKAMLSGPG QFAENETNEV NFREIPSHVL SKVCMYFTYK VRYTNSSTE IPEFPIAPEI ALELLMAANF LDC UniProtKB: Elongin-C |
-Macromolecule #4: Lrr1
| Macromolecule | Name: Lrr1 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 46.882258 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKLQCEVEVI NRMLPTFGLK NRGKGTRAVL SVGRQEGKRG AAYLMICTLK DKSGSRYKLE NNIEQLFTRF VGEGKATLRL KEPALDICL SKAEICGLRN FISTVGLANK GTDIGTVSLP RLTPAKTSEI EKPRSKLFIT TKKDYPITKS FPYSLEHLQV S YCKLARVD ...String: MKLQCEVEVI NRMLPTFGLK NRGKGTRAVL SVGRQEGKRG AAYLMICTLK DKSGSRYKLE NNIEQLFTRF VGEGKATLRL KEPALDICL SKAEICGLRN FISTVGLANK GTDIGTVSLP RLTPAKTSEI EKPRSKLFIT TKKDYPITKS FPYSLEHLQV S YCKLARVD MRMLCLKKLQ KLDLSNNHIK KLPKTIGDLV CLQELILNHN FLESFEVVLC STTLRDTLKS LDLSANKLKA LP VQICNFK ELVSLKLDEN ELLQLPFPIG QLSKLRFLSA TKNNLQCLPN TFKKLTLENL DLFGNPFMQA TPLVPDIQLK IPL PLLETA ARATLKYRIP YGPHLIPATL CQDLSLAKTC DCGLPCLNSF IQTIVLMNLH QVSQTVVLVD TMGGTDGPIV CYFC SLTCY SQFLDKYLQS TRV UniProtKB: Leucine rich repeat protein 1 L homeolog isoform X1 |
-Macromolecule #5: RING-type domain-containing protein
| Macromolecule | Name: RING-type domain-containing protein / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 12.277985 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAAAMDVDTP SGANNSAGKK RFEVKKWNAV ALWAWDIVVD NCAICRNHIM DLCIECQANQ ASATSEECTV AWGVCNHAFH FHCISRWLK TRQVCPLDNR EWEFQKYVG UniProtKB: RING-type domain-containing protein |
-Macromolecule #6: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 6 / Number of copies: 4 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.8 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: TCEP is freshly added. | ||||||||||||
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 15 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.039 kPa | ||||||||||||
| Vitrification | Cryogen name: NITROGEN / Chamber humidity: 100 % / Chamber temperature: 281.2 K / Instrument: FEI VITROBOT MARK IV / Details: Blot 6 seconds with the force 12. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 54.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: OTHER / Target criteria: Correlation coefficient |
|---|---|
| Output model |  PDB-7shk: |
 Movie
Movie Controller
Controller



















 Z
Z Y
Y X
X