[English] 日本語
 Yorodumi
Yorodumi- PDB-1tzf: X-ray Crystal Structure of alpha-D-glucose-1-phosphate cytidylylt... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1tzf | ||||||
|---|---|---|---|---|---|---|---|
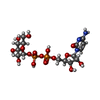
| Title | X-ray Crystal Structure of alpha-D-glucose-1-phosphate cytidylyltransferase from Salmonella typhi | ||||||
 Components Components | Glucose-1-phosphate cytidylyltransferase | ||||||
 Keywords Keywords | TRANSFERASE / nucleotidyltransferase / mixed alpha/beta fold | ||||||
| Function / homology |  Function and homology information Function and homology informationglucose-1-phosphate cytidylyltransferase / glucose-1-phosphate cytidylyltransferase activity / O antigen biosynthetic process / nucleotide binding / metal ion binding Similarity search - Function | ||||||
| Biological species |  Salmonella enterica subsp. enterica serovar Typhi (bacteria) Salmonella enterica subsp. enterica serovar Typhi (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MAD / Resolution: 2.1 Å MAD / Resolution: 2.1 Å | ||||||
 Authors Authors | Koropatkin, N.M. / Holden, H.M. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2004 Journal: J.Biol.Chem. / Year: 2004Title: Molecular structure of alpha-D-glucose-1-phosphate cytidylyltransferase from Salmonella typhi Authors: Koropatkin, N.M. / Holden, H.M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1tzf.cif.gz 1tzf.cif.gz | 69.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1tzf.ent.gz pdb1tzf.ent.gz | 50.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1tzf.json.gz 1tzf.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/tz/1tzf https://data.pdbj.org/pub/pdb/validation_reports/tz/1tzf ftp://data.pdbj.org/pub/pdb/validation_reports/tz/1tzf ftp://data.pdbj.org/pub/pdb/validation_reports/tz/1tzf | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | x 6
| ||||||||||
| Unit cell |
| ||||||||||
| Components on special symmetry positions |
| ||||||||||
| Details | The biological unit is a hexamer, generated from the monomer in the asymmetric unit by the operations: ROTATION MATRIX: -0.50000 -0.86603 0.00000 0.86603 -0.50000 0.00001 -0.00001 0.00000 1.00000 TRANSLATION VECTOR IN AS 0.00043 0.00017 0.00008 ROTATION MATRIX: -0.50000 0.86602 0.00000 -0.86602 -0.50000 0.00000 0.00000 0.00000 1.00000 TRANSLATION VECTOR IN AS 0.00046 0.00027 0.00009 ROTATION MATRIX: -1.00000 0.00000 0.00000 0.00000 1.00000 0.00001 0.00000 0.00001 -1.00000 TRANSLATION VECTOR IN AS 0.00020 -0.00001 81.14987 ROTATION MATRIX: 0.50000 0.86602 0.00000 0.86602 -0.50000 0.00000 0.00000 0.00000 -1.00000 TRANSLATION VECTOR IN AS 0.00082 0.00015 81.15000 ROTATION MATRIX: 0.50000 -0.86603 0.00000 -0.86603 -0.50000 0.00000 0.00000 0.00000 -1.00000 TRANSLATION VECTOR IN AS 0.00045 0.00058 81.14999 These are the rotation and translation operations to be performed on the original monomer to generate the next five subunits of the complete hexamer 'P 63 2 2' 1 1 0 0 0 1 0 0 0 1 0 0 0 0 0 0 'P 63 2 2' 3 0 -1 0 1 -1 0 0 0 1 0 0 0 0 0 0 'P 63 2 2' 5 -1 1 0 -1 0 0 0 0 1 0 0 0 0 0 0 'P 63 2 2' 7 1 -1 0 0 -1 0 0 0 -1 0 0 0 0 0 0 'P 63 2 2' 9 -1 0 0 -1 1 0 0 0 -1 0 0 0 0 0 0 'P 63 2 2' 11 0 1 0 1 0 0 0 0 -1 0 0 0 0 0 0 'P 63 2 2' 2 1 -1 0 1 0 0 0 0 1 0 0 0 0 1 2 'P 63 2 2' 4 -1 0 0 0 -1 0 0 0 1 0 0 0 0 1 2 'P 63 2 2' 6 0 1 0 -1 1 0 0 0 1 0 0 0 0 1 2 'P 63 2 2' 8 0 -1 0 -1 0 0 0 0 -1 0 0 0 0 1 2 'P 63 2 2' 10 -1 1 0 0 1 0 0 0 -1 0 0 0 0 1 2 'P 63 2 2' 12 1 0 0 1 -1 0 0 0 -1 0 0 0 0 1 2 |
- Components
Components
| #1: Protein | Mass: 29203.598 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Details: complexed with cytidyl-5'-phosphate-glucosyl-6-phosphate Source: (gene. exp.)  Salmonella enterica subsp. enterica serovar Typhi (bacteria) Salmonella enterica subsp. enterica serovar Typhi (bacteria)Species: Salmonella enterica / Strain: CT18 / Gene: rfbF / Plasmid: pET28a / Production host:  References: UniProt: P26396, UniProt: Q8Z5I4*PLUS, glucose-1-phosphate cytidylyltransferase |
|---|---|
| #2: Chemical | ChemComp-MG / |
| #3: Chemical | ChemComp-C5G / [ |
| #4: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3 Å3/Da / Density % sol: 58.3 % |
|---|---|
| Crystal grow | Temperature: 277 K / Method: vapor diffusion, hanging drop / pH: 5 Details: PEG 4000, Ammonium sulfate, sodium acetate, pH 5.0, VAPOR DIFFUSION, HANGING DROP, temperature 277K |
-Data collection
| Diffraction | Mean temperature: 130 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 19-ID / Wavelength: 0.9641 Å / Beamline: 19-ID / Wavelength: 0.9641 Å |
| Detector | Type: SBC-3 / Detector: CCD / Date: Mar 29, 2004 |
| Radiation | Monochromator: Double crystal Si-111 / Protocol: MAD / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9641 Å / Relative weight: 1 |
| Reflection | Resolution: 2.1→50 Å / Num. all: 22401 / Num. obs: 22358 / % possible obs: 98.5 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 14.1 % / Rsym value: 0.067 / Net I/σ(I): 65.3 |
| Reflection shell | Resolution: 2.1→2.18 Å / Redundancy: 5.4 % / Mean I/σ(I) obs: 9.9 / Num. unique all: 2163 / Rsym value: 0.176 / % possible all: 98.1 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MAD / Resolution: 2.1→50 Å / Cross valid method: THROUGHOUT / σ(F): 0 MAD / Resolution: 2.1→50 Å / Cross valid method: THROUGHOUT / σ(F): 0
| |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.1→50 Å
| |||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller








 PDBj
PDBj