[English] 日本語
 Yorodumi
Yorodumi- EMDB-20092: Monomeric kinesin-1 motor domain in no-nucleotide state bound to ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20092 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Monomeric kinesin-1 motor domain in no-nucleotide state bound to GMPCPP-stabilized microtubule | |||||||||
 Map data Map data | Monomeric kinesin-1 motor domain in the no-nucleotide state bound to GMPCPP-stabilized microtubule | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Kinesin / Microtubule / MOTOR PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcytoplasm organization / cytolytic granule membrane / plus-end-directed vesicle transport along microtubule / mitocytosis / anterograde dendritic transport of neurotransmitter receptor complex / anterograde neuronal dense core vesicle transport / anterograde axonal protein transport / retrograde neuronal dense core vesicle transport / vesicle transport along microtubule / lysosome localization ...cytoplasm organization / cytolytic granule membrane / plus-end-directed vesicle transport along microtubule / mitocytosis / anterograde dendritic transport of neurotransmitter receptor complex / anterograde neuronal dense core vesicle transport / anterograde axonal protein transport / retrograde neuronal dense core vesicle transport / vesicle transport along microtubule / lysosome localization / positive regulation of potassium ion transport / natural killer cell mediated cytotoxicity / plus-end-directed microtubule motor activity / Kinesins / RHO GTPases activate KTN1 / stress granule disassembly / positive regulation of axon guidance / mitochondrion transport along microtubule / ciliary rootlet / COPI-dependent Golgi-to-ER retrograde traffic / centrosome localization / microtubule motor activity / kinesin complex / synaptic vesicle transport / Insulin processing / microtubule-based movement / microtubule-based process / centriolar satellite / axon cytoplasm / phagocytic vesicle / MHC class II antigen presentation / dendrite cytoplasm / regulation of membrane potential / axon guidance / positive regulation of synaptic transmission, GABAergic / positive regulation of protein localization to plasma membrane / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / structural constituent of cytoskeleton / cellular response to type II interferon / microtubule cytoskeleton organization / microtubule cytoskeleton / Signaling by ALK fusions and activated point mutants / nervous system development / mitotic cell cycle / microtubule binding / vesicle / microtubule / cadherin binding / protein heterodimerization activity / GTPase activity / protein-containing complex binding / GTP binding / perinuclear region of cytoplasm / ATP hydrolysis activity / mitochondrion / ATP binding / identical protein binding / membrane / metal ion binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.67 Å | |||||||||
 Authors Authors | Cha HK / Debs G | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structural Intermediates of the Dimeric Kinesin Stepping Cycle Revealed by Cryo-EM Authors: Cha HK / Debs G / Liu X / Liu D / Sindelar CV | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20092.map.gz emd_20092.map.gz | 230.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20092-v30.xml emd-20092-v30.xml emd-20092.xml emd-20092.xml | 12.2 KB 12.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20092.png emd_20092.png | 157.1 KB | ||
| Filedesc metadata |  emd-20092.cif.gz emd-20092.cif.gz | 6.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20092 http://ftp.pdbj.org/pub/emdb/structures/EMD-20092 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20092 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20092 | HTTPS FTP |
-Validation report
| Summary document |  emd_20092_validation.pdf.gz emd_20092_validation.pdf.gz | 402.1 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_20092_full_validation.pdf.gz emd_20092_full_validation.pdf.gz | 401.7 KB | Display | |
| Data in XML |  emd_20092_validation.xml.gz emd_20092_validation.xml.gz | 7.9 KB | Display | |
| Data in CIF |  emd_20092_validation.cif.gz emd_20092_validation.cif.gz | 9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20092 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20092 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20092 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20092 | HTTPS FTP |
-Related structure data
| Related structure data |  6ojqMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20092.map.gz / Format: CCP4 / Size: 371.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20092.map.gz / Format: CCP4 / Size: 371.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Monomeric kinesin-1 motor domain in the no-nucleotide state bound to GMPCPP-stabilized microtubule | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.3 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
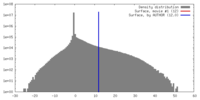
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Monomeric kinesin-1 motor domain in the no-nucleotide state bound...
| Entire | Name: Monomeric kinesin-1 motor domain in the no-nucleotide state bound to GMPCPP-stabilized microtubules |
|---|---|
| Components |
|
-Supramolecule #1: Monomeric kinesin-1 motor domain in the no-nucleotide state bound...
| Supramolecule | Name: Monomeric kinesin-1 motor domain in the no-nucleotide state bound to GMPCPP-stabilized microtubules type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Tubulin beta-2B chain
| Macromolecule | Name: Tubulin beta-2B chain / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 47.825859 KDa |
| Sequence | String: MREIVHIQAG QCGNQIGAKF WEVISDEHGI DPTGSYHGDS DLQLERINVY YNEAAGNKYV PRAILVDLEP GTMDSVRSGP FGQIFRPDN FVFGQSGAGN NWAKGHYTEG AELVDSVLDV VRKESESCDC LQGFQLTHSL GGGTGSGMGT LLISKIREEY P DRIMNTFS ...String: MREIVHIQAG QCGNQIGAKF WEVISDEHGI DPTGSYHGDS DLQLERINVY YNEAAGNKYV PRAILVDLEP GTMDSVRSGP FGQIFRPDN FVFGQSGAGN NWAKGHYTEG AELVDSVLDV VRKESESCDC LQGFQLTHSL GGGTGSGMGT LLISKIREEY P DRIMNTFS VVPSPKVSDT VVEPYNATLS VHQLVENTDE TYCIDNEALY DICFRTLKLT TPTYGDLNHL VSATMSGVTT CL RFPGQLN ADLRKLAVNM VPFPRLHFFM PGFAPLTSRG SQQYRALTVP ELTQQMFDAK NMMAACDPRH GRYLTVAAVF RGR MSMKEV DEQMLNVQNK NSSYFVEWIP NNVKTAVCDI PPRGLKMSAT FIGNSTAIQE LFKRISEQFT AMFRRKAFLH WYTG EGMDE MEFTEAESNM NDLVSEYQQY Q UniProtKB: Tubulin beta-2B chain |
-Macromolecule #2: Tubulin alpha-1B chain
| Macromolecule | Name: Tubulin alpha-1B chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 48.665027 KDa |
| Sequence | String: MRECISIHVG QAGVQIGNAC WELYCLEHGI QPDGQMPSDK TIGGGDDSFN TFFSETGAGK HVPRAVFVDL EPTVIDEVRT GTYRQLFHP EQLITGKEDA ANNYARGHYT IGKEIIDLVL DRIRKLADQC TGLQGFLVFH SFGGGTGSGF TSLLMERLSV D YGKKSKLE ...String: MRECISIHVG QAGVQIGNAC WELYCLEHGI QPDGQMPSDK TIGGGDDSFN TFFSETGAGK HVPRAVFVDL EPTVIDEVRT GTYRQLFHP EQLITGKEDA ANNYARGHYT IGKEIIDLVL DRIRKLADQC TGLQGFLVFH SFGGGTGSGF TSLLMERLSV D YGKKSKLE FSIYPAPQVS TAVVEPYNSI LTTHTTLEHS DCAFMVDNEA IYDICRRNLD IERPTYTNLN RLISQIVSSI TA SLRFDGA LNVDLTEFQT NLVPYPRIHF PLATYAPVIS AEKAYHEQLS VAEITNACFE PANQMVKCDP RHGKYMACCL LYR GDVVPK DVNAAIATIK TKRSIQFVDW CPTGFKVGIN YQPPTVVPGG DLAKVQRAVC MLSNTTAIAE AWARLDHKFD LMYA KRAFV HWYVGEGMEE GEFSEAREDM AALEKDYEEV GV UniProtKB: Tubulin alpha-1B chain |
-Macromolecule #3: Kinesin-1 heavy chain
| Macromolecule | Name: Kinesin-1 heavy chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 35.445801 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: NIKVMCRFRP LNESEVNRGD KYIAKFQGED TVVIASKPYA FDRVFQSSTS QEQVYNDAAK KIVKDVLEGY NGTIFAYGQT SSGKTHTME GKLHDPEGMG IIPRIVQDIF NYIYSMDENL EFHIKVSYFE IYLDKIRDLL DVSKTNLSVH EDKNRVPYVK G ATERFVSS ...String: NIKVMCRFRP LNESEVNRGD KYIAKFQGED TVVIASKPYA FDRVFQSSTS QEQVYNDAAK KIVKDVLEGY NGTIFAYGQT SSGKTHTME GKLHDPEGMG IIPRIVQDIF NYIYSMDENL EFHIKVSYFE IYLDKIRDLL DVSKTNLSVH EDKNRVPYVK G ATERFVSS PDEVMDTIDE GKSNRHVAVT NMNEHSSRSH SIFLINVKQE NTQTEQKLSG KLYLVDLAGS EKVSKTGAEG AV LDEAKNI NKSLSALGNV ISALAEGSTY VPYRDSKMTR ILQDSLGGNA RTTIVICCSP SSYNESETKS TLLFGQRAKT UniProtKB: Kinesin-1 heavy chain |
-Macromolecule #4: PHOSPHOMETHYLPHOSPHONIC ACID GUANYLATE ESTER
| Macromolecule | Name: PHOSPHOMETHYLPHOSPHONIC ACID GUANYLATE ESTER / type: ligand / ID: 4 / Number of copies: 1 / Formula: G2P |
|---|---|
| Molecular weight | Theoretical: 521.208 Da |
| Chemical component information |  ChemComp-G2P: |
-Macromolecule #5: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #6: GUANOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: GUANOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 6 / Number of copies: 1 / Formula: GTP |
|---|---|
| Molecular weight | Theoretical: 523.18 Da |
| Chemical component information |  ChemComp-GTP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 1.625 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.67 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 403424 |
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller

































 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)