+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20480 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human TRPM2 bound to ADPR and calcium | |||||||||
 Map data Map data | human TRPM2 bound with ADPR and calcium, unsharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TRPM2 channel / ADPR / calcium / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationmono-ADP-D-ribose binding / manganese ion transmembrane transporter activity / dendritic cell differentiation / zinc ion transmembrane transport / cellular response to temperature stimulus / regulation of filopodium assembly / response to purine-containing compound / ligand-gated calcium channel activity / response to hydroperoxide / dendritic cell chemotaxis ...mono-ADP-D-ribose binding / manganese ion transmembrane transporter activity / dendritic cell differentiation / zinc ion transmembrane transport / cellular response to temperature stimulus / regulation of filopodium assembly / response to purine-containing compound / ligand-gated calcium channel activity / response to hydroperoxide / dendritic cell chemotaxis / TRP channels / calcium ion transmembrane import into cytosol / sodium channel activity / temperature homeostasis / intracellularly gated calcium channel activity / tertiary granule membrane / calcium ion import across plasma membrane / ficolin-1-rich granule membrane / specific granule membrane / monoatomic cation channel activity / cell projection / release of sequestered calcium ion into cytosol / cellular response to calcium ion / cytoplasmic vesicle membrane / regulation of actin cytoskeleton organization / calcium ion transmembrane transport / calcium channel activity / cellular response to hydrogen peroxide / calcium ion transport / response to heat / protein homotetramerization / perikaryon / lysosome / lysosomal membrane / calcium ion binding / Neutrophil degranulation / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Du J / Lu W / Huang Y | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2019 Journal: Elife / Year: 2019Title: Ligand recognition and gating mechanism through three ligand-binding sites of human TRPM2 channel. Authors: Yihe Huang / Becca Roth / Wei Lü / Juan Du /  Abstract: TRPM2 is critically involved in diverse physiological processes including core temperature sensing, apoptosis, and immune response. TRPM2's activation by Ca and ADP ribose (ADPR), an NAD-metabolite ...TRPM2 is critically involved in diverse physiological processes including core temperature sensing, apoptosis, and immune response. TRPM2's activation by Ca and ADP ribose (ADPR), an NAD-metabolite produced under oxidative stress and neurodegenerative conditions, suggests a role in neurological disorders. We provide a central concept between triple-site ligand binding and the channel gating of human TRPM2. We show consecutive structural rearrangements and channel activation of TRPM2 induced by binding of ADPR in two indispensable locations, and the binding of Ca in the transmembrane domain. The 8-Br-cADPR-an antagonist of cADPR-binds only to the MHR1/2 domain and inhibits TRPM2 by stabilizing the channel in an apo-like conformation. We conclude that MHR1/2 acts as a orthostatic ligand-binding site for TRPM2. The NUDT9-H domain binds to a second ADPR to assist channel activation in vertebrates, but not necessary in invertebrates. Our work provides insights into the gating mechanism of human TRPM2 and its pharmacology. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20480.map.gz emd_20480.map.gz | 75.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20480-v30.xml emd-20480-v30.xml emd-20480.xml emd-20480.xml | 13.1 KB 13.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20480.png emd_20480.png | 235.8 KB | ||
| Filedesc metadata |  emd-20480.cif.gz emd-20480.cif.gz | 6.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20480 http://ftp.pdbj.org/pub/emdb/structures/EMD-20480 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20480 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20480 | HTTPS FTP |
-Related structure data
| Related structure data |  6pusMC  6puoC  6purC  6puuC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20480.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20480.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | human TRPM2 bound with ADPR and calcium, unsharpened map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.074 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : human TRPM2
| Entire | Name: human TRPM2 |
|---|---|
| Components |
|
-Supramolecule #1: human TRPM2
| Supramolecule | Name: human TRPM2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Transient receptor potential cation channel subfamily M member 2
| Macromolecule | Name: Transient receptor potential cation channel subfamily M member 2 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 172.280062 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GSGGRATMEP SALRKAGSEQ EEGFEGLPRR VTDLGMVSNL RRSNSSLFKS WRLQCPFGNN DKQESLSSWI PENIKKKECV YFVESSKLS DAGKVVCQCG YTHEQHLEEA TKPHTFQGTQ WDPKKHVQEM PTDAFGDIVF TGLSQKVKKY VRVSQDTPSS V IYHLMTQH ...String: GSGGRATMEP SALRKAGSEQ EEGFEGLPRR VTDLGMVSNL RRSNSSLFKS WRLQCPFGNN DKQESLSSWI PENIKKKECV YFVESSKLS DAGKVVCQCG YTHEQHLEEA TKPHTFQGTQ WDPKKHVQEM PTDAFGDIVF TGLSQKVKKY VRVSQDTPSS V IYHLMTQH WGLDVPNLLI SVTGGAKNFN MKPRLKSIFR RGLVKVAQTT GAWIITGGSH TGVMKQVGEA VRDFSLSSSY KE GELITIG VATWGTVHRR EGLIHPTGSF PAEYILDEDG QGNLTCLDSN HSHFILVDDG THGQYGVEIP LRTRLEKFIS EQT KERGGV AIKIPIVCVV LEGGPGTLHT IDNATTNGTP CVVVEGSGRV ADVIAQVANL PVSDITISLI QQKLSVFFQE MFET FTESR IVEWTKKIQD IVRRRQLLTV FREGKDGQQD VDVAILQALL KASRSQDHFG HENWDHQLKL AVAWNRVDIA RSEIF MDEW QWKPSDLHPT MTAALISNKP EFVKLFLENG VQLKEFVTWD TLLYLYENLD PSCLFHSKLQ KVLVEDPERP ACAPAA PRL QMHHVAQVLR ELLGDFTQPL YPRPRHNDRL RLLLPVPHVK LNVQGVSLRS LYKRSSGHVT FTMDPIRDLL IWAIVQN RR ELAGIIWAQS QDCIAAALAC SKILKELSKE EEDTDSSEEM LALAEEYEHR AIGVFTECYR KDEERAQKLL TRVSEAWG K TTCLQLALEA KDMKFVSHGG IQAFLTKVWW GQLSVDNGLW RVTLCMLAFP LLLTGLISFR EKRLQDVGTP AARARAFFT APVVVFHLNI LSYFAFLCLF AYVLMVDFQP VPSWCECAIY LWLFSLVCEE MRQLFYDPDE CGLMKKAALY FSDFWNKLDV GAILLFVAG LTCRLIPATL YPGRVILSLD FILFCLRLMH IFTISKTLGP KIIIVKRMMK DVFFFLFLLA VWVVSFGVAK Q AILIHNER RVDWLFRGAV YHSYLTIFGQ IPGYIDGVNF NPEHCSPNGT DPYKPKCPES DATQQRPAFP EWLTVLLLCL YL LFTNILL LNLLIAMFNY TFQQVQEHTD QIWKFQRHDL IEEYHGRPAA PPPFILLSHL QLFIKRVVLK TPAKRHKQLK NKL EKNEEA ALLSWEIYLK ENYLQNRQFQ QKQRPEQKIE DISNKVDAMV DLLDLDPLKR SGSMEQRLAS LEEQVAQTAQ ALHW IVRTL RASGFSSEAD VPTLASQKAA EEPDAEPGGR KKTEEPGDSY HVNARHLLYP NCPVTRFPVP NEKVPWETEF LIYDP PFYT AERKDAAAMD PMGDTLEPLS TIQYNVVDGL RDRRSFHGPY TVQAGLPLNP MGRTGLRGRG SLSCFGPNHT LYPMVT RWR RNEDGAICRK SIKKMLEVLV VKLPLSEHWA LPGGSREPGE MLPRKLKRIL RQEHWPSFEN LLKCGMEVYK GYMDDPR NT DNAWIETVAV SVHFQDQNDV ELNRLNSNLH ACDSGASIRW QVVDRRIPLY ANHKTLLQKA AAEFGAHYFE UniProtKB: Transient receptor potential cation channel subfamily M member 2 |
-Macromolecule #2: ADENOSINE-5-DIPHOSPHORIBOSE
| Macromolecule | Name: ADENOSINE-5-DIPHOSPHORIBOSE / type: ligand / ID: 2 / Number of copies: 8 / Formula: APR |
|---|---|
| Molecular weight | Theoretical: 559.316 Da |
| Chemical component information | 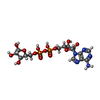 ChemComp-APR: |
-Macromolecule #3: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 3 / Number of copies: 4 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 1-40 / Average exposure time: 8.0 sec. / Average electron dose: 6.8 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER / Details: cryosparc |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.7 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 3) / Number images used: 287184 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 3) |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6pus: |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















