[English] 日本語
 Yorodumi
Yorodumi- EMDB-23264: Cryo-EM map of PDX1.2/PDX1.3 co-expression complex (Arabidopsis t... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23264 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM map of PDX1.2/PDX1.3 co-expression complex (Arabidopsis thaliana) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationresponse to non-ionic osmotic stress / response to lipid hydroperoxide / chlorophyll metabolic process / pyridoxal 5'-phosphate synthase (glutamine hydrolysing) / pyridoxal 5'-phosphate synthase (glutamine hydrolysing) activity / pyridoxal phosphate biosynthetic process / hyperosmotic salinity response / amino acid metabolic process / response to UV-B / endomembrane system ...response to non-ionic osmotic stress / response to lipid hydroperoxide / chlorophyll metabolic process / pyridoxal 5'-phosphate synthase (glutamine hydrolysing) / pyridoxal 5'-phosphate synthase (glutamine hydrolysing) activity / pyridoxal phosphate biosynthetic process / hyperosmotic salinity response / amino acid metabolic process / response to UV-B / endomembrane system / response to salt stress / response to oxidative stress / protein heterodimerization activity / protein homodimerization activity / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
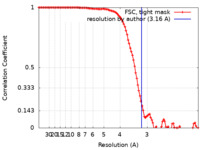
| Method | single particle reconstruction / cryo EM / Resolution: 3.16 Å | |||||||||
 Authors Authors | Novikova IV / Evans JE | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: ACS Chem Biol / Year: 2021 Journal: ACS Chem Biol / Year: 2021Title: Tunable Heteroassembly of a Plant Pseudoenzyme-Enzyme Complex. Authors: Irina V Novikova / Mowei Zhou / Chen Du / Marcelina Parra / Doo Nam Kim / Zachary L VanAernum / Jared B Shaw / Hanjo Hellmann / Vicki H Wysocki / James E Evans /  Abstract: Pseudoenzymes have emerged as key regulatory elements in all kingdoms of life despite being catalytically nonactive. Yet many factors defining why one protein is active while its homologue is ...Pseudoenzymes have emerged as key regulatory elements in all kingdoms of life despite being catalytically nonactive. Yet many factors defining why one protein is active while its homologue is inactive remain uncertain. For pseudoenzyme-enzyme pairs, the similarity of both subunits can often hinder conventional characterization approaches. In plants, a pseudoenzyme, PDX1.2, positively regulates vitamin B production by association with its active catalytic homologues such as PDX1.3 through an unknown assembly mechanism. Here we used an integrative experimental approach to learn that such pseudoenzyme-enzyme pair associations result in heterocomplexes of variable stoichiometry, which are unexpectedly tunable. We also present the atomic structure of the PDX1.2 pseudoenzyme as well as the population averaged PDX1.2-PDX1.3 pseudoenzyme-enzyme pair. Finally, we dissected hetero-dodecamers of each stoichiometry to understand the arrangement of monomers in the heterocomplexes and identified symmetry-imposed preferences in PDX1.2-PDX1.3 interactions. Our results provide a new model of pseudoenzyme-enzyme interactions and their native heterogeneity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23264.map.gz emd_23264.map.gz | 51.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23264-v30.xml emd-23264-v30.xml emd-23264.xml emd-23264.xml | 11.2 KB 11.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_23264_fsc.xml emd_23264_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_23264.png emd_23264.png | 82.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23264 http://ftp.pdbj.org/pub/emdb/structures/EMD-23264 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23264 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23264 | HTTPS FTP |
-Validation report
| Summary document |  emd_23264_validation.pdf.gz emd_23264_validation.pdf.gz | 361.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_23264_full_validation.pdf.gz emd_23264_full_validation.pdf.gz | 361.5 KB | Display | |
| Data in XML |  emd_23264_validation.xml.gz emd_23264_validation.xml.gz | 12 KB | Display | |
| Data in CIF |  emd_23264_validation.cif.gz emd_23264_validation.cif.gz | 15.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23264 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23264 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23264 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23264 | HTTPS FTP |
-Related structure data
| Related structure data |  7lb6MC  7lb5C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23264.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23264.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.012 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : PDX1.2/PDX1.3 heterododecamer
| Entire | Name: PDX1.2/PDX1.3 heterododecamer |
|---|---|
| Components |
|
-Supramolecule #1: PDX1.2/PDX1.3 heterododecamer
| Supramolecule | Name: PDX1.2/PDX1.3 heterododecamer / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Molecular weight | Experimental: 445 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 50 mM Tris, 150 mM NaCl, 4 mM. DTT |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR / Details: at 15 mA |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 85 % / Chamber temperature: 298 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 100.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller